Pavel Osten
Reconstructing neuronal anatomy from whole-brain images
Mar 17, 2019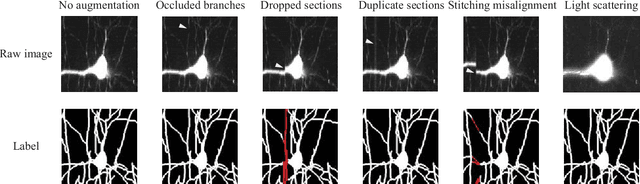
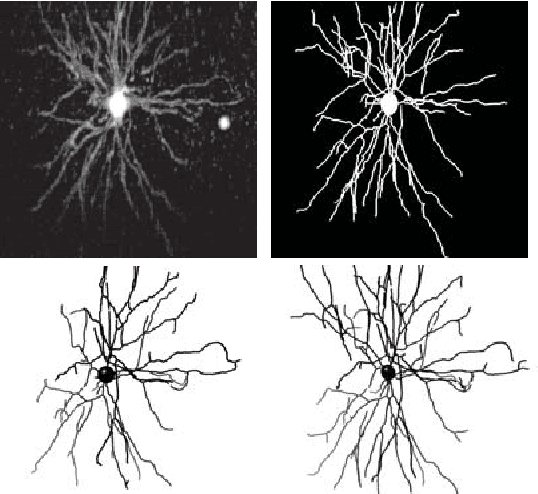
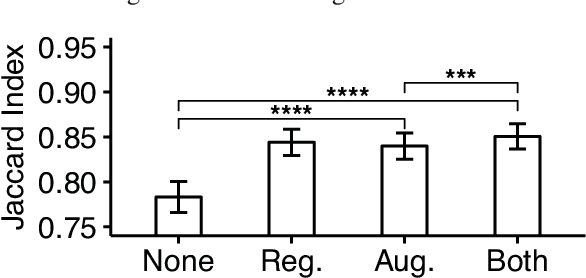
Abstract:Reconstructing multiple molecularly defined neurons from individual brains and across multiple brain regions can reveal organizational principles of the nervous system. However, high resolution imaging of the whole brain is a technically challenging and slow process. Recently, oblique light sheet microscopy has emerged as a rapid imaging method that can provide whole brain fluorescence microscopy at a voxel size of 0.4 by 0.4 by 2.5 cubic microns. On the other hand, complex image artifacts due to whole-brain coverage produce apparent discontinuities in neuronal arbors. Here, we present connectivity-preserving methods and data augmentation strategies for supervised learning of neuroanatomy from light microscopy using neural networks. We quantify the merit of our approach by implementing an end-to-end automated tracing pipeline. Lastly, we demonstrate a scalable, distributed implementation that can reconstruct the large datasets that sub-micron whole-brain images produce.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge