Mohammad Mofrad
ProDyn0: Inferring calponin homology domain stretching behavior using graph neural networks
Oct 22, 2019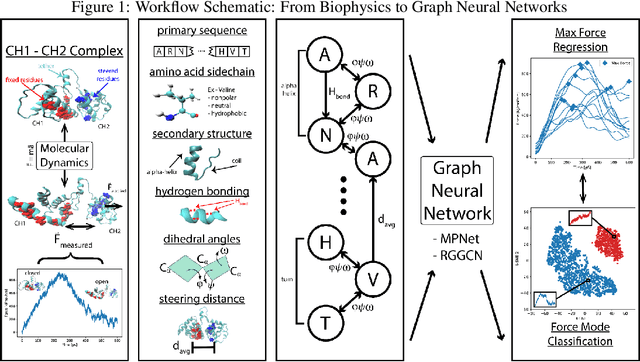
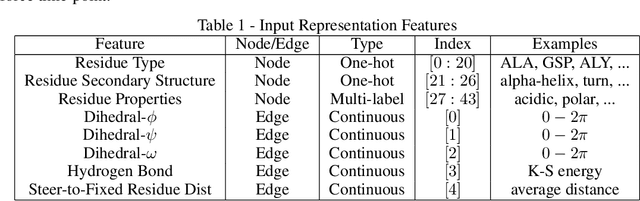
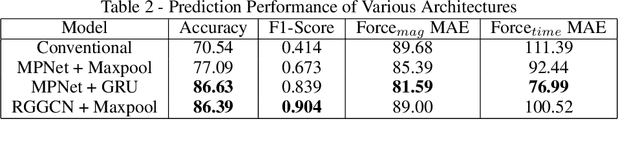
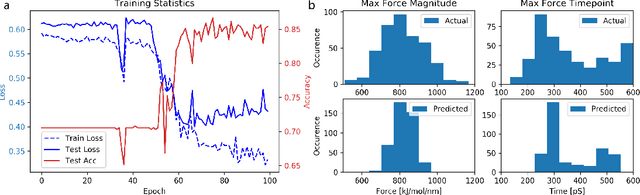
Abstract:Graph neural networks are a quickly emerging field for non-Euclidean data that leverage the inherent graphical structure to predict node, edge, and global-level properties of a system. Protein properties can not easily be understood as a simple sum of their parts (i.e. amino acids), therefore, understanding their dynamical properties in the context of graphs is attractive for revealing how perturbations to their structure can affect their global function. To tackle this problem, we generate a database of 2020 mutated calponin homology (CH) domains undergoing large-scale separation in molecular dynamics. To predict the mechanosensitive force response, we develop neural message passing networks and residual gated graph convnets which predict the protein dependent force separation at 86.63 percent, 81.59 kJ/mol/nm MAE, 76.99 psec MAE for force mode classification, max force magnitude, max force time respectively-- significantly better than non-graph-based deep learning techniques. Towards uniting geometric learning techniques and biophysical observables, we premiere our simulation database as a benchmark dataset for further development/evaluation of graph neural network architectures.
* 8 pages, 2 figures, 2 tables
Fast and accurate classification of echocardiograms using deep learning
Jun 27, 2017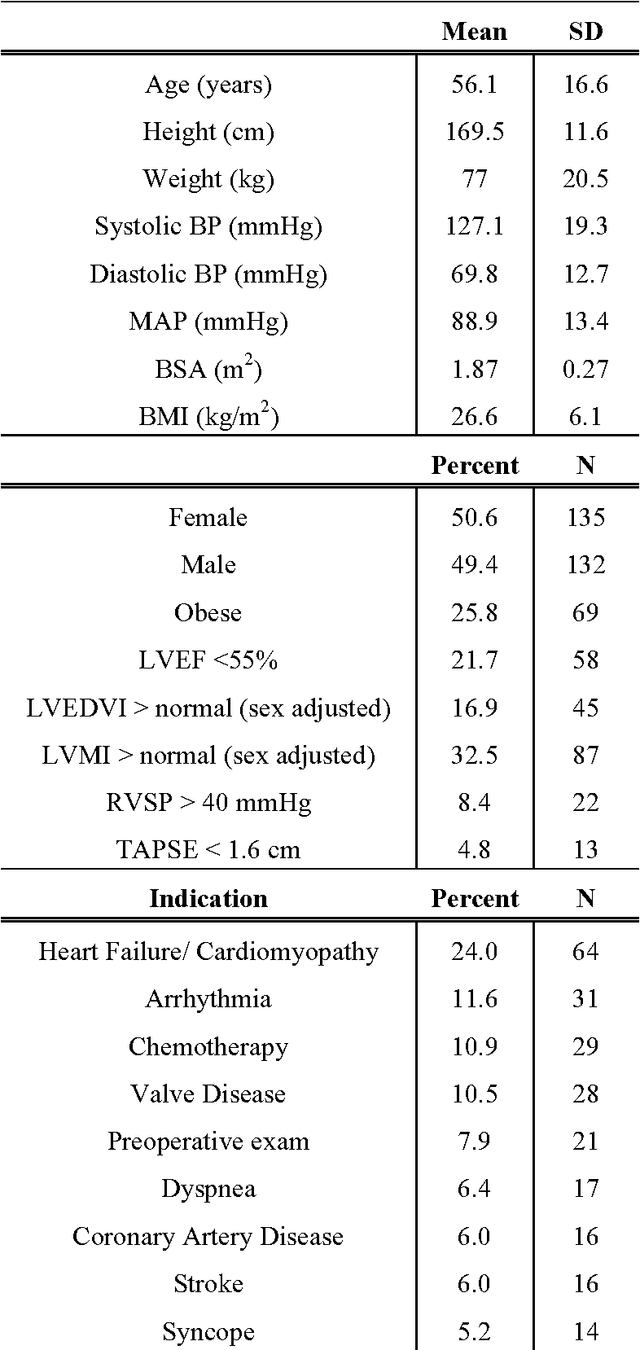
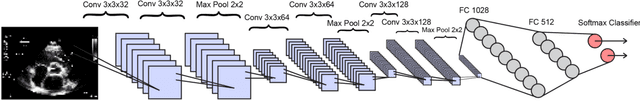
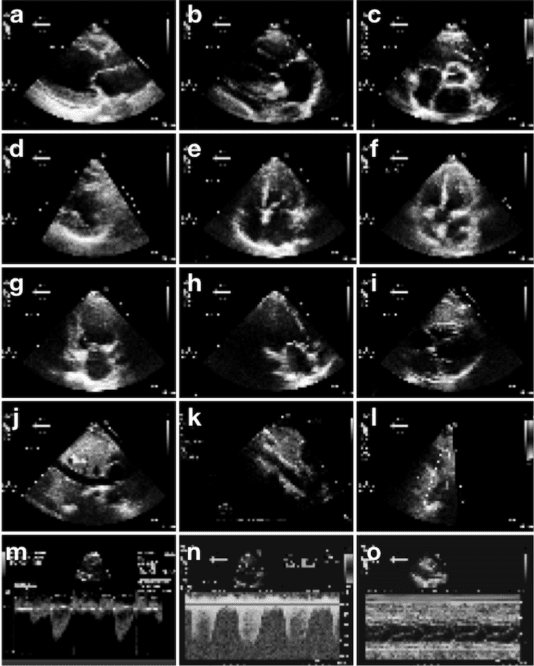
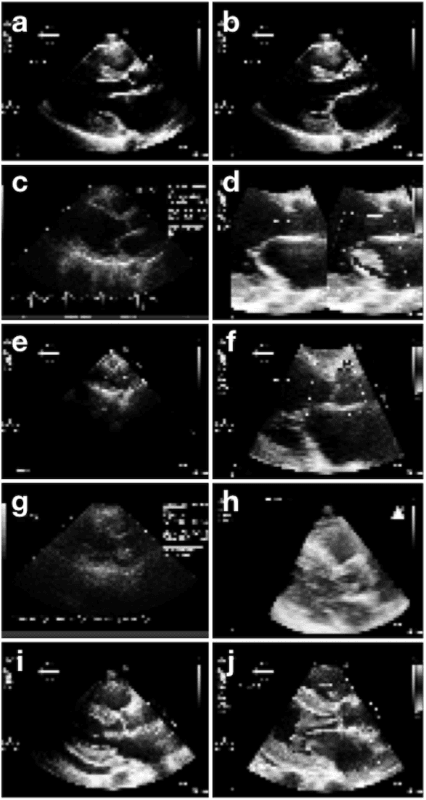
Abstract:Echocardiography is essential to modern cardiology. However, human interpretation limits high throughput analysis, limiting echocardiography from reaching its full clinical and research potential for precision medicine. Deep learning is a cutting-edge machine-learning technique that has been useful in analyzing medical images but has not yet been widely applied to echocardiography, partly due to the complexity of echocardiograms' multi view, multi modality format. The essential first step toward comprehensive computer assisted echocardiographic interpretation is determining whether computers can learn to recognize standard views. To this end, we anonymized 834,267 transthoracic echocardiogram (TTE) images from 267 patients (20 to 96 years, 51 percent female, 26 percent obese) seen between 2000 and 2017 and labeled them according to standard views. Images covered a range of real world clinical variation. We built a multilayer convolutional neural network and used supervised learning to simultaneously classify 15 standard views. Eighty percent of data used was randomly chosen for training and 20 percent reserved for validation and testing on never seen echocardiograms. Using multiple images from each clip, the model classified among 12 video views with 97.8 percent overall test accuracy without overfitting. Even on single low resolution images, test accuracy among 15 views was 91.7 percent versus 70.2 to 83.5 percent for board-certified echocardiographers. Confusional matrices, occlusion experiments, and saliency mapping showed that the model finds recognizable similarities among related views and classifies using clinically relevant image features. In conclusion, deep neural networks can classify essential echocardiographic views simultaneously and with high accuracy. Our results provide a foundation for more complex deep learning assisted echocardiographic interpretation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge