Licheng Zong
Search-R2: Enhancing Search-Integrated Reasoning via Actor-Refiner Collaboration
Feb 03, 2026Abstract:Search-integrated reasoning enables language agents to transcend static parametric knowledge by actively querying external sources. However, training these agents via reinforcement learning is hindered by the multi-scale credit assignment problem: existing methods typically rely on sparse, trajectory-level rewards that fail to distinguish between high-quality reasoning and fortuitous guesses, leading to redundant or misleading search behaviors. To address this, we propose Search-R2, a novel Actor-Refiner collaboration framework that enhances reasoning through targeted intervention, with both components jointly optimized during training. Our approach decomposes the generation process into an Actor, which produces initial reasoning trajectories, and a Meta-Refiner, which selectively diagnoses and repairs flawed steps via a 'cut-and-regenerate' mechanism. To provide fine-grained supervision, we introduce a hybrid reward design that couples outcome correctness with a dense process reward quantifying the information density of retrieved evidence. Theoretically, we formalize the Actor-Refiner interaction as a smoothed mixture policy, proving that selective correction yields strict performance gains over strong baselines. Extensive experiments across various general and multi-hop QA datasets demonstrate that Search-R2 consistently outperforms strong RAG and RL-based baselines across model scales, achieving superior reasoning accuracy with minimal overhead.
Enhancing Biomedical Knowledge Retrieval-Augmented Generation with Self-Rewarding Tree Search and Proximal Policy Optimization
Jun 17, 2024



Abstract:Large Language Models (LLMs) have shown great potential in the biomedical domain with the advancement of retrieval-augmented generation (RAG). However, existing retrieval-augmented approaches face challenges in addressing diverse queries and documents, particularly for medical knowledge queries, resulting in sub-optimal performance. To address these limitations, we propose a novel plug-and-play LLM-based retrieval method called Self-Rewarding Tree Search (SeRTS) based on Monte Carlo Tree Search (MCTS) and a self-rewarding paradigm. By combining the reasoning capabilities of LLMs with the effectiveness of tree search, SeRTS boosts the zero-shot performance of retrieving high-quality and informative results for RAG. We further enhance retrieval performance by fine-tuning LLMs with Proximal Policy Optimization (PPO) objectives using the trajectories collected by SeRTS as feedback. Controlled experiments using the BioASQ-QA dataset with GPT-3.5-Turbo and LLama2-7b demonstrate that our method significantly improves the performance of the BM25 retriever and surpasses the strong baseline of self-reflection in both efficiency and scalability. Moreover, SeRTS generates higher-quality feedback for PPO training than self-reflection. Our proposed method effectively adapts LLMs to document retrieval tasks, enhancing their ability to retrieve highly relevant documents for RAG in the context of medical knowledge queries. This work presents a significant step forward in leveraging LLMs for accurate and comprehensive biomedical question answering.
HMD-AMP: Protein Language-Powered Hierarchical Multi-label Deep Forest for Annotating Antimicrobial Peptides
Nov 11, 2021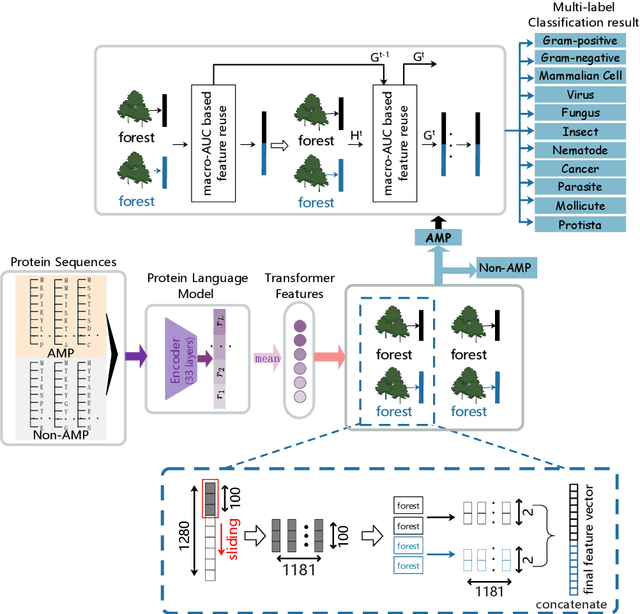
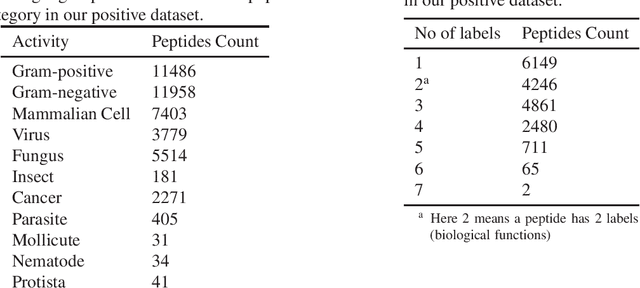

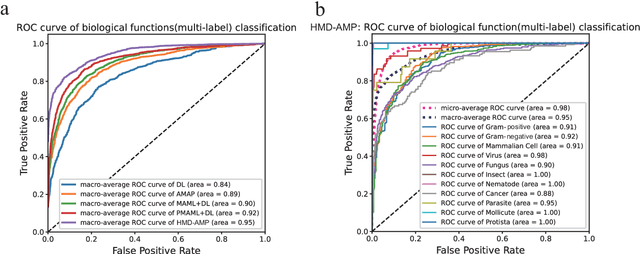
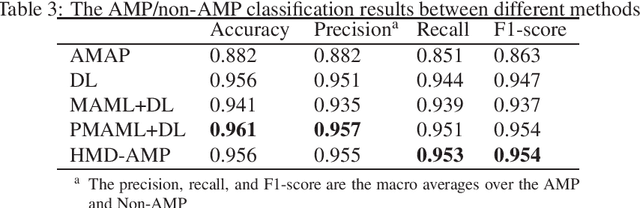
Abstract:Identifying the targets of an antimicrobial peptide is a fundamental step in studying the innate immune response and combating antibiotic resistance, and more broadly, precision medicine and public health. There have been extensive studies on the statistical and computational approaches to identify (i) whether a peptide is an antimicrobial peptide (AMP) or a non-AMP and (ii) which targets are these sequences effective to (Gram-positive, Gram-negative, etc.). Despite the existing deep learning methods on this problem, most of them are unable to handle the small AMP classes (anti-insect, anti-parasite, etc.). And more importantly, some AMPs can have multiple targets, which the previous methods fail to consider. In this study, we build a diverse and comprehensive multi-label protein sequence database by collecting and cleaning amino acids from various AMP databases. To generate efficient representations and features for the small classes dataset, we take advantage of a protein language model trained on 250 million protein sequences. Based on that, we develop an end-to-end hierarchical multi-label deep forest framework, HMD-AMP, to annotate AMP comprehensively. After identifying an AMP, it further predicts what targets the AMP can effectively kill from eleven available classes. Extensive experiments suggest that our framework outperforms state-of-the-art models in both the binary classification task and the multi-label classification task, especially on the minor classes.The model is robust against reduced features and small perturbations and produces promising results. We believe HMD-AMP contributes to both the future wet-lab investigations of the innate structural properties of different antimicrobial peptides and build promising empirical underpinnings for precise medicine with antibiotics.
Protein-RNA interaction prediction with deep learning: Structure matters
Jul 26, 2021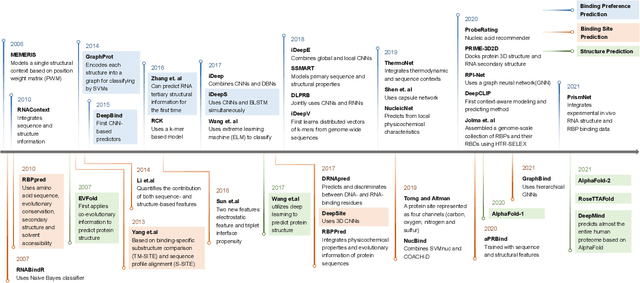
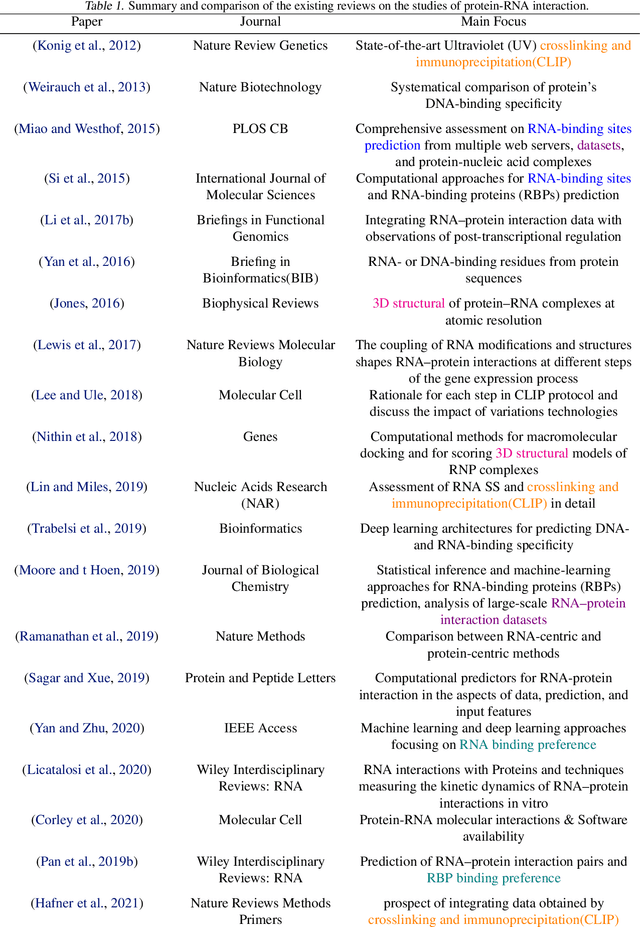
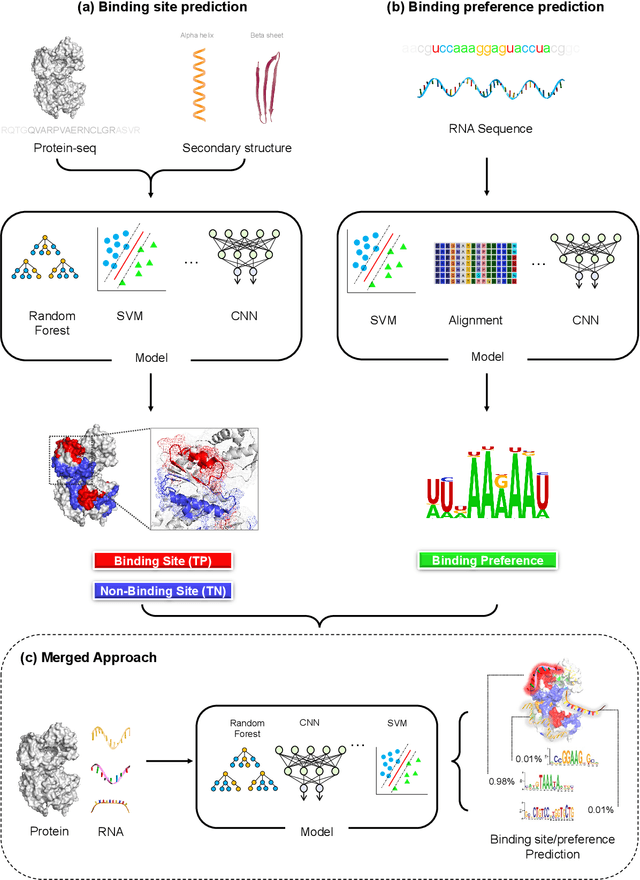

Abstract:Protein-RNA interactions are of vital importance to a variety of cellular activities. Both experimental and computational techniques have been developed to study the interactions. Due to the limitation of the previous database, especially the lack of protein structure data, most of the existing computational methods rely heavily on the sequence data, with only a small portion of the methods utilizing the structural information. Recently, AlphaFold has revolutionized the entire protein and biology field. Foreseeably, the protein-RNA interaction prediction will also be promoted significantly in the upcoming years. In this work, we give a thorough review of this field, surveying both the binding site and binding preference prediction problems and covering the commonly used datasets, features, and models. We also point out the potential challenges and opportunities in this field. This survey summarizes the development of the RBP-RNA interaction field in the past and foresees its future development in the post-AlphaFold era.
Model Adaption Object Detection System for Robot
Nov 20, 2019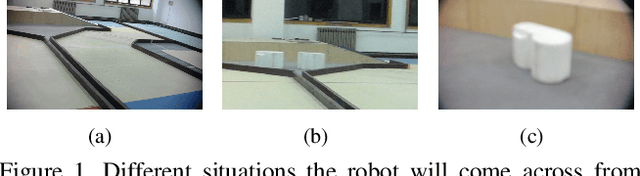
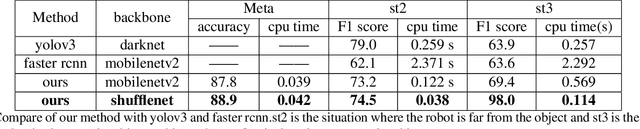
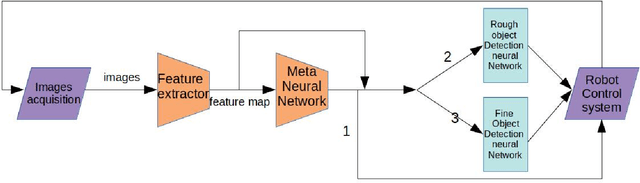

Abstract:Object detection for robot guidance is a crucial mission for autonomous robots, which has provoked extensive attention for researchers. However, the changing view of robot movement and limited available data hinder the research in this area. To address these matters, we proposed a new vision system for robots, the model adaptation object detection system. Instead of using a single one to solve problems, We made use of different object detection neural networks to guide the robot in accordance with various situations, with the help of a meta neural network to allocate the object detection neural networks. Furthermore, taking advantage of transfer learning technology and depthwise separable convolutions, our model is easy to train and can address small dataset problems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge