Nicholas Turner
Convolutional nets for reconstructing neural circuits from brain images acquired by serial section electron microscopy
Apr 29, 2019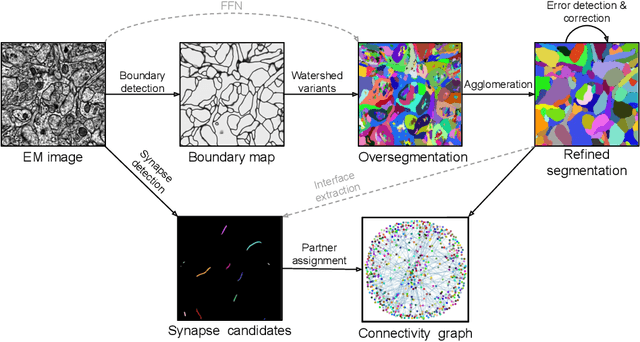
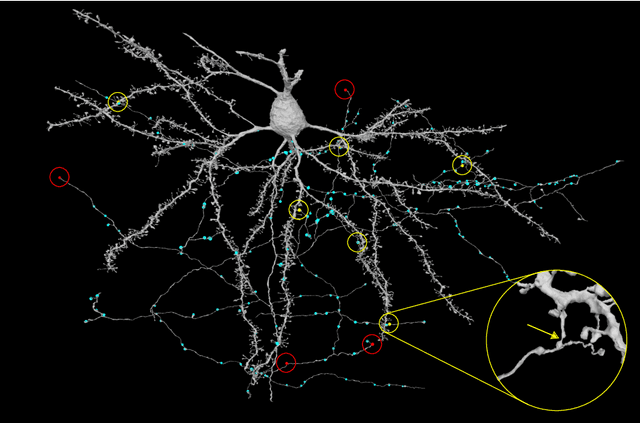

Abstract:Neural circuits can be reconstructed from brain images acquired by serial section electron microscopy. Image analysis has been performed by manual labor for half a century, and efforts at automation date back almost as far. Convolutional nets were first applied to neuronal boundary detection a dozen years ago, and have now achieved impressive accuracy on clean images. Robust handling of image defects is a major outstanding challenge. Convolutional nets are also being employed for other tasks in neural circuit reconstruction: finding synapses and identifying synaptic partners, extending or pruning neuronal reconstructions, and aligning serial section images to create a 3D image stack. Computational systems are being engineered to handle petavoxel images of cubic millimeter brain volumes.
Synaptic Partner Assignment Using Attentional Voxel Association Networks
Apr 22, 2019

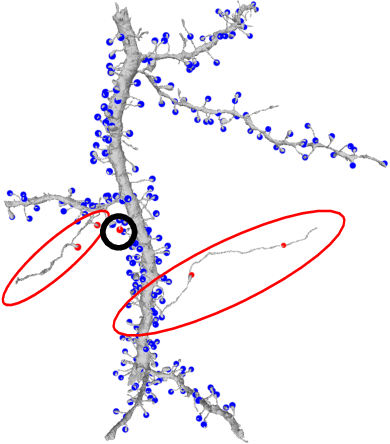
Abstract:Connectomics aims to recover a complete set of synaptic connections within a dataset imaged by electron microscopy. Most systems for locating synapses use voxelwise classifier models, and train these classifiers to reproduce binary masks of synaptic clefts. However, only recent work has included a way to identify the synaptic partners that communicate at synaptic cleft segments. Here, we present a novel method for associating synaptic cleft segments with their synaptic partners using a convolutional network trained to associate the mask of a cleft with the voxels of its synaptic partners. The network takes the local image context and a mask of a single cleft segment as input. It is trained to produce two volumes of output: one which labels the voxels of the presynaptic partner within the input image, and another similar volume for the postsynaptic partner. The cleft mask acts as an attentional gating signal for the network, in that two clefts with the same local image context often have different partners. We find that an implementation of this approach performs well on a dataset of mouse somatosensory cortex, and evaluate it as part of a combined system to predict both clefts and connections.
Reconstructing neuronal anatomy from whole-brain images
Mar 17, 2019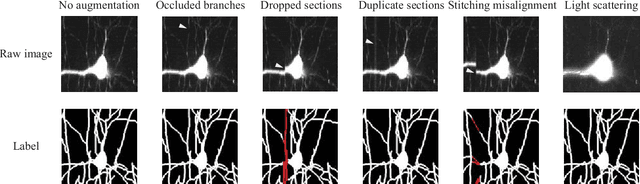
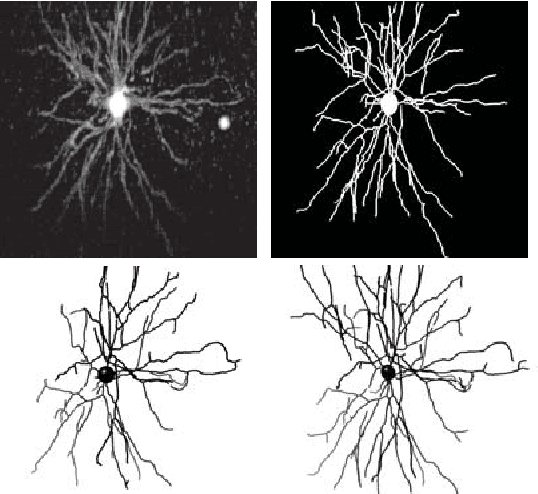
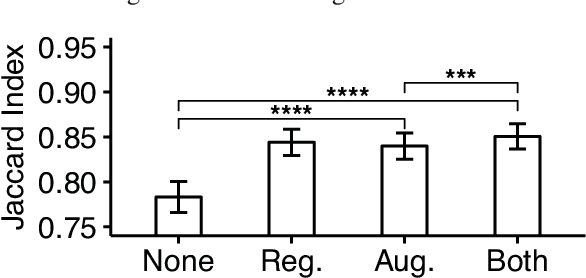
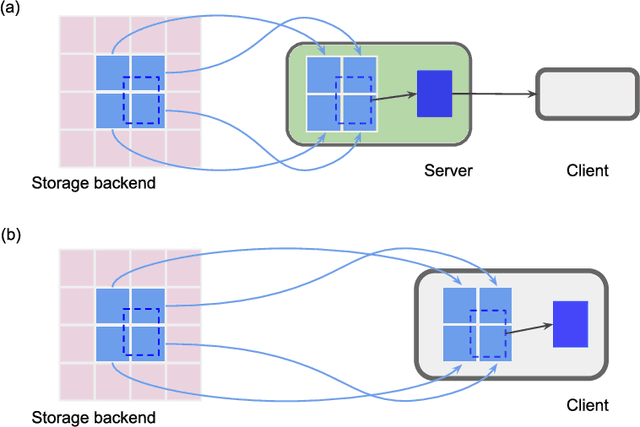
Abstract:Reconstructing multiple molecularly defined neurons from individual brains and across multiple brain regions can reveal organizational principles of the nervous system. However, high resolution imaging of the whole brain is a technically challenging and slow process. Recently, oblique light sheet microscopy has emerged as a rapid imaging method that can provide whole brain fluorescence microscopy at a voxel size of 0.4 by 0.4 by 2.5 cubic microns. On the other hand, complex image artifacts due to whole-brain coverage produce apparent discontinuities in neuronal arbors. Here, we present connectivity-preserving methods and data augmentation strategies for supervised learning of neuroanatomy from light microscopy using neural networks. We quantify the merit of our approach by implementing an end-to-end automated tracing pipeline. Lastly, we demonstrate a scalable, distributed implementation that can reconstruct the large datasets that sub-micron whole-brain images produce.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge