Rima Arnaout
MARCUS: An agentic, multimodal vision-language model for cardiac diagnosis and management
Mar 23, 2026Abstract:Cardiovascular disease remains the leading cause of global mortality, with progress hindered by human interpretation of complex cardiac tests. Current AI vision-language models are limited to single-modality inputs and are non-interactive. We present MARCUS (Multimodal Autonomous Reasoning and Chat for Ultrasound and Signals), an agentic vision-language system for end-to-end interpretation of electrocardiograms (ECGs), echocardiograms, and cardiac magnetic resonance imaging (CMR) independently and as multimodal input. MARCUS employs a hierarchical agentic architecture comprising modality-specific vision-language expert models, each integrating domain-trained visual encoders with multi-stage language model optimization, coordinated by a multimodal orchestrator. Trained on 13.5 million images (0.25M ECGs, 1.3M echocardiogram images, 12M CMR images) and our novel expert-curated dataset spanning 1.6 million questions, MARCUS achieves state-of-the-art performance surpassing frontier models (GPT-5 Thinking, Gemini 2.5 Pro Deep Think). Across internal (Stanford) and external (UCSF) test cohorts, MARCUS achieves accuracies of 87-91% for ECG, 67-86% for echocardiography, and 85-88% for CMR, outperforming frontier models by 34-45% (P<0.001). On multimodal cases, MARCUS achieved 70% accuracy, nearly triple that of frontier models (22-28%), with 1.7-3.0x higher free-text quality scores. Our agentic architecture also confers resistance to mirage reasoning, whereby vision-language models derive reasoning from unintended textual signals or hallucinated visual content. MARCUS demonstrates that domain-specific visual encoders with an agentic orchestrator enable multimodal cardiac interpretation. We release our models, code, and benchmark open-source.
Which Similarity-Sensitive Entropy?
Nov 10, 2025Abstract:A canonical step in quantifying a system is to measure its entropy. Shannon entropy and other traditional entropy measures capture only the information encoded in the frequencies of a system's elements. Recently, Leinster, Cobbold, and Reeve (LCR) introduced a method that also captures the rich information encoded in the similarities and differences among elements, yielding similarity-sensitive entropy. More recently, the Vendi score (VS) was introduced as an alternative, raising the question of how LCR and VS compare, and which is preferable. Here we address these questions conceptually, analytically, and experimentally, using 53 machine-learning datasets. We show that LCR and VS can differ by orders of magnitude and can capture complementary information about a system, except in limiting cases. We demonstrate that both LCR and VS depend on how similarities are scaled and introduce the concept of ``half distance'' to parameterize this dependence. We prove that VS provides an upper bound on LCR for several values of the Rényi-Hill order parameter and conjecture that this bound holds for all values. We conclude that VS is preferable only when interpreting elements as linear combinations of a more fundamental set of ``ur-elements'' or when the system or dataset possesses a quantum-mechanical character. In the broader circumstance where one seeks simply to capture the rich information encoded by similarity, LCR is favored; nevertheless, for certain half-distances the two methods can complement each other.
X-Factor: Quality Is a Dataset-Intrinsic Property
May 28, 2025Abstract:In the universal quest to optimize machine-learning classifiers, three factors -- model architecture, dataset size, and class balance -- have been shown to influence test-time performance but do not fully account for it. Previously, evidence was presented for an additional factor that can be referred to as dataset quality, but it was unclear whether this was actually a joint property of the dataset and the model architecture, or an intrinsic property of the dataset itself. If quality is truly dataset-intrinsic and independent of model architecture, dataset size, and class balance, then the same datasets should perform better (or worse) regardless of these other factors. To test this hypothesis, here we create thousands of datasets, each controlled for size and class balance, and use them to train classifiers with a wide range of architectures, from random forests and support-vector machines to deep networks. We find that classifier performance correlates strongly by subset across architectures ($R^2=0.79$), supporting quality as an intrinsic property of datasets independent of dataset size and class balance and of model architecture. Digging deeper, we find that dataset quality appears to be an emergent property of something more fundamental: the quality of datasets' constituent classes. Thus, quality joins size, class balance, and model architecture as an independent correlate of performance and a separate target for optimizing machine-learning-based classification.
Grade Inflation in Generative Models
Jan 05, 2025Abstract:Generative models hold great potential, but only if one can trust the evaluation of the data they generate. We show that many commonly used quality scores for comparing two-dimensional distributions of synthetic vs. ground-truth data give better results than they should, a phenomenon we call the "grade inflation problem." We show that the correlation score, Jaccard score, earth-mover's score, and Kullback-Leibler (relative-entropy) score all suffer grade inflation. We propose that any score that values all datapoints equally, as these do, will also exhibit grade inflation; we refer to such scores as "equipoint" scores. We introduce the concept of "equidensity" scores, and present the Eden score, to our knowledge the first example of such a score. We found that Eden avoids grade inflation and agrees better with human perception of goodness-of-fit than the equipoint scores above. We propose that any reasonable equidensity score will avoid grade inflation. We identify a connection between equidensity scores and R\'enyi entropy of negative order. We conclude that equidensity scores are likely to outperform equipoint scores for generative models, and for comparing low-dimensional distributions more generally.
Beyond Size and Class Balance: Alpha as a New Dataset Quality Metric for Deep Learning
Jul 22, 2024Abstract:In deep learning, achieving high performance on image classification tasks requires diverse training sets. However, dataset diversity is incompletely understood. The current best practice is to try to maximize dataset size and class balance. Yet large, class-balanced datasets are not guaranteed to be diverse: images can still be arbitrarily similar. We hypothesized that, for a given model architecture, better model performance can be achieved by maximizing dataset diversity more directly. This could open a path for performance improvement without additional computational resources or architectural advances. To test this hypothesis, we introduce a comprehensive framework of diversity measures, developed in ecology, that generalizes familiar quantities like Shannon entropy by accounting for similarities and differences among images. (Dataset size and class balance emerge from this framework as special cases.) By analyzing thousands of subsets from seven medical datasets representing ultrasound, X-ray, CT, and pathology images, we found that the best correlates of performance were not size or class balance but $A$ -- ``big alpha'' -- a set of generalized entropy measures interpreted as the effective number of image-class pairs in the dataset, after accounting for similarities among images. One of these, $A_0$, explained 67\% of the variance in balanced accuracy across all subsets, vs. 54\% for class balance and just 39\% for size. The best pair was size and $A_1$ (79\%), which outperformed size and class balance (74\%). $A$ performed best for subsets from individual datasets as well as across datasets, supporting the generality of these results. We propose maximizing $A$ as a potential new way to improve the performance of deep learning in medical imaging.
Are foundation models efficient for medical image segmentation?
Nov 08, 2023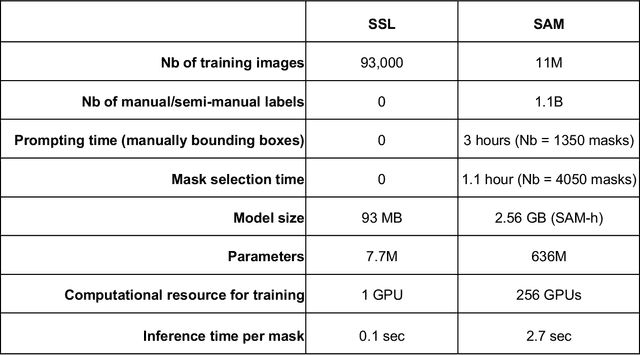

Abstract:Foundation models are experiencing a surge in popularity. The Segment Anything model (SAM) asserts an ability to segment a wide spectrum of objects but required supervised training at unprecedented scale. We compared SAM's performance (against clinical ground truth) and resources (labeling time, compute) to a modality-specific, label-free self-supervised learning (SSL) method on 25 measurements for 100 cardiac ultrasounds. SAM performed poorly and required significantly more labeling and computing resources, demonstrating worse efficiency than SSL.
Label-free segmentation from cardiac ultrasound using self-supervised learning
Oct 10, 2022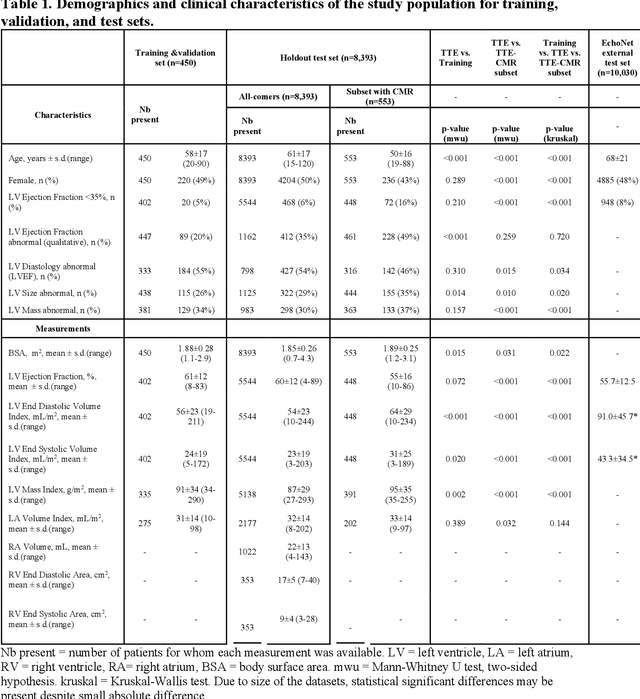
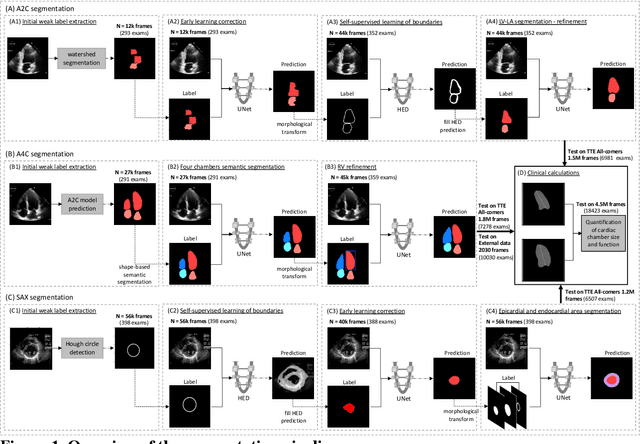
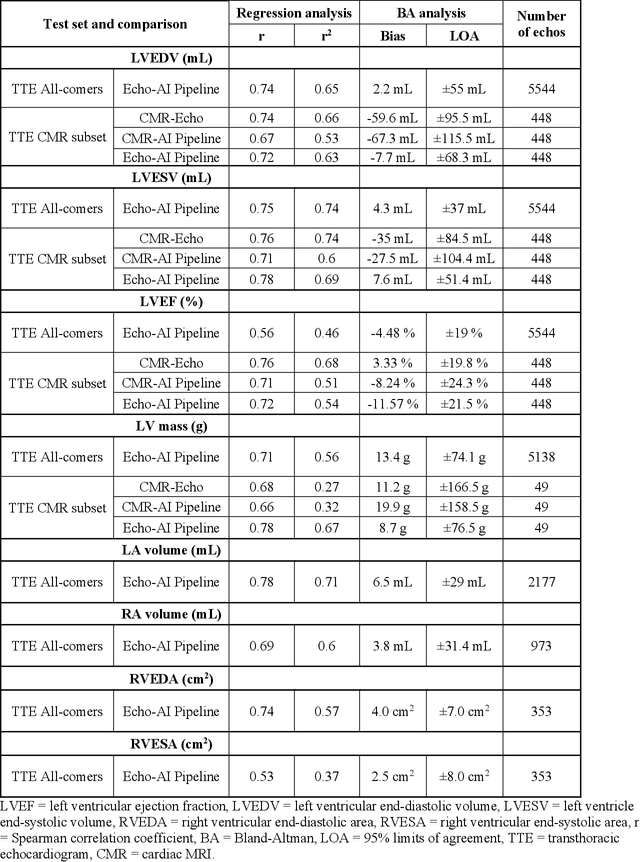
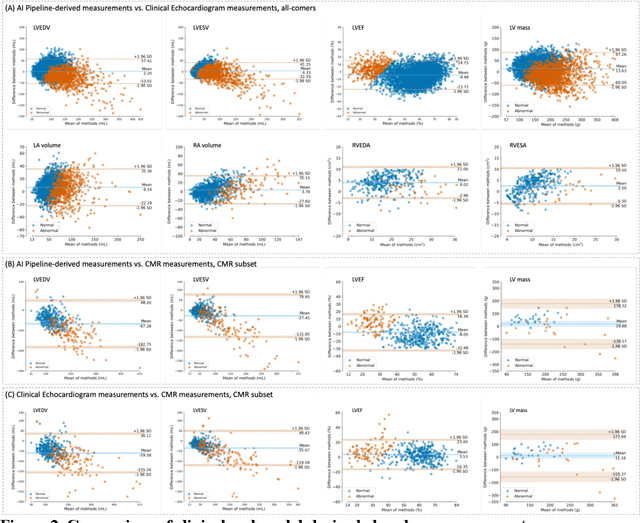
Abstract:Background: Segmentation and measurement of cardiac chambers is critical in echocardiography but is also laborious and poorly reproducible. Neural networks can assist, but supervised approaches require the same laborious manual annotations, while unsupervised approaches have fared poorly in ultrasound to date. Objectives: We built a pipeline for self-supervised (no manual labels required) segmentation of cardiac chambers, combining computer vision, clinical domain knowledge, and deep learning. Methods: We trained on 450 echocardiograms (145,000 images) and tested on 8,393 echocardiograms (4,476,266 images; mean age 61 years, 51% female), using the resulting segmentations to calculate structural and functional measurements. We also tested our pipeline against external images from an additional 10,030 patients (20,060 images) with available manual tracings of the left ventricle. Results: r2 between clinically measured and pipeline-predicted measurements were similar to reported inter-clinician variation for LVESV and LVEDV (pipeline vs. clinical r2= 0.74 and r2=0.65, respectively), LVEF and LV mass (r2= 0.46 and r2=0.54), left and right atrium volumes (r2=0.7 and r2=0.6), and right ventricle area (r2=0.47). When binarized into normal vs. abnormal categories, average accuracy was 0.81 (range 0.71-0.95). A subset of the test echocardiograms (n=553) had corresponding cardiac MRI; correlation between pipeline and CMR measurements was similar to that between clinical echocardiogram and CMR. Finally, in the external dataset, our pipeline accurately segments the left ventricle with an average Dice score of 0.83 (95% CI 0.83). Conclusions: Our results demonstrate a human-label-free, valid, and scalable method for segmentation from ultrasound, a noisy but globally important imaging modality.
Domain-guided data augmentation for deep learning on medical imaging
Oct 10, 2022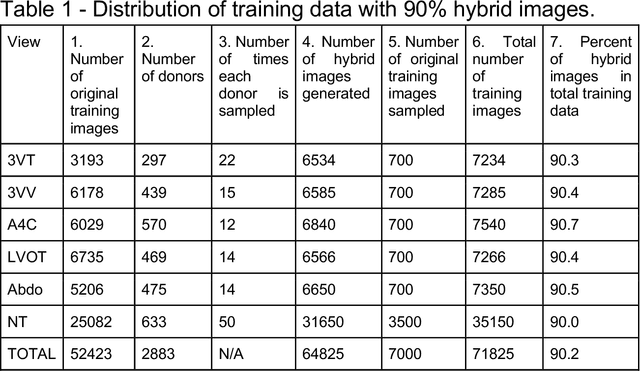
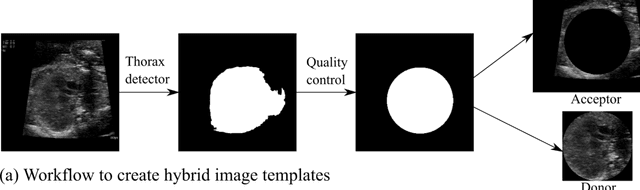

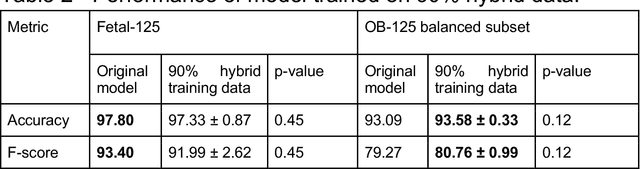
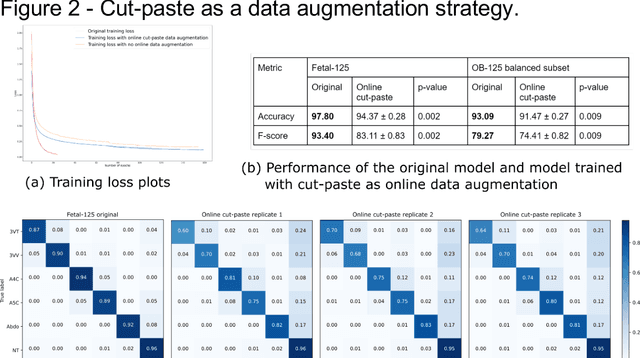
Abstract:While domain-specific data augmentation can be useful in training neural networks for medical imaging tasks, such techniques have not been widely used to date. Here, we test whether domain-specific data augmentation is useful for medical imaging using a well-benchmarked task: view classification on fetal ultrasound FETAL-125 and OB-125 datasets. We found that using a context-preserving cut-paste strategy, we could create valid training data as measured by performance of the resulting trained model on the benchmark test dataset. When used in an online fashion, models trained on this data performed similarly to those trained using traditional data augmentation (FETAL-125 F-score 85.33+/-0.24 vs 86.89+/-0.60, p-value 0.0139; OB-125 F-score 74.60+/-0.11 vs 72.43+/-0.62, p-value 0.0039). Furthermore, the ability to perform augmentations during training time, as well as the ability to apply chosen augmentations equally across data classes, are important considerations in designing a bespoke data augmentation. Finally, we provide open-source code to facilitate running bespoke data augmentations in an online fashion. Taken together, this work expands the ability to design and apply domain-guided data augmentations for medical imaging tasks.
Deep-learning models improve on community-level diagnosis for common congenital heart disease lesions
Sep 19, 2018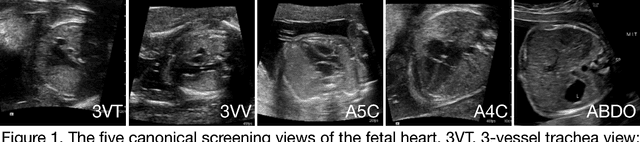
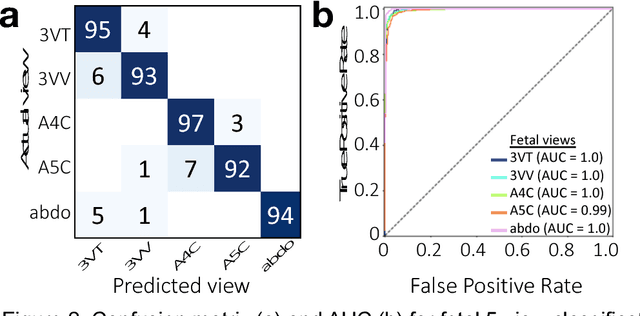

Abstract:Prenatal diagnosis of tetralogy of Fallot (TOF) and hypoplastic left heart syndrome (HLHS), two serious congenital heart defects, improves outcomes and can in some cases facilitate in utero interventions. In practice, however, the fetal diagnosis rate for these lesions is only 30-50 percent in community settings. Improving fetal diagnosis of congenital heart disease is therefore critical. Deep learning is a cutting-edge machine learning technique for finding patterns in images but has not yet been applied to prenatal diagnosis of congenital heart disease. Using 685 retrospectively collected echocardiograms from fetuses 18-24 weeks of gestational age from 2000-2018, we trained convolutional and fully-convolutional deep learning models in a supervised manner to (i) identify the five canonical screening views of the fetal heart and (ii) segment cardiac structures to calculate fetal cardiac biometrics. We then trained models to distinguish by view between normal hearts, TOF, and HLHS. In a holdout test set of images, F-score for identification of the five most important fetal cardiac views was 0.95. Binary classification of unannotated cardiac views of normal heart vs. TOF reached an overall sensitivity of 75% and a specificity of 76%, while normal vs. HLHS reached a sensitivity of 100% and specificity of 90%, both well above average diagnostic rates for these lesions. Furthermore, segmentation-based measurements for cardiothoracic ratio (CTR), cardiac axis (CA), and ventricular fractional area change (FAC) were compatible with clinically measured metrics for normal, TOF, and HLHS hearts. Thus, using guideline-recommended imaging, deep learning models can significantly improve detection of fetal congenital heart disease compared to the common standard of care.
Fast and accurate classification of echocardiograms using deep learning
Jun 27, 2017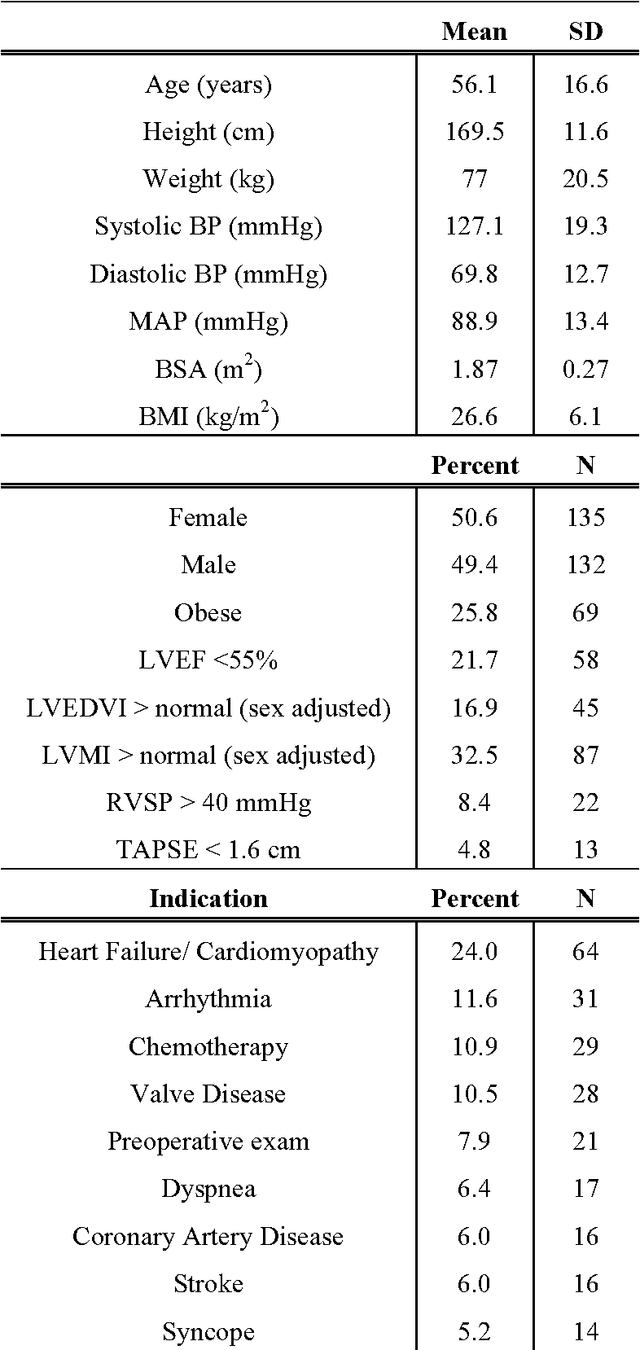
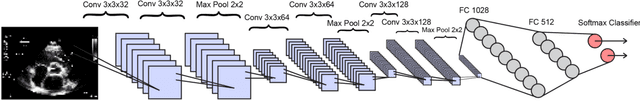
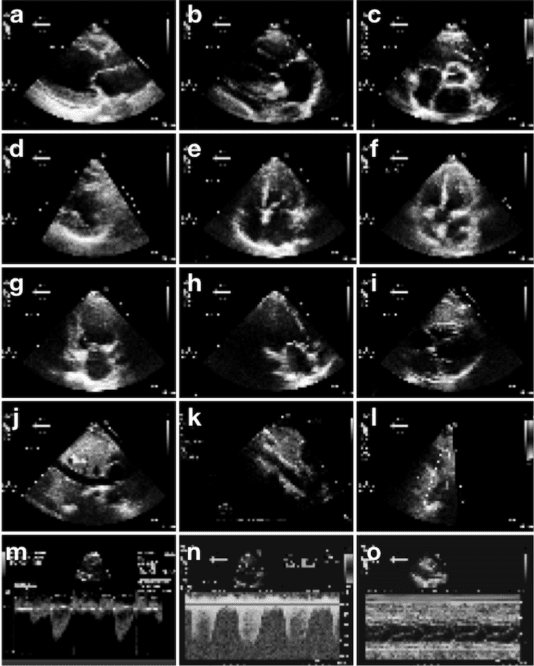
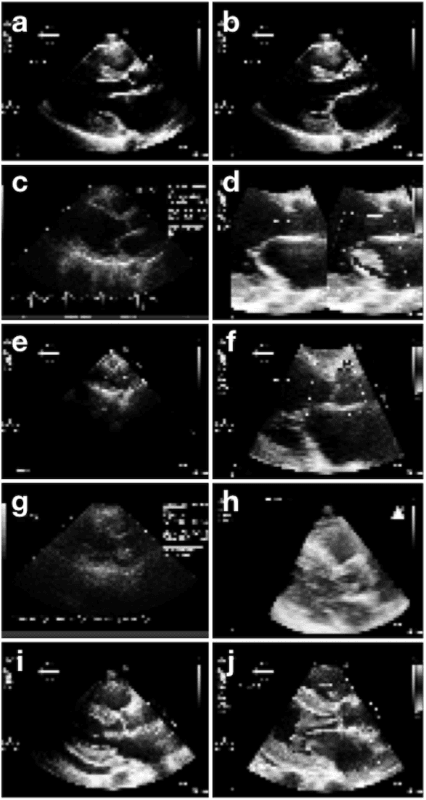
Abstract:Echocardiography is essential to modern cardiology. However, human interpretation limits high throughput analysis, limiting echocardiography from reaching its full clinical and research potential for precision medicine. Deep learning is a cutting-edge machine-learning technique that has been useful in analyzing medical images but has not yet been widely applied to echocardiography, partly due to the complexity of echocardiograms' multi view, multi modality format. The essential first step toward comprehensive computer assisted echocardiographic interpretation is determining whether computers can learn to recognize standard views. To this end, we anonymized 834,267 transthoracic echocardiogram (TTE) images from 267 patients (20 to 96 years, 51 percent female, 26 percent obese) seen between 2000 and 2017 and labeled them according to standard views. Images covered a range of real world clinical variation. We built a multilayer convolutional neural network and used supervised learning to simultaneously classify 15 standard views. Eighty percent of data used was randomly chosen for training and 20 percent reserved for validation and testing on never seen echocardiograms. Using multiple images from each clip, the model classified among 12 video views with 97.8 percent overall test accuracy without overfitting. Even on single low resolution images, test accuracy among 15 views was 91.7 percent versus 70.2 to 83.5 percent for board-certified echocardiographers. Confusional matrices, occlusion experiments, and saliency mapping showed that the model finds recognizable similarities among related views and classifies using clinically relevant image features. In conclusion, deep neural networks can classify essential echocardiographic views simultaneously and with high accuracy. Our results provide a foundation for more complex deep learning assisted echocardiographic interpretation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge