Nina Montaña-Brown
An X-Ray Is Worth 15 Features: Sparse Autoencoders for Interpretable Radiology Report Generation
Oct 04, 2024Abstract:Radiological services are experiencing unprecedented demand, leading to increased interest in automating radiology report generation. Existing Vision-Language Models (VLMs) suffer from hallucinations, lack interpretability, and require expensive fine-tuning. We introduce SAE-Rad, which uses sparse autoencoders (SAEs) to decompose latent representations from a pre-trained vision transformer into human-interpretable features. Our hybrid architecture combines state-of-the-art SAE advancements, achieving accurate latent reconstructions while maintaining sparsity. Using an off-the-shelf language model, we distil ground-truth reports into radiological descriptions for each SAE feature, which we then compile into a full report for each image, eliminating the need for fine-tuning large models for this task. To the best of our knowledge, SAE-Rad represents the first instance of using mechanistic interpretability techniques explicitly for a downstream multi-modal reasoning task. On the MIMIC-CXR dataset, SAE-Rad achieves competitive radiology-specific metrics compared to state-of-the-art models while using significantly fewer computational resources for training. Qualitative analysis reveals that SAE-Rad learns meaningful visual concepts and generates reports aligning closely with expert interpretations. Our results suggest that SAEs can enhance multimodal reasoning in healthcare, providing a more interpretable alternative to existing VLMs.
Competing for pixels: a self-play algorithm for weakly-supervised segmentation
May 26, 2024Abstract:Weakly-supervised segmentation (WSS) methods, reliant on image-level labels indicating object presence, lack explicit correspondence between labels and regions of interest (ROIs), posing a significant challenge. Despite this, WSS methods have attracted attention due to their much lower annotation costs compared to fully-supervised segmentation. Leveraging reinforcement learning (RL) self-play, we propose a novel WSS method that gamifies image segmentation of a ROI. We formulate segmentation as a competition between two agents that compete to select ROI-containing patches until exhaustion of all such patches. The score at each time-step, used to compute the reward for agent training, represents likelihood of object presence within the selection, determined by an object presence detector pre-trained using only image-level binary classification labels of object presence. Additionally, we propose a game termination condition that can be called by either side upon exhaustion of all ROI-containing patches, followed by the selection of a final patch from each. Upon termination, the agent is incentivised if ROI-containing patches are exhausted or disincentivised if an ROI-containing patch is found by the competitor. This competitive setup ensures minimisation of over- or under-segmentation, a common problem with WSS methods. Extensive experimentation across four datasets demonstrates significant performance improvements over recent state-of-the-art methods. Code: https://github.com/s-sd/spurl/tree/main/wss
Active learning using adaptable task-based prioritisation
Dec 03, 2022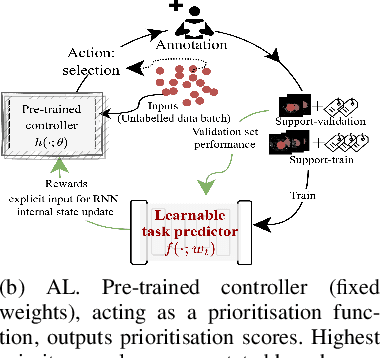
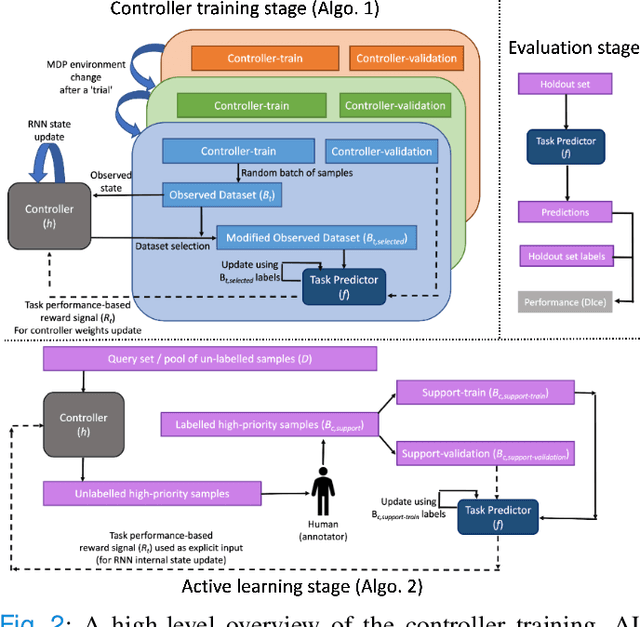
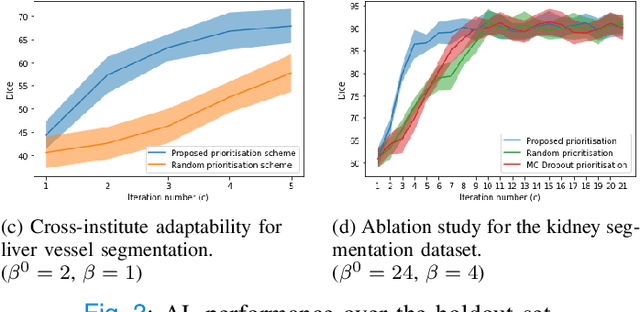
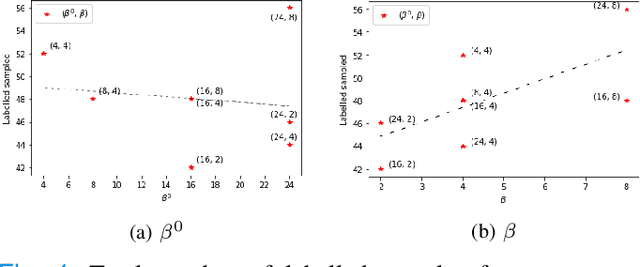
Abstract:Supervised machine learning-based medical image computing applications necessitate expert label curation, while unlabelled image data might be relatively abundant. Active learning methods aim to prioritise a subset of available image data for expert annotation, for label-efficient model training. We develop a controller neural network that measures priority of images in a sequence of batches, as in batch-mode active learning, for multi-class segmentation tasks. The controller is optimised by rewarding positive task-specific performance gain, within a Markov decision process (MDP) environment that also optimises the task predictor. In this work, the task predictor is a segmentation network. A meta-reinforcement learning algorithm is proposed with multiple MDPs, such that the pre-trained controller can be adapted to a new MDP that contains data from different institutes and/or requires segmentation of different organs or structures within the abdomen. We present experimental results using multiple CT datasets from more than one thousand patients, with segmentation tasks of nine different abdominal organs, to demonstrate the efficacy of the learnt prioritisation controller function and its cross-institute and cross-organ adaptability. We show that the proposed adaptable prioritisation metric yields converging segmentation accuracy for the novel class of kidney, unseen in training, using between approximately 40\% to 60\% of labels otherwise required with other heuristic or random prioritisation metrics. For clinical datasets of limited size, the proposed adaptable prioritisation offers a performance improvement of 22.6\% and 10.2\% in Dice score, for tasks of kidney and liver vessel segmentation, respectively, compared to random prioritisation and alternative active sampling strategies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge