Hongjie Wu
Enhancing Diffusion Model Stability for Image Restoration via Gradient Management
Jul 09, 2025

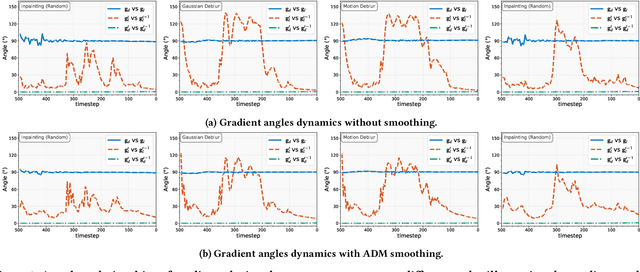

Abstract:Diffusion models have shown remarkable promise for image restoration by leveraging powerful priors. Prominent methods typically frame the restoration problem within a Bayesian inference framework, which iteratively combines a denoising step with a likelihood guidance step. However, the interactions between these two components in the generation process remain underexplored. In this paper, we analyze the underlying gradient dynamics of these components and identify significant instabilities. Specifically, we demonstrate conflicts between the prior and likelihood gradient directions, alongside temporal fluctuations in the likelihood gradient itself. We show that these instabilities disrupt the generative process and compromise restoration performance. To address these issues, we propose Stabilized Progressive Gradient Diffusion (SPGD), a novel gradient management technique. SPGD integrates two synergistic components: (1) a progressive likelihood warm-up strategy to mitigate gradient conflicts; and (2) adaptive directional momentum (ADM) smoothing to reduce fluctuations in the likelihood gradient. Extensive experiments across diverse restoration tasks demonstrate that SPGD significantly enhances generation stability, leading to state-of-the-art performance in quantitative metrics and visually superior results. Code is available at \href{https://github.com/74587887/SPGD}{here}.
Task-oriented Uncertainty Collaborative Learning for Label-Efficient Brain Tumor Segmentation
Mar 07, 2025



Abstract:Multi-contrast magnetic resonance imaging (MRI) plays a vital role in brain tumor segmentation and diagnosis by leveraging complementary information from different contrasts. Each contrast highlights specific tumor characteristics, enabling a comprehensive understanding of tumor morphology, edema, and pathological heterogeneity. However, existing methods still face the challenges of multi-level specificity perception across different contrasts, especially with limited annotations. These challenges include data heterogeneity, granularity differences, and interference from redundant information. To address these limitations, we propose a Task-oriented Uncertainty Collaborative Learning (TUCL) framework for multi-contrast MRI segmentation. TUCL introduces a task-oriented prompt attention (TPA) module with intra-prompt and cross-prompt attention mechanisms to dynamically model feature interactions across contrasts and tasks. Additionally, a cyclic process is designed to map the predictions back to the prompt to ensure that the prompts are effectively utilized. In the decoding stage, the TUCL framework proposes a dual-path uncertainty refinement (DUR) strategy which ensures robust segmentation by refining predictions iteratively. Extensive experimental results on limited labeled data demonstrate that TUCL significantly improves segmentation accuracy (88.2\% in Dice and 10.853 mm in HD95). It shows that TUCL has the potential to extract multi-contrast information and reduce the reliance on extensive annotations. The code is available at: https://github.com/Zhenxuan-Zhang/TUCL_BrainSeg.
Diffusion Posterior Proximal Sampling for Image Restoration
Feb 25, 2024



Abstract:Diffusion models have demonstrated remarkable efficacy in generating high-quality samples. Existing diffusion-based image restoration algorithms exploit pre-trained diffusion models to leverage data priors, yet they still preserve elements inherited from the unconditional generation paradigm. These strategies initiate the denoising process with pure white noise and incorporate random noise at each generative step, leading to over-smoothed results. In this paper, we introduce a refined paradigm for diffusion-based image restoration. Specifically, we opt for a sample consistent with the measurement identity at each generative step, exploiting the sampling selection as an avenue for output stability and enhancement. Besides, we start the restoration process with an initialization combined with the measurement signal, providing supplementary information to better align the generative process. Extensive experimental results and analyses validate the effectiveness of our proposed approach across diverse image restoration tasks.
SemanticCAP: Chromatin Accessibility Prediction Enhanced by Features Learning from a Language Model
Apr 06, 2022
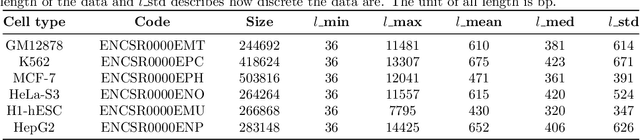
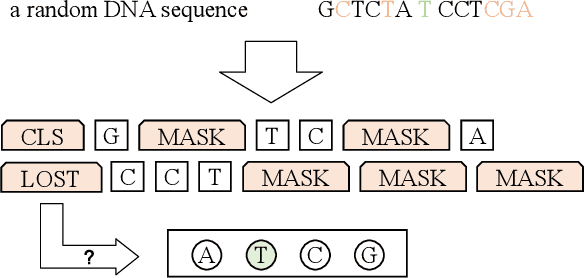
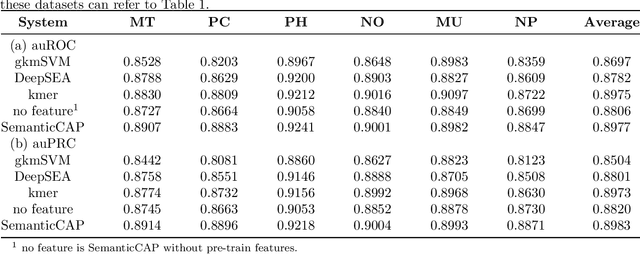
Abstract:A large number of inorganic and organic compounds are able to bind DNA and form complexes, among which drug-related molecules are important. Chromatin accessibility changes not only directly affects drug-DNA interactions, but also promote or inhibit the expression of critical genes associated with drug resistance by affecting the DNA binding capacity of TFs and transcriptional regulators. However, Biological experimental techniques for measuring it are expensive and time consuming. In recent years, several kinds of computational methods have been proposed to identify accessible regions of the genome. Existing computational models mostly ignore the contextual information of bases in gene sequences. To address these issues, we proposed a new solution named SemanticCAP. It introduces a gene language model which models the context of gene sequences, thus being able to provide an effective representation of a certain site in gene sequences. Basically, we merge the features provided by the gene language model into our chromatin accessibility model. During the process, we designed some methods to make feature fusion smoother. Compared with other systems under public benchmarks, our model proved to have better performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge