Danielle L. Ferreira
SAS: Segment Anything Small for Ultrasound -- A Non-Generative Data Augmentation Technique for Robust Deep Learning in Ultrasound Imaging
Mar 07, 2025
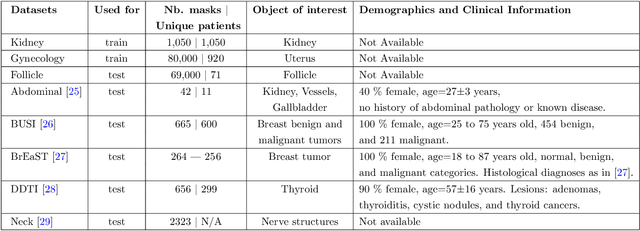
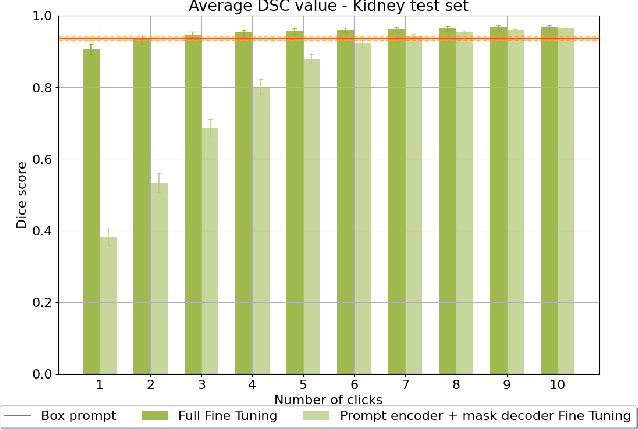
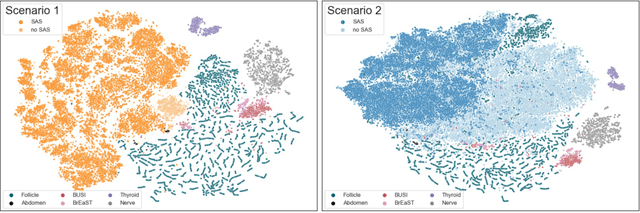
Abstract:Accurate segmentation of anatomical structures in ultrasound (US) images, particularly small ones, is challenging due to noise and variability in imaging conditions (e.g., probe position, patient anatomy, tissue characteristics and pathology). To address this, we introduce Segment Anything Small (SAS), a simple yet effective scale- and texture-aware data augmentation technique designed to enhance the performance of deep learning models for segmenting small anatomical structures in ultrasound images. SAS employs a dual transformation strategy: (1) simulating diverse organ scales by resizing and embedding organ thumbnails into a black background, and (2) injecting noise into regions of interest to simulate varying tissue textures. These transformations generate realistic and diverse training data without introducing hallucinations or artifacts, improving the model's robustness to noise and variability. We fine-tuned a promptable foundation model on a controlled organ-specific medical imaging dataset and evaluated its performance on one internal and five external datasets. Experimental results demonstrate significant improvements in segmentation performance, with Dice score gains of up to 0.35 and an average improvement of 0.16 [95% CI 0.132,0.188]. Additionally, our iterative point prompts provide precise control and adaptive refinement, achieving performance comparable to bounding box prompts with just two points. SAS enhances model robustness and generalizability across diverse anatomical structures and imaging conditions, particularly for small structures, without compromising the accuracy of larger ones. By offering a computationally efficient solution that eliminates the need for extensive human labeling efforts, SAS emerges as a powerful tool for advancing medical image analysis, particularly in resource-constrained settings.
Label-free segmentation from cardiac ultrasound using self-supervised learning
Oct 10, 2022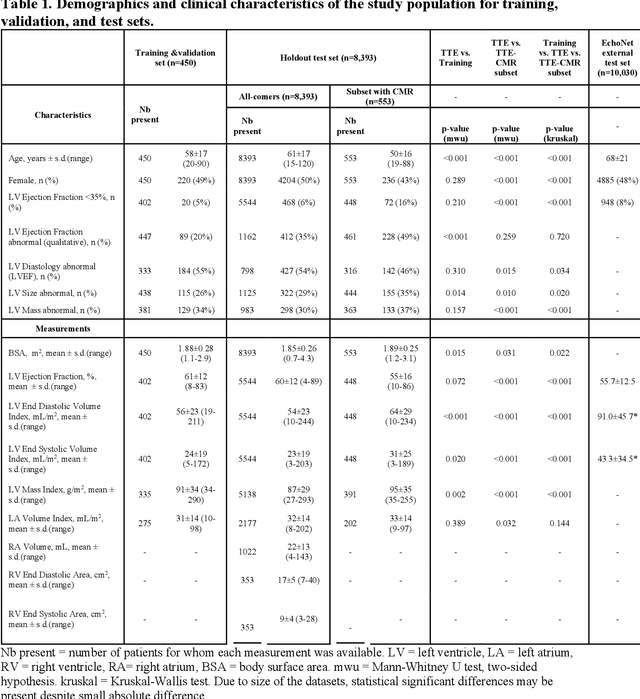
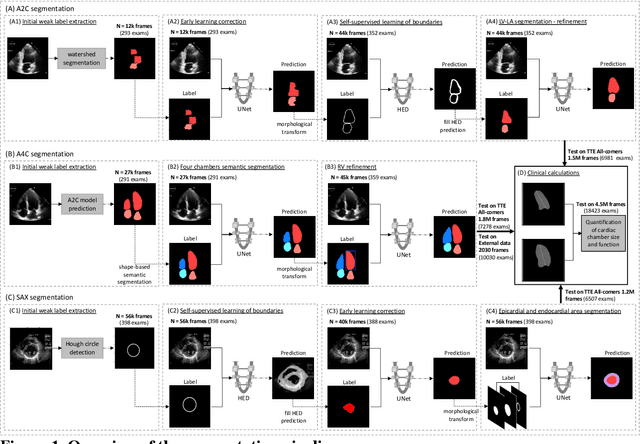
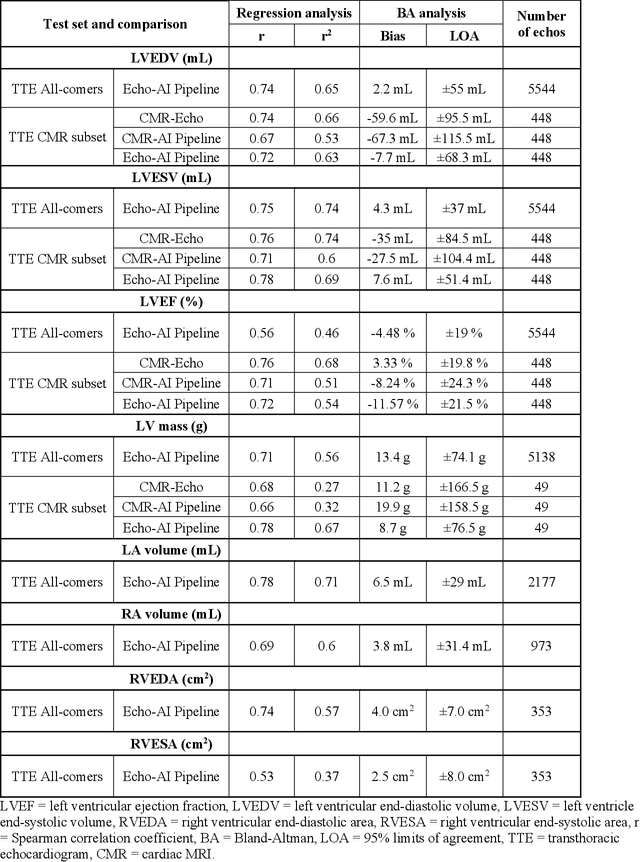
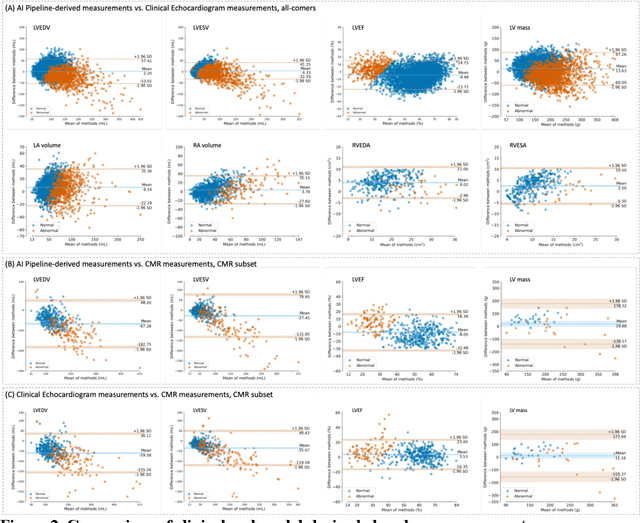
Abstract:Background: Segmentation and measurement of cardiac chambers is critical in echocardiography but is also laborious and poorly reproducible. Neural networks can assist, but supervised approaches require the same laborious manual annotations, while unsupervised approaches have fared poorly in ultrasound to date. Objectives: We built a pipeline for self-supervised (no manual labels required) segmentation of cardiac chambers, combining computer vision, clinical domain knowledge, and deep learning. Methods: We trained on 450 echocardiograms (145,000 images) and tested on 8,393 echocardiograms (4,476,266 images; mean age 61 years, 51% female), using the resulting segmentations to calculate structural and functional measurements. We also tested our pipeline against external images from an additional 10,030 patients (20,060 images) with available manual tracings of the left ventricle. Results: r2 between clinically measured and pipeline-predicted measurements were similar to reported inter-clinician variation for LVESV and LVEDV (pipeline vs. clinical r2= 0.74 and r2=0.65, respectively), LVEF and LV mass (r2= 0.46 and r2=0.54), left and right atrium volumes (r2=0.7 and r2=0.6), and right ventricle area (r2=0.47). When binarized into normal vs. abnormal categories, average accuracy was 0.81 (range 0.71-0.95). A subset of the test echocardiograms (n=553) had corresponding cardiac MRI; correlation between pipeline and CMR measurements was similar to that between clinical echocardiogram and CMR. Finally, in the external dataset, our pipeline accurately segments the left ventricle with an average Dice score of 0.83 (95% CI 0.83). Conclusions: Our results demonstrate a human-label-free, valid, and scalable method for segmentation from ultrasound, a noisy but globally important imaging modality.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge