Rajarsi R. Gupta
PixCell: A generative foundation model for digital histopathology images
Jun 05, 2025Abstract:The digitization of histology slides has revolutionized pathology, providing massive datasets for cancer diagnosis and research. Contrastive self-supervised and vision-language models have been shown to effectively mine large pathology datasets to learn discriminative representations. On the other hand, generative models, capable of synthesizing realistic and diverse images, present a compelling solution to address unique problems in pathology that involve synthesizing images; overcoming annotated data scarcity, enabling privacy-preserving data sharing, and performing inherently generative tasks, such as virtual staining. We introduce PixCell, the first diffusion-based generative foundation model for histopathology. We train PixCell on PanCan-30M, a vast, diverse dataset derived from 69,184 H\&E-stained whole slide images covering various cancer types. We employ a progressive training strategy and a self-supervision-based conditioning that allows us to scale up training without any annotated data. PixCell generates diverse and high-quality images across multiple cancer types, which we find can be used in place of real data to train a self-supervised discriminative model. Synthetic images shared between institutions are subject to fewer regulatory barriers than would be the case with real clinical images. Furthermore, we showcase the ability to precisely control image generation using a small set of annotated images, which can be used for both data augmentation and educational purposes. Testing on a cell segmentation task, a mask-guided PixCell enables targeted data augmentation, improving downstream performance. Finally, we demonstrate PixCell's ability to use H\&E structural staining to infer results from molecular marker studies; we use this capability to infer IHC staining from H\&E images. Our trained models are publicly released to accelerate research in computational pathology.
GECKO: Gigapixel Vision-Concept Contrastive Pretraining in Histopathology
Apr 01, 2025Abstract:Pretraining a Multiple Instance Learning (MIL) aggregator enables the derivation of Whole Slide Image (WSI)-level embeddings from patch-level representations without supervision. While recent multimodal MIL pretraining approaches leveraging auxiliary modalities have demonstrated performance gains over unimodal WSI pretraining, the acquisition of these additional modalities necessitates extensive clinical profiling. This requirement increases costs and limits scalability in existing WSI datasets lacking such paired modalities. To address this, we propose Gigapixel Vision-Concept Knowledge Contrastive pretraining (GECKO), which aligns WSIs with a Concept Prior derived from the available WSIs. First, we derive an inherently interpretable concept prior by computing the similarity between each WSI patch and textual descriptions of predefined pathology concepts. GECKO then employs a dual-branch MIL network: one branch aggregates patch embeddings into a WSI-level deep embedding, while the other aggregates the concept prior into a corresponding WSI-level concept embedding. Both aggregated embeddings are aligned using a contrastive objective, thereby pretraining the entire dual-branch MIL model. Moreover, when auxiliary modalities such as transcriptomics data are available, GECKO seamlessly integrates them. Across five diverse tasks, GECKO consistently outperforms prior unimodal and multimodal pretraining approaches while also delivering clinically meaningful interpretability that bridges the gap between computational models and pathology expertise. Code is made available at https://github.com/bmi-imaginelab/GECKO
ZoomLDM: Latent Diffusion Model for multi-scale image generation
Nov 25, 2024Abstract:Diffusion models have revolutionized image generation, yet several challenges restrict their application to large-image domains, such as digital pathology and satellite imagery. Given that it is infeasible to directly train a model on 'whole' images from domains with potential gigapixel sizes, diffusion-based generative methods have focused on synthesizing small, fixed-size patches extracted from these images. However, generating small patches has limited applicability since patch-based models fail to capture the global structures and wider context of large images, which can be crucial for synthesizing (semantically) accurate samples. In this paper, to overcome this limitation, we present ZoomLDM, a diffusion model tailored for generating images across multiple scales. Central to our approach is a novel magnification-aware conditioning mechanism that utilizes self-supervised learning (SSL) embeddings and allows the diffusion model to synthesize images at different 'zoom' levels, i.e., fixed-size patches extracted from large images at varying scales. ZoomLDM achieves state-of-the-art image generation quality across all scales, excelling particularly in the data-scarce setting of generating thumbnails of entire large images. The multi-scale nature of ZoomLDM unlocks additional capabilities in large image generation, enabling computationally tractable and globally coherent image synthesis up to $4096 \times 4096$ pixels and $4\times$ super-resolution. Additionally, multi-scale features extracted from ZoomLDM are highly effective in multiple instance learning experiments. We provide high-resolution examples of the generated images on our website https://histodiffusion.github.io/docs/publications/zoomldm/.
SI-MIL: Taming Deep MIL for Self-Interpretability in Gigapixel Histopathology
Dec 22, 2023



Abstract:Introducing interpretability and reasoning into Multiple Instance Learning (MIL) methods for Whole Slide Image (WSI) analysis is challenging, given the complexity of gigapixel slides. Traditionally, MIL interpretability is limited to identifying salient regions deemed pertinent for downstream tasks, offering little insight to the end-user (pathologist) regarding the rationale behind these selections. To address this, we propose Self-Interpretable MIL (SI-MIL), a method intrinsically designed for interpretability from the very outset. SI-MIL employs a deep MIL framework to guide an interpretable branch grounded on handcrafted pathological features, facilitating linear predictions. Beyond identifying salient regions, SI-MIL uniquely provides feature-level interpretations rooted in pathological insights for WSIs. Notably, SI-MIL, with its linear prediction constraints, challenges the prevalent myth of an inevitable trade-off between model interpretability and performance, demonstrating competitive results compared to state-of-the-art methods on WSI-level prediction tasks across three cancer types. In addition, we thoroughly benchmark the local- and global-interpretability of SI-MIL in terms of statistical analysis, a domain expert study, and desiderata of interpretability, namely, user-friendliness and faithfulness.
Attention De-sparsification Matters: Inducing Diversity in Digital Pathology Representation Learning
Sep 12, 2023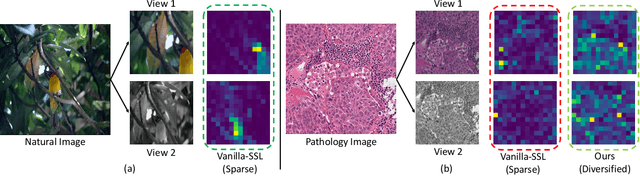
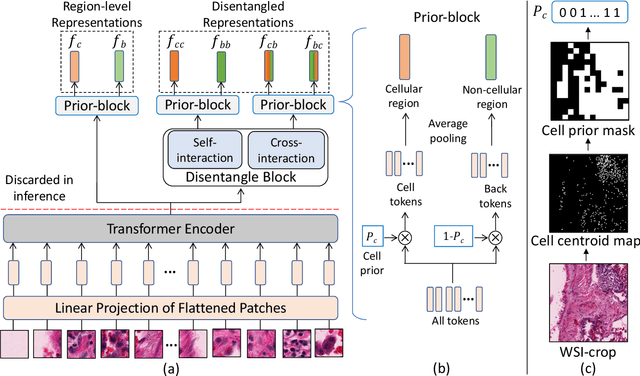
Abstract:We propose DiRL, a Diversity-inducing Representation Learning technique for histopathology imaging. Self-supervised learning techniques, such as contrastive and non-contrastive approaches, have been shown to learn rich and effective representations of digitized tissue samples with limited pathologist supervision. Our analysis of vanilla SSL-pretrained models' attention distribution reveals an insightful observation: sparsity in attention, i.e, models tends to localize most of their attention to some prominent patterns in the image. Although attention sparsity can be beneficial in natural images due to these prominent patterns being the object of interest itself, this can be sub-optimal in digital pathology; this is because, unlike natural images, digital pathology scans are not object-centric, but rather a complex phenotype of various spatially intermixed biological components. Inadequate diversification of attention in these complex images could result in crucial information loss. To address this, we leverage cell segmentation to densely extract multiple histopathology-specific representations, and then propose a prior-guided dense pretext task for SSL, designed to match the multiple corresponding representations between the views. Through this, the model learns to attend to various components more closely and evenly, thus inducing adequate diversification in attention for capturing context rich representations. Through quantitative and qualitative analysis on multiple tasks across cancer types, we demonstrate the efficacy of our method and observe that the attention is more globally distributed.
Unsupervised Histopathology Image Synthesis
Dec 13, 2017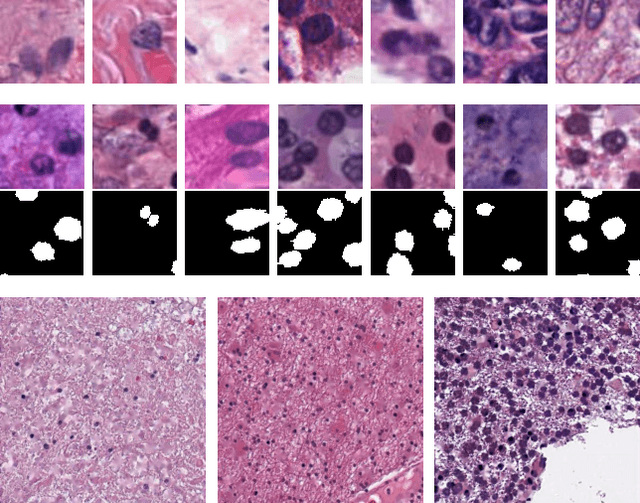
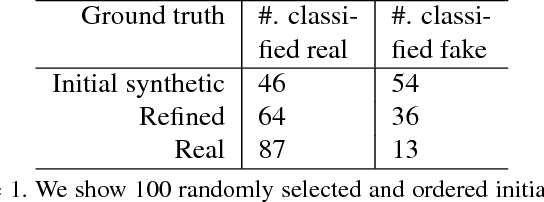
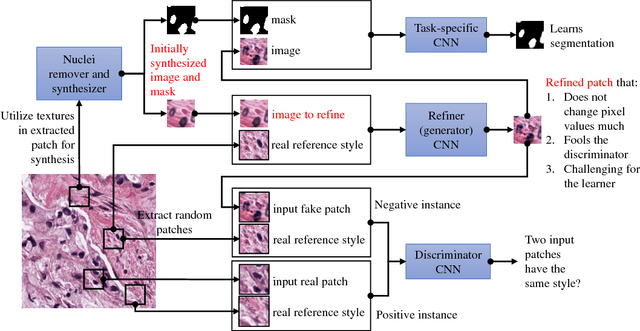
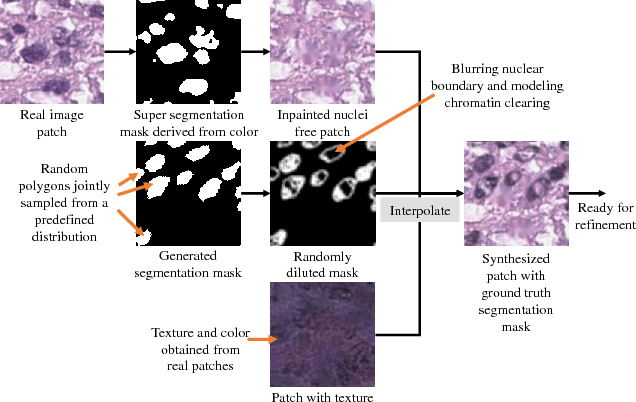
Abstract:Hematoxylin and Eosin stained histopathology image analysis is essential for the diagnosis and study of complicated diseases such as cancer. Existing state-of-the-art approaches demand extensive amount of supervised training data from trained pathologists. In this work we synthesize in an unsupervised manner, large histopathology image datasets, suitable for supervised training tasks. We propose a unified pipeline that: a) generates a set of initial synthetic histopathology images with paired information about the nuclei such as segmentation masks; b) refines the initial synthetic images through a Generative Adversarial Network (GAN) to reference styles; c) trains a task-specific CNN and boosts the performance of the task-specific CNN with on-the-fly generated adversarial examples. Our main contribution is that the synthetic images are not only realistic, but also representative (in reference styles) and relatively challenging for training task-specific CNNs. We test our method for nucleus segmentation using images from four cancer types. When no supervised data exists for a cancer type, our method without supervision cost significantly outperforms supervised methods which perform across-cancer generalization. Even when supervised data exists for all cancer types, our approach without supervision cost performs better than supervised methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge