Stuart A. Cook
Joint Semi-supervised 3D Super-Resolution and Segmentation with Mixed Adversarial Gaussian Domain Adaptation
Jul 16, 2021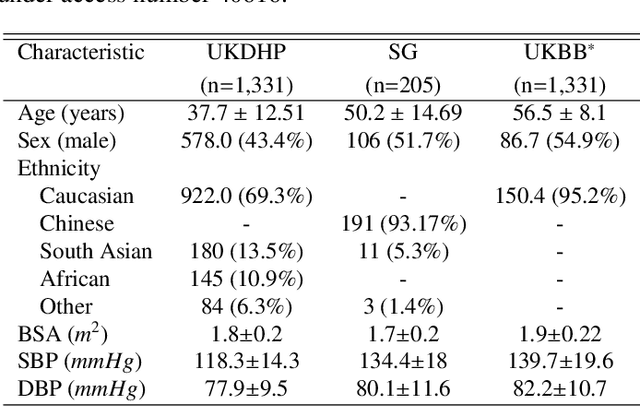
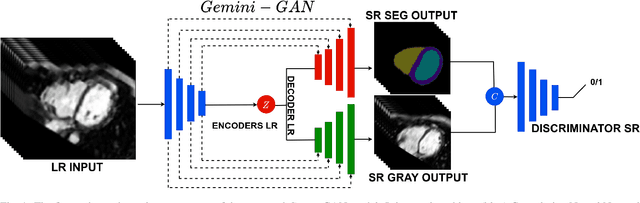
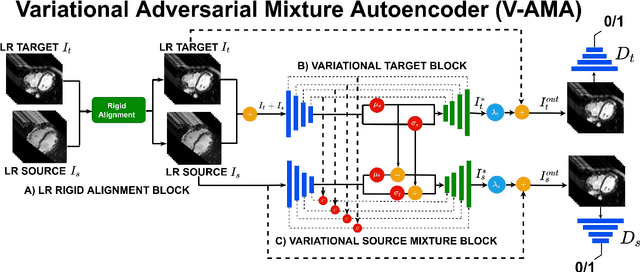
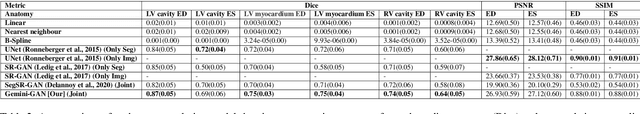
Abstract:Optimising the analysis of cardiac structure and function requires accurate 3D representations of shape and motion. However, techniques such as cardiac magnetic resonance imaging are conventionally limited to acquiring contiguous cross-sectional slices with low through-plane resolution and potential inter-slice spatial misalignment. Super-resolution in medical imaging aims to increase the resolution of images but is conventionally trained on features from low resolution datasets and does not super-resolve corresponding segmentations. Here we propose a semi-supervised multi-task generative adversarial network (Gemini-GAN) that performs joint super-resolution of the images and their labels using a ground truth of high resolution 3D cines and segmentations, while an unsupervised variational adversarial mixture autoencoder (V-AMA) is used for continuous domain adaptation. Our proposed approach is extensively evaluated on two transnational multi-ethnic populations of 1,331 and 205 adults respectively, delivering an improvement on state of the art methods in terms of Dice index, peak signal to noise ratio, and structural similarity index measure. This framework also exceeds the performance of state of the art generative domain adaptation models on external validation (Dice index 0.81 vs 0.74 for the left ventricle). This demonstrates how joint super-resolution and segmentation, trained on 3D ground-truth data with cross-domain generalization, enables robust precision phenotyping in diverse populations.
Explainable Shape Analysis through Deep Hierarchical Generative Models: Application to Cardiac Remodeling
Jun 28, 2019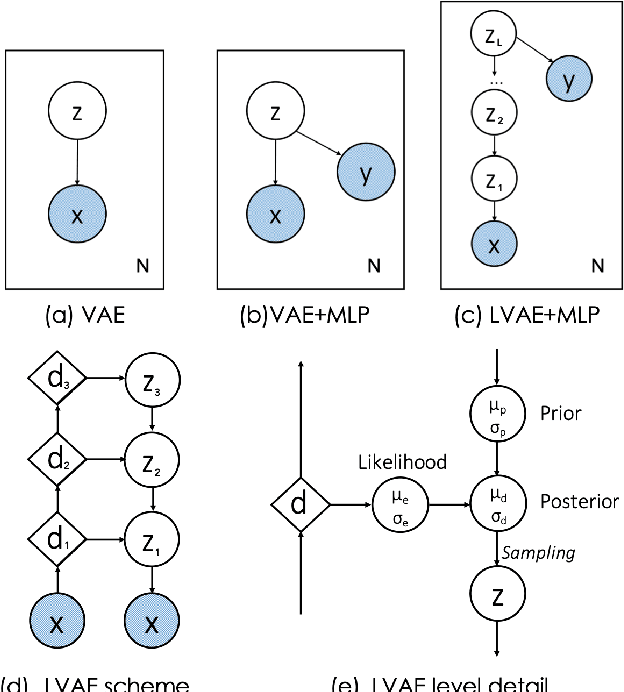
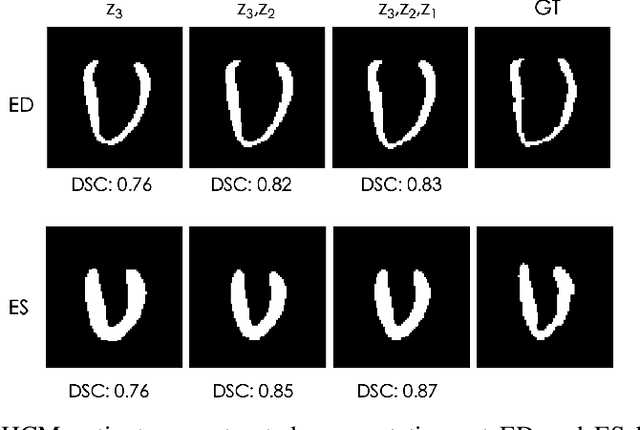
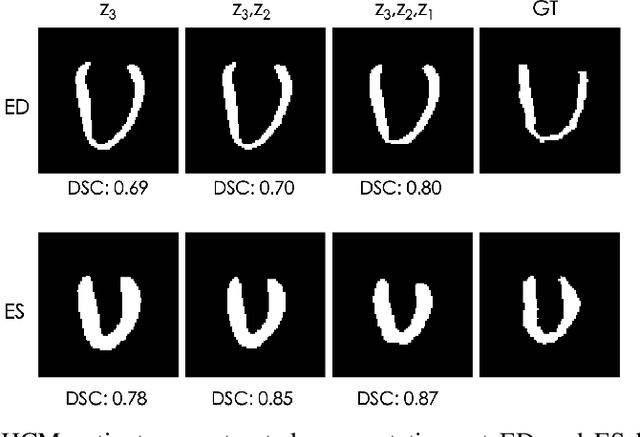
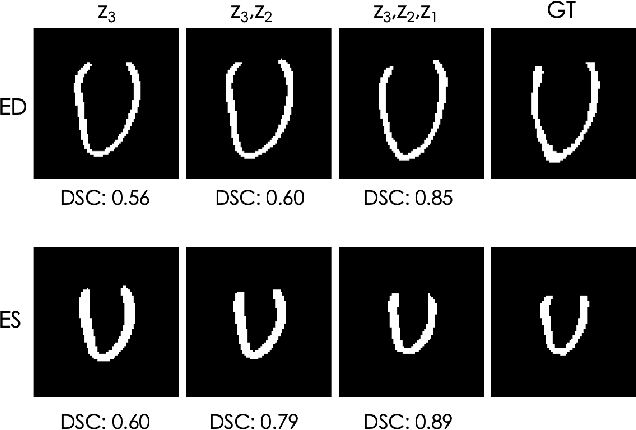


Abstract:Quantification of anatomical shape changes still relies on scalar global indexes which are largely insensitive to regional or asymmetric modifications. Accurate assessment of pathology-driven anatomical remodeling is a crucial step for the diagnosis and treatment of heart conditions. Deep learning approaches have recently achieved wide success in the analysis of medical images, but they lack interpretability in the feature extraction and decision processes. In this work, we propose a new interpretable deep learning model for shape analysis. In particular, we exploit deep generative networks to model a population of anatomical segmentations through a hierarchy of conditional latent variables. At the highest level of this hierarchy, a two-dimensional latent space is simultaneously optimised to discriminate distinct clinical conditions, enabling the direct visualisation of the classification space. Moreover, the anatomical variability encoded by this discriminative latent space can be visualised in the segmentation space thanks to the generative properties of the model, making the classification task transparent. This approach yielded high accuracy in the categorisation of healthy and remodelled hearts when tested on unseen segmentations from our own multi-centre dataset as well as in an external validation set. More importantly, it enabled the visualisation in three-dimensions of the most discriminative anatomical features between the two conditions. The proposed approach scales effectively to large populations, facilitating high-throughput analysis of normal anatomy and pathology in large-scale studies of volumetric imaging.
3D High-Resolution Cardiac Segmentation Reconstruction from 2D Views using Conditional Variational Autoencoders
Feb 28, 2019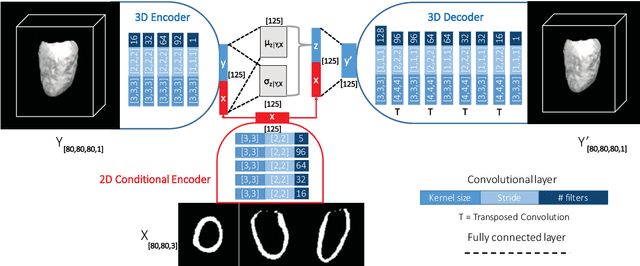


Abstract:Accurate segmentation of heart structures imaged by cardiac MR is key for the quantitative analysis of pathology. High-resolution 3D MR sequences enable whole-heart structural imaging but are time-consuming, expensive to acquire and they often require long breath holds that are not suitable for patients. Consequently, multiplanar breath-hold 2D cine sequences are standard practice but are disadvantaged by lack of whole-heart coverage and low through-plane resolution. To address this, we propose a conditional variational autoencoder architecture able to learn a generative model of 3D high-resolution left ventricular (LV) segmentations which is conditioned on three 2D LV segmentations of one short-axis and two long-axis images. By only employing these three 2D segmentations, our model can efficiently reconstruct the 3D high-resolution LV segmentation of a subject. When evaluated on 400 unseen healthy volunteers, our model yielded an average Dice score of $87.92 \pm 0.15$ and outperformed competing architectures.
Deep learning cardiac motion analysis for human survival prediction
Oct 08, 2018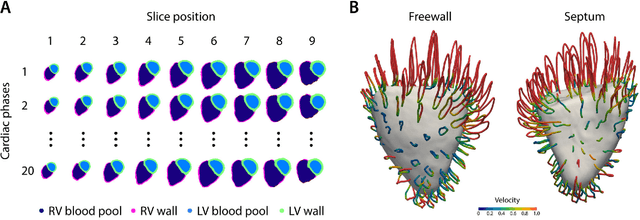
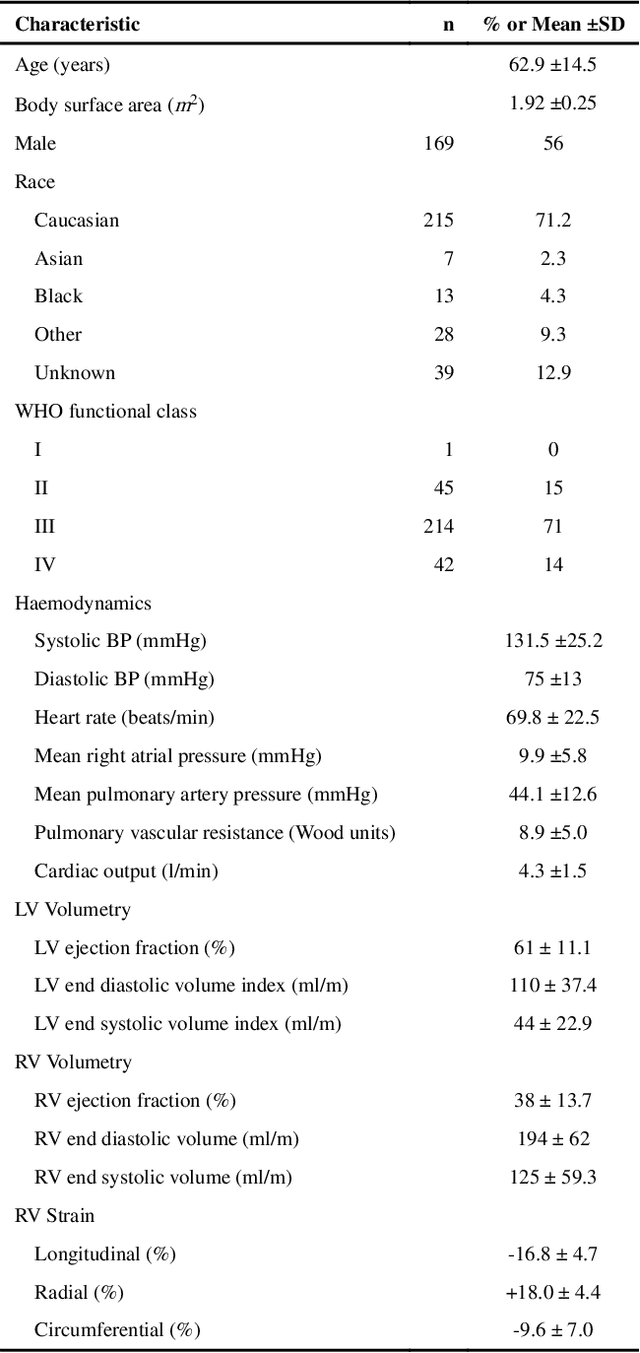


Abstract:Motion analysis is used in computer vision to understand the behaviour of moving objects in sequences of images. Optimising the interpretation of dynamic biological systems requires accurate and precise motion tracking as well as efficient representations of high-dimensional motion trajectories so that these can be used for prediction tasks. Here we use image sequences of the heart, acquired using cardiac magnetic resonance imaging, to create time-resolved three-dimensional segmentations using a fully convolutional network trained on anatomical shape priors. This dense motion model formed the input to a supervised denoising autoencoder (4Dsurvival), which is a hybrid network consisting of an autoencoder that learns a task-specific latent code representation trained on observed outcome data, yielding a latent representation optimised for survival prediction. To handle right-censored survival outcomes, our network used a Cox partial likelihood loss function. In a study of 302 patients the predictive accuracy (quantified by Harrell's C-index) was significantly higher (p < .0001) for our model C=0.73 (95$\%$ CI: 0.68 - 0.78) than the human benchmark of C=0.59 (95$\%$ CI: 0.53 - 0.65). This work demonstrates how a complex computer vision task using high-dimensional medical image data can efficiently predict human survival.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge