Rita Kuznetsova
Data-Driven Discovery of Feature Groups in Clinical Time Series
Nov 11, 2025Abstract:Clinical time series data are critical for patient monitoring and predictive modeling. These time series are typically multivariate and often comprise hundreds of heterogeneous features from different data sources. The grouping of features based on similarity and relevance to the prediction task has been shown to enhance the performance of deep learning architectures. However, defining these groups a priori using only semantic knowledge is challenging, even for domain experts. To address this, we propose a novel method that learns feature groups by clustering weights of feature-wise embedding layers. This approach seamlessly integrates into standard supervised training and discovers the groups that directly improve downstream performance on clinically relevant tasks. We demonstrate that our method outperforms static clustering approaches on synthetic data and achieves performance comparable to expert-defined groups on real-world medical data. Moreover, the learned feature groups are clinically interpretable, enabling data-driven discovery of task-relevant relationships between variables.
Towards Foundation Models for Critical Care Time Series
Nov 25, 2024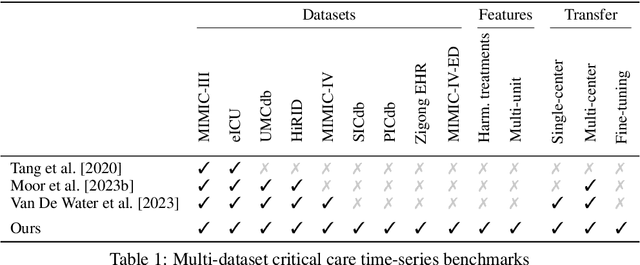
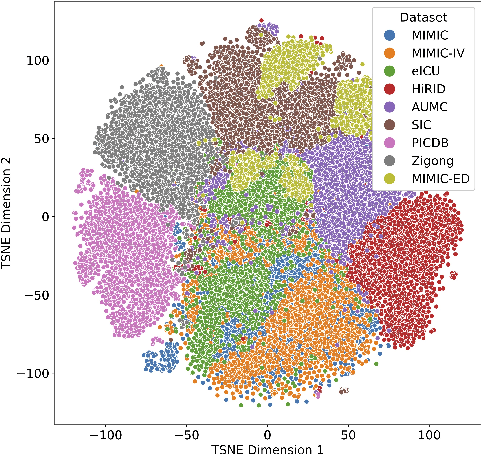
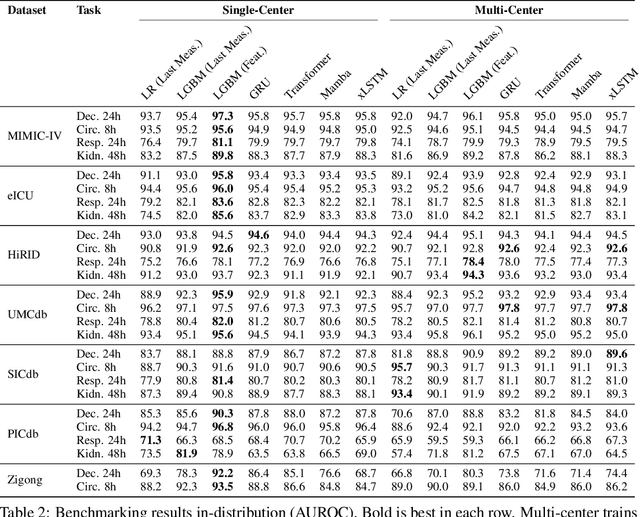
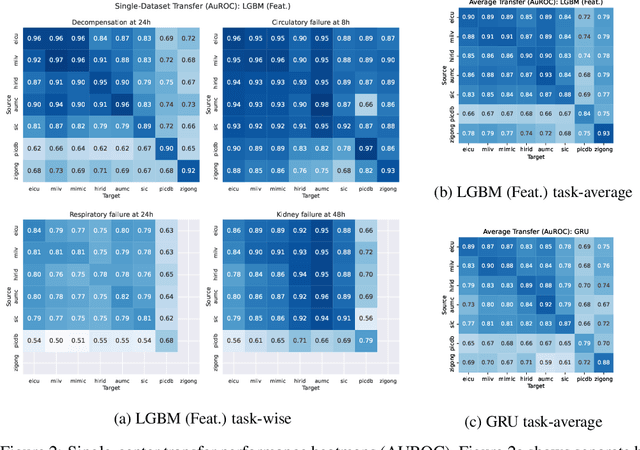
Abstract:Notable progress has been made in generalist medical large language models across various healthcare areas. However, large-scale modeling of in-hospital time series data - such as vital signs, lab results, and treatments in critical care - remains underexplored. Existing datasets are relatively small, but combining them can enhance patient diversity and improve model robustness. To effectively utilize these combined datasets for large-scale modeling, it is essential to address the distribution shifts caused by varying treatment policies, necessitating the harmonization of treatment variables across the different datasets. This work aims to establish a foundation for training large-scale multi-variate time series models on critical care data and to provide a benchmark for machine learning models in transfer learning across hospitals to study and address distribution shift challenges. We introduce a harmonized dataset for sequence modeling and transfer learning research, representing the first large-scale collection to include core treatment variables. Future plans involve expanding this dataset to support further advancements in transfer learning and the development of scalable, generalizable models for critical healthcare applications.
Multi-Modal Contrastive Learning for Online Clinical Time-Series Applications
Mar 27, 2024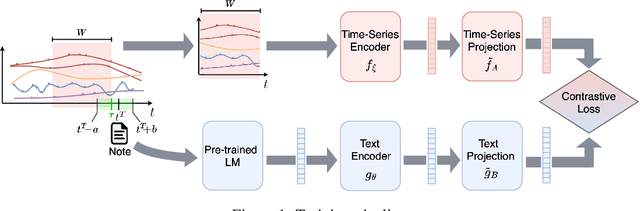
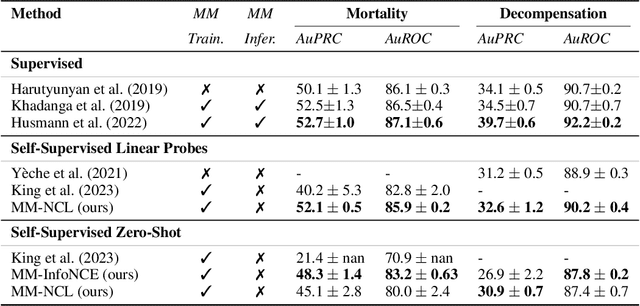

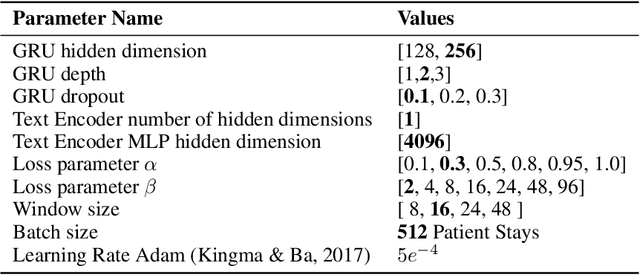
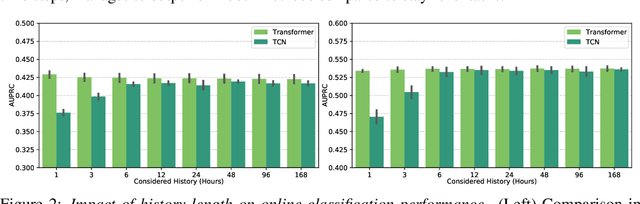
Abstract:Electronic Health Record (EHR) datasets from Intensive Care Units (ICU) contain a diverse set of data modalities. While prior works have successfully leveraged multiple modalities in supervised settings, we apply advanced self-supervised multi-modal contrastive learning techniques to ICU data, specifically focusing on clinical notes and time-series for clinically relevant online prediction tasks. We introduce a loss function Multi-Modal Neighborhood Contrastive Loss (MM-NCL), a soft neighborhood function, and showcase the excellent linear probe and zero-shot performance of our approach.
On the Importance of Step-wise Embeddings for Heterogeneous Clinical Time-Series
Nov 15, 2023



Abstract:Recent advances in deep learning architectures for sequence modeling have not fully transferred to tasks handling time-series from electronic health records. In particular, in problems related to the Intensive Care Unit (ICU), the state-of-the-art remains to tackle sequence classification in a tabular manner with tree-based methods. Recent findings in deep learning for tabular data are now surpassing these classical methods by better handling the severe heterogeneity of data input features. Given the similar level of feature heterogeneity exhibited by ICU time-series and motivated by these findings, we explore these novel methods' impact on clinical sequence modeling tasks. By jointly using such advances in deep learning for tabular data, our primary objective is to underscore the importance of step-wise embeddings in time-series modeling, which remain unexplored in machine learning methods for clinical data. On a variety of clinically relevant tasks from two large-scale ICU datasets, MIMIC-III and HiRID, our work provides an exhaustive analysis of state-of-the-art methods for tabular time-series as time-step embedding models, showing overall performance improvement. In particular, we evidence the importance of feature grouping in clinical time-series, with significant performance gains when considering features within predefined semantic groups in the step-wise embedding module.
Knowledge Graph Representations to enhance Intensive Care Time-Series Predictions
Nov 13, 2023



Abstract:Intensive Care Units (ICU) require comprehensive patient data integration for enhanced clinical outcome predictions, crucial for assessing patient conditions. Recent deep learning advances have utilized patient time series data, and fusion models have incorporated unstructured clinical reports, improving predictive performance. However, integrating established medical knowledge into these models has not yet been explored. The medical domain's data, rich in structural relationships, can be harnessed through knowledge graphs derived from clinical ontologies like the Unified Medical Language System (UMLS) for better predictions. Our proposed methodology integrates this knowledge with ICU data, improving clinical decision modeling. It combines graph representations with vital signs and clinical reports, enhancing performance, especially when data is missing. Additionally, our model includes an interpretability component to understand how knowledge graph nodes affect predictions.
Language Model Training Paradigms for Clinical Feature Embeddings
Nov 01, 2023



Abstract:In research areas with scarce data, representation learning plays a significant role. This work aims to enhance representation learning for clinical time series by deriving universal embeddings for clinical features, such as heart rate and blood pressure. We use self-supervised training paradigms for language models to learn high-quality clinical feature embeddings, achieving a finer granularity than existing time-step and patient-level representation learning. We visualize the learnt embeddings via unsupervised dimension reduction techniques and observe a high degree of consistency with prior clinical knowledge. We also evaluate the model performance on the MIMIC-III benchmark and demonstrate the effectiveness of using clinical feature embeddings. We publish our code online for replication.
Multi-modal Graph Learning over UMLS Knowledge Graphs
Jul 10, 2023



Abstract:Clinicians are increasingly looking towards machine learning to gain insights about patient evolutions. We propose a novel approach named Multi-Modal UMLS Graph Learning (MMUGL) for learning meaningful representations of medical concepts using graph neural networks over knowledge graphs based on the unified medical language system. These representations are aggregated to represent entire patient visits and then fed into a sequence model to perform predictions at the granularity of multiple hospital visits of a patient. We improve performance by incorporating prior medical knowledge and considering multiple modalities. We compare our method to existing architectures proposed to learn representations at different granularities on the MIMIC-III dataset and show that our approach outperforms these methods. The results demonstrate the significance of multi-modal medical concept representations based on prior medical knowledge.
On the Importance of Clinical Notes in Multi-modal Learning for EHR Data
Dec 06, 2022Abstract:Understanding deep learning model behavior is critical to accepting machine learning-based decision support systems in the medical community. Previous research has shown that jointly using clinical notes with electronic health record (EHR) data improved predictive performance for patient monitoring in the intensive care unit (ICU). In this work, we explore the underlying reasons for these improvements. While relying on a basic attention-based model to allow for interpretability, we first confirm that performance significantly improves over state-of-the-art EHR data models when combining EHR data and clinical notes. We then provide an analysis showing improvements arise almost exclusively from a subset of notes containing broader context on patient state rather than clinician notes. We believe such findings highlight deep learning models for EHR data to be more limited by partially-descriptive data than by modeling choice, motivating a more data-centric approach in the field.
Temporal Label Smoothing for Early Prediction of Adverse Events
Aug 29, 2022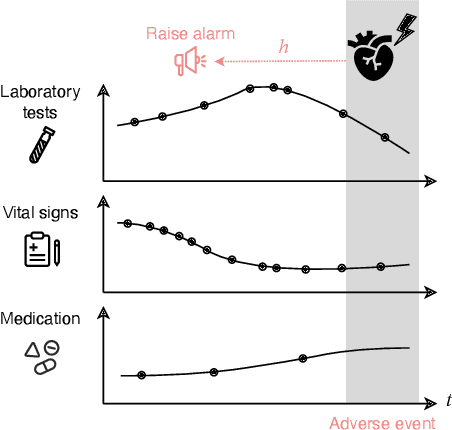

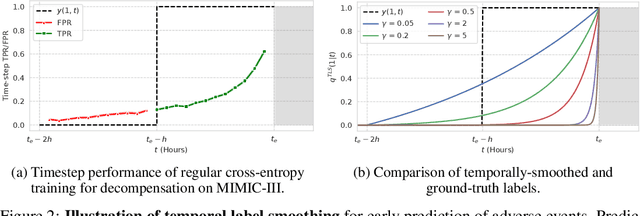
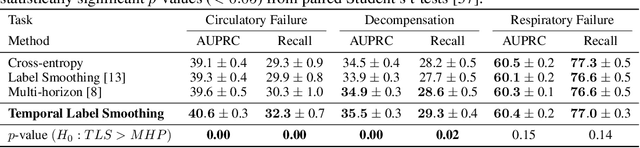
Abstract:Models that can predict adverse events ahead of time with low false-alarm rates are critical to the acceptance of decision support systems in the medical community. This challenging machine learning task remains typically treated as simple binary classification, with few bespoke methods proposed to leverage temporal dependency across samples. We propose Temporal Label Smoothing (TLS), a novel learning strategy that modulates smoothing strength as a function of proximity to the event of interest. This regularization technique reduces model confidence at the class boundary, where the signal is often noisy or uninformative, thus allowing training to focus on clinically informative data points away from this boundary region. From a theoretical perspective, we also show that our method can be framed as an extension of multi-horizon prediction, a learning heuristic proposed in other early prediction work. TLS empirically matches or outperforms considered competing methods on various early prediction benchmark tasks. In particular, our approach significantly improves performance on clinically-relevant metrics such as event recall at low false-alarm rates.
HiRID-ICU-Benchmark -- A Comprehensive Machine Learning Benchmark on High-resolution ICU Data
Nov 18, 2021
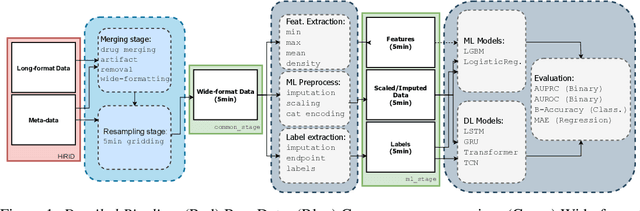
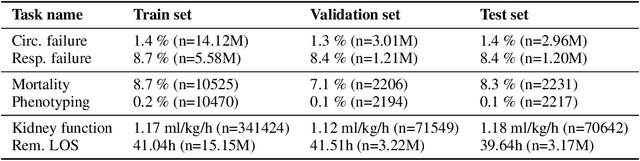
Abstract:The recent success of machine learning methods applied to time series collected from Intensive Care Units (ICU) exposes the lack of standardized machine learning benchmarks for developing and comparing such methods. While raw datasets, such as MIMIC-IV or eICU, can be freely accessed on Physionet, the choice of tasks and pre-processing is often chosen ad-hoc for each publication, limiting comparability across publications. In this work, we aim to improve this situation by providing a benchmark covering a large spectrum of ICU-related tasks. Using the HiRID dataset, we define multiple clinically relevant tasks developed in collaboration with clinicians. In addition, we provide a reproducible end-to-end pipeline to construct both data and labels. Finally, we provide an in-depth analysis of current state-of-the-art sequence modeling methods, highlighting some limitations of deep learning approaches for this type of data. With this benchmark, we hope to give the research community the possibility of a fair comparison of their work.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge