Josef Hoppe
Don't be Afraid of Cell Complexes! An Introduction from an Applied Perspective
Jun 11, 2025Abstract:Cell complexes (CCs) are a higher-order network model deeply rooted in algebraic topology that has gained interest in signal processing and network science recently. However, while the processing of signals supported on CCs can be described in terms of easily-accessible algebraic or combinatorial notions, the commonly presented definition of CCs is grounded in abstract concepts from topology and remains disconnected from the signal processing methods developed for CCs. In this paper, we aim to bridge this gap by providing a simplified definition of CCs that is accessible to a wider audience and can be used in practical applications. Specifically, we first introduce a simplified notion of abstract regular cell complexes (ARCCs). These ARCCs only rely on notions from algebra and can be shown to be equivalent to regular cell complexes for most practical applications. Second, using this new definition we provide an accessible introduction to (abstract) cell complexes from a perspective of network science and signal processing. Furthermore, as many practical applications work with CCs of dimension 2 and below, we provide an even simpler definition for this case that significantly simplifies understanding and working with CCs in practice.
Topological Trajectory Classification and Landmark Inference on Simplicial Complexes
Dec 04, 2024Abstract:We consider the problem of classifying trajectories on a discrete or discretised 2-dimensional manifold modelled by a simplicial complex. Previous works have proposed to project the trajectories into the harmonic eigenspace of the Hodge Laplacian, and then cluster the resulting embeddings. However, if the considered space has vanishing homology (i.e., no "holes"), then the harmonic space of the 1-Hodge Laplacian is trivial and thus the approach fails. Here we propose to view this issue akin to a sensor placement problem and present an algorithm that aims to learn "optimal holes" to distinguish a set of given trajectory classes. Specifically, given a set of labelled trajectories, which we interpret as edge-flows on the underlying simplicial complex, we search for 2-simplices whose deletion results in an optimal separation of the trajectory labels according to the corresponding spectral embedding of the trajectories into the harmonic space. Finally, we generalise this approach to the unsupervised setting.
TopoX: A Suite of Python Packages for Machine Learning on Topological Domains
Feb 07, 2024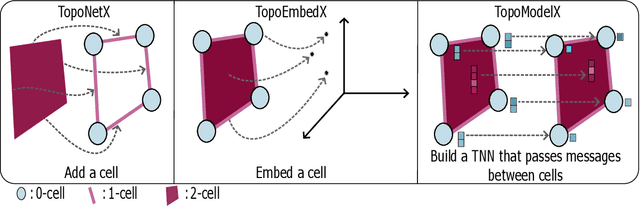
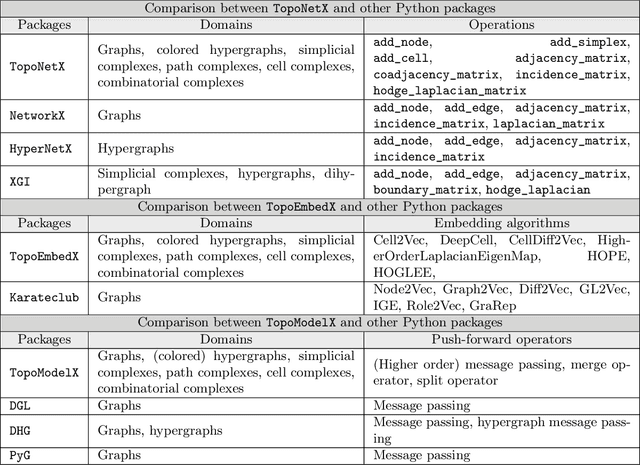
Abstract:We introduce topox, a Python software suite that provides reliable and user-friendly building blocks for computing and machine learning on topological domains that extend graphs: hypergraphs, simplicial, cellular, path and combinatorial complexes. topox consists of three packages: toponetx facilitates constructing and computing on these domains, including working with nodes, edges and higher-order cells; topoembedx provides methods to embed topological domains into vector spaces, akin to popular graph-based embedding algorithms such as node2vec; topomodelx is built on top of PyTorch and offers a comprehensive toolbox of higher-order message passing functions for neural networks on topological domains. The extensively documented and unit-tested source code of topox is available under MIT license at https://github.com/pyt-team.
ICML 2023 Topological Deep Learning Challenge : Design and Results
Oct 02, 2023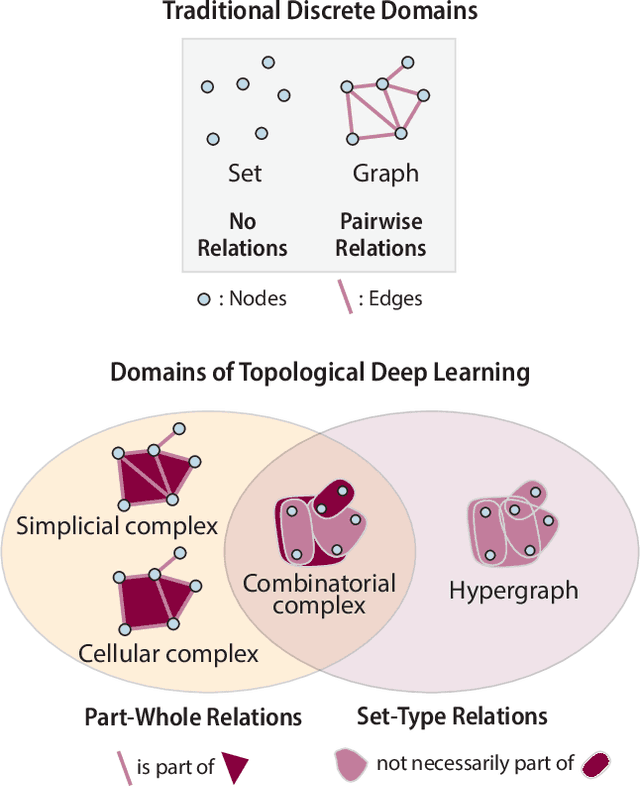
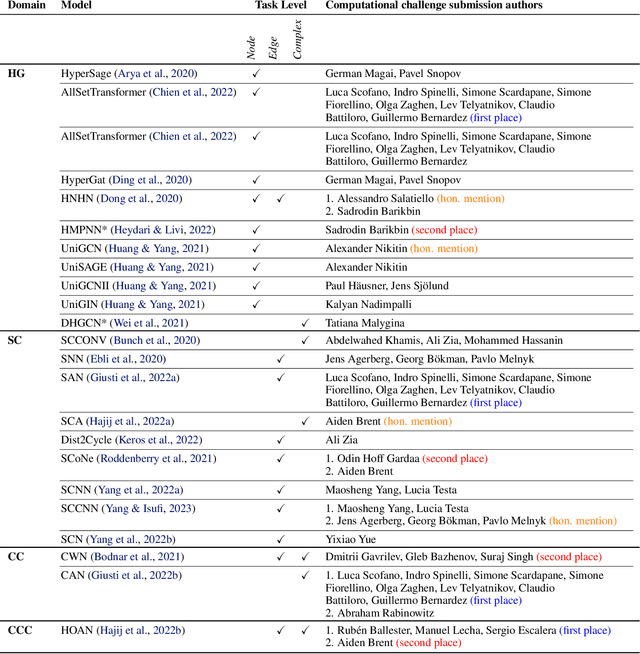
Abstract:This paper presents the computational challenge on topological deep learning that was hosted within the ICML 2023 Workshop on Topology and Geometry in Machine Learning. The competition asked participants to provide open-source implementations of topological neural networks from the literature by contributing to the python packages TopoNetX (data processing) and TopoModelX (deep learning). The challenge attracted twenty-eight qualifying submissions in its two-month duration. This paper describes the design of the challenge and summarizes its main findings.
Representing Edge Flows on Graphs via Sparse Cell Complexes
Sep 15, 2023



Abstract:Obtaining sparse, interpretable representations of observable data is crucial in many machine learning and signal processing tasks. For data representing flows along the edges of a graph, an intuitively interpretable way to obtain such representations is to lift the graph structure to a simplicial complex: The eigenvectors of the associated Hodge-Laplacian, respectively the incidence matrices of the corresponding simplicial complex then induce a Hodge decomposition, which can be used to represent the observed data in terms of gradient, curl, and harmonic flows. In this paper, we generalize this approach to cellular complexes and introduce the cell inference optimization problem, i.e., the problem of augmenting the observed graph by a set of cells, such that the eigenvectors of the associated Hodge Laplacian provide a sparse, interpretable representation of the observed edge flows on the graph. We show that this problem is NP-hard and introduce an efficient approximation algorithm for its solution. Experiments on real-world and synthetic data demonstrate that our algorithm outperforms current state-of-the-art methods while being computationally efficient.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge