Michal Ozery-Flato
BioVERSE: Representation Alignment of Biomedical Modalities to LLMs for Multi-Modal Reasoning
Oct 01, 2025



Abstract:Recent advances in large language models (LLMs) and biomedical foundation models (BioFMs) have achieved strong results in biological text reasoning, molecular modeling, and single-cell analysis, yet they remain siloed in disjoint embedding spaces, limiting cross-modal reasoning. We present BIOVERSE (Biomedical Vector Embedding Realignment for Semantic Engagement), a two-stage approach that adapts pretrained BioFMs as modality encoders and aligns them with LLMs through lightweight, modality-specific projection layers. The approach first aligns each modality to a shared LLM space through independently trained projections, allowing them to interoperate naturally, and then applies standard instruction tuning with multi-modal data to bring them together for downstream reasoning. By unifying raw biomedical data with knowledge embedded in LLMs, the approach enables zero-shot annotation, cross-modal question answering, and interactive, explainable dialogue. Across tasks spanning cell-type annotation, molecular description, and protein function reasoning, compact BIOVERSE configurations surpass larger LLM baselines while enabling richer, generative outputs than existing BioFMs, establishing a foundation for principled multi-modal biomedical reasoning.
MAMMAL -- Molecular Aligned Multi-Modal Architecture and Language
Oct 28, 2024Abstract:Drug discovery typically consists of multiple steps, including identifying a target protein key to a disease's etiology, validating that interacting with this target could prevent symptoms or cure the disease, discovering a small molecule or biologic therapeutic to interact with it, and optimizing the candidate molecule through a complex landscape of required properties. Drug discovery related tasks often involve prediction and generation while considering multiple entities that potentially interact, which poses a challenge for typical AI models. For this purpose we present MAMMAL - Molecular Aligned Multi-Modal Architecture and Language - a method that we applied to create a versatile multi-task foundation model ibm/biomed.omics.bl.sm.ma-ted-458m that learns from large-scale biological datasets (2 billion samples) across diverse modalities, including proteins, small molecules, and genes. We introduce a prompt syntax that supports a wide range of classification, regression, and generation tasks. It allows combining different modalities and entity types as inputs and/or outputs. Our model handles combinations of tokens and scalars and enables the generation of small molecules and proteins, property prediction, and transcriptomic lab test predictions. We evaluated the model on 11 diverse downstream tasks spanning different steps within a typical drug discovery pipeline, where it reaches new SOTA in 9 tasks and is comparable to SOTA in 2 tasks. This performance is achieved while using a unified architecture serving all tasks, in contrast to the original SOTA performance achieved using tailored architectures. The model code and pretrained weights are publicly available at https://github.com/BiomedSciAI/biomed-multi-alignment and https://huggingface.co/ibm/biomed.omics.bl.sm.ma-ted-458m.
AI Age Discrepancy: A Novel Parameter for Frailty Assessment in Kidney Tumor Patients
Jul 02, 2024Abstract:Kidney cancer is a global health concern, and accurate assessment of patient frailty is crucial for optimizing surgical outcomes. This paper introduces AI Age Discrepancy, a novel metric derived from machine learning analysis of preoperative abdominal CT scans, as a potential indicator of frailty and postoperative risk in kidney cancer patients. This retrospective study of 599 patients from the 2023 Kidney Tumor Segmentation (KiTS) challenge dataset found that a higher AI Age Discrepancy is significantly associated with longer hospital stays and lower overall survival rates, independent of established factors. This suggests that AI Age Discrepancy may provide valuable insights into patient frailty and could thus inform clinical decision-making in kidney cancer treatment.
A large dataset curation and benchmark for drug target interaction
Jan 30, 2024Abstract:Bioactivity data plays a key role in drug discovery and repurposing. The resource-demanding nature of \textit{in vitro} and \textit{in vivo} experiments, as well as the recent advances in data-driven computational biochemistry research, highlight the importance of \textit{in silico} drug target interaction (DTI) prediction approaches. While numerous large public bioactivity data sources exist, research in the field could benefit from better standardization of existing data resources. At present, different research works that share similar goals are often difficult to compare properly because of different choices of data sources and train/validation/test split strategies. Additionally, many works are based on small data subsets, leading to results and insights of possible limited validity. In this paper we propose a way to standardize and represent efficiently a very large dataset curated from multiple public sources, split the data into train, validation and test sets based on different meaningful strategies, and provide a concrete evaluation protocol to accomplish a benchmark. We analyze the proposed data curation, prove its usefulness and validate the proposed benchmark through experimental studies based on an existing neural network model.
Adversarial Balancing for Causal Inference
Oct 17, 2018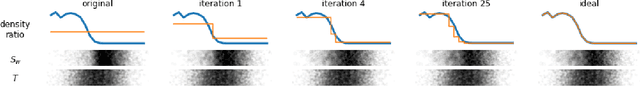
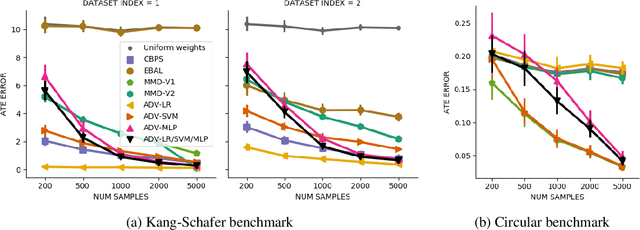
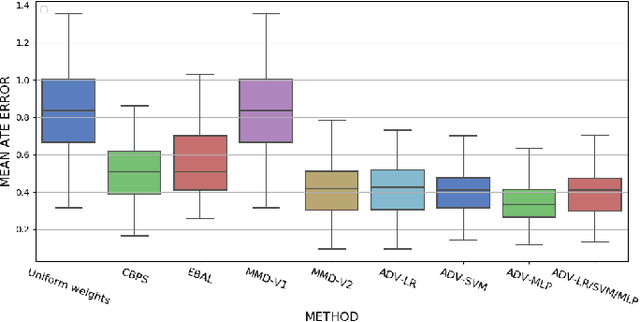
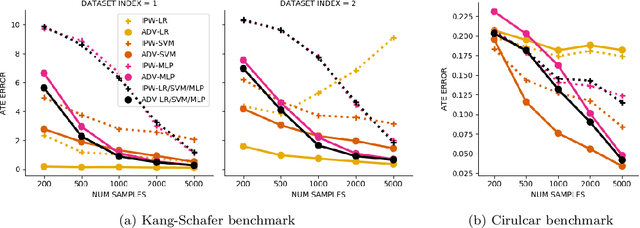
Abstract:Biases in observational data pose a major challenge to estimation methods for the effect of treatments. An important technique that accounts for these biases is reweighting samples to minimize the discrepancy between treatment groups. Inverse probability weighting, a popular weighting technique, models the conditional treatment probability given covariates. However, it is overly sensitive to model misspecification and suffers from large estimation variance. Recent methods attempt to alleviate these limitations by finding weights that minimize a selected discrepancy measure between the reweighted populations. We present a new reweighting approach that uses classification error as a measure of similarity between datasets. Our proposed framework uses bi-level optimization to alternately train a discriminator to minimize classification error, and a balancing weights generator to maximize this error. This approach borrows principles from generative adversarial networks (GANs) that aim to exploit the power of classifiers for discrepancy measure estimation. We tested our approach on several benchmarks. The results of our experiments demonstrate the effectiveness and robustness of this approach in estimating causal effects under different data generating settings.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge