Qianfei Zhao
PyMIC: A deep learning toolkit for annotation-efficient medical image segmentation
Aug 19, 2022
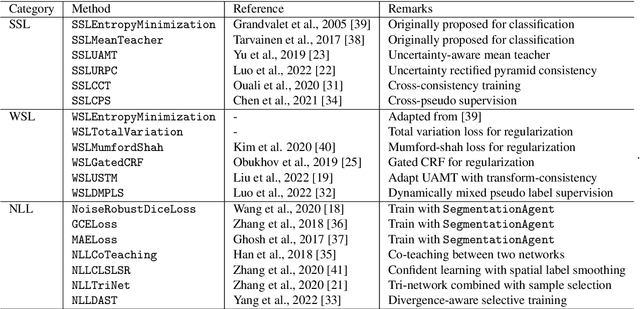
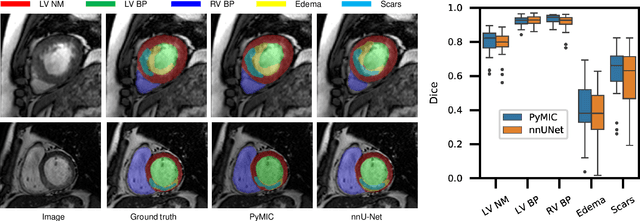
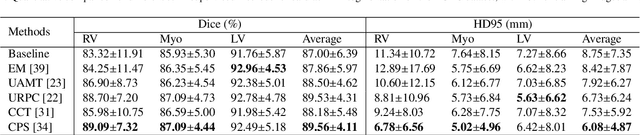
Abstract:Background and Objective: Existing deep learning platforms for medical image segmentation mainly focus on fully supervised segmentation that assumes full and accurate pixel-level annotations are available. We aim to develop a new deep learning toolkit to support annotation-efficient learning for medical image segmentation, which can accelerate and simply the development of deep learning models with limited annotation budget, e.g., learning from partial, sparse or noisy annotations. Methods: Our proposed toolkit named PyMIC is a modular deep learning platform for medical image segmentation tasks. In addition to basic components that support development of high-performance models for fully supervised segmentation, it contains several advanced components that are tailored for learning from imperfect annotations, such as loading annotated and unannounced images, loss functions for unannotated, partially or inaccurately annotated images, and training procedures for co-learning between multiple networks, etc. PyMIC is built on the PyTorch framework and supports development of semi-supervised, weakly supervised and noise-robust learning methods for medical image segmentation. Results: We present four illustrative medical image segmentation tasks based on PyMIC: (1) Achieving competitive performance on fully supervised learning; (2) Semi-supervised cardiac structure segmentation with only 10% training images annotated; (3) Weakly supervised segmentation using scribble annotations; and (4) Learning from noisy labels for chest radiograph segmentation. Conclusions: The PyMIC toolkit is easy to use and facilitates efficient development of medical image segmentation models with imperfect annotations. It is modular and flexible, which enables researchers to develop high-performance models with low annotation cost. The source code is available at: https://github.com/HiLab-git/PyMIC.
HMRNet: High and Multi-Resolution Network with Bidirectional Feature Calibration for Brain Structure Segmentation in Radiotherapy
Jun 07, 2022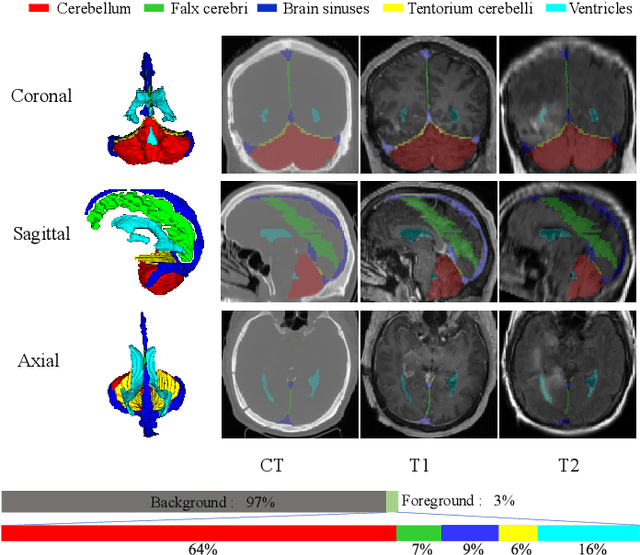
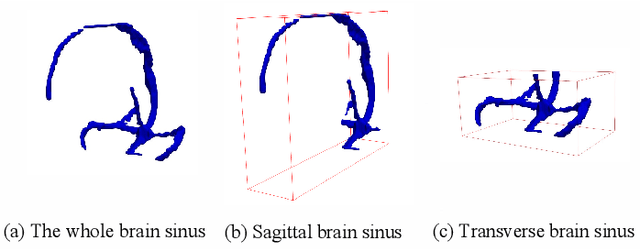
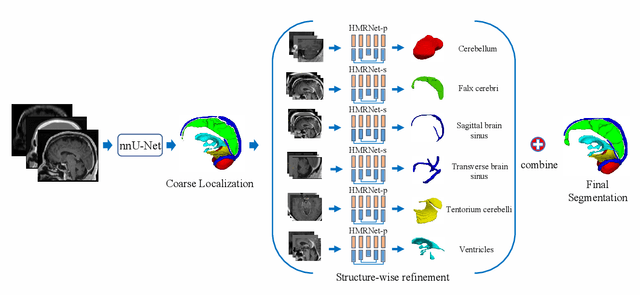

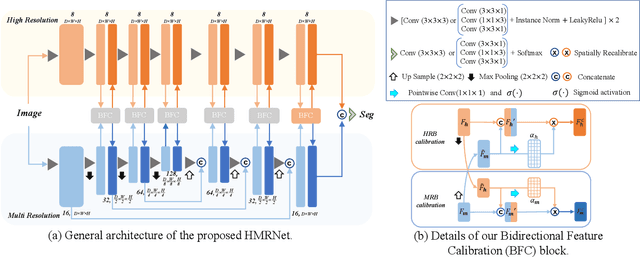
Abstract:Accurate segmentation of Anatomical brain Barriers to Cancer spread (ABCs) plays an important role for automatic delineation of Clinical Target Volume (CTV) of brain tumors in radiotherapy. Despite that variants of U-Net are state-of-the-art segmentation models, they have limited performance when dealing with ABCs structures with various shapes and sizes, especially thin structures (e.g., the falx cerebri) that span only few slices. To deal with this problem, we propose a High and Multi-Resolution Network (HMRNet) that consists of a multi-scale feature learning branch and a high-resolution branch, which can maintain the high-resolution contextual information and extract more robust representations of anatomical structures with various scales. We further design a Bidirectional Feature Calibration (BFC) block to enable the two branches to generate spatial attention maps for mutual feature calibration. Considering the different sizes and positions of ABCs structures, our network was applied after a rough localization of each structure to obtain fine segmentation results. Experiments on the MICCAI 2020 ABCs challenge dataset showed that: 1) Our proposed two-stage segmentation strategy largely outperformed methods segmenting all the structures in just one stage; 2) The proposed HMRNet with two branches can maintain high-resolution representations and is effective to improve the performance on thin structures; 3) The proposed BFC block outperformed existing attention methods using monodirectional feature calibration. Our method won the second place of ABCs 2020 challenge and has a potential for more accurate and reasonable delineation of CTV of brain tumors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge