Gilberto Ochoa-Ruiz
FedAgain: A Trust-Based and Robust Federated Learning Strategy for an Automated Kidney Stone Identification in Ureteroscopy
Mar 19, 2026Abstract:The reliability of artificial intelligence (AI) in medical imaging critically depends on its robustness to heterogeneous and corrupted images acquired with diverse devices across different hospitals which is highly challenging. Therefore, this paper introduces FedAgain, a trust-based Federated Learning (Federated Learning) strategy designed to enhance robustness and generalization for automated kidney stone identification from endoscopic images. FedAgain integrates a dual trust mechanism that combines benchmark reliability and model divergence to dynamically weight client contributions, mitigating the impact of noisy or adversarial updates during aggregation. The framework enables the training of collaborative models across multiple institutions while preserving data privacy and promoting stable convergence under real-world conditions. Extensive experiments across five datasets, including two canonical benchmarks (MNIST and CIFAR-10), two private multi-institutional kidney stone datasets, and one public dataset (MyStone), demonstrate that FedAgain consistently outperforms standard Federated Learning baselines under non-identically and independently distributed (non-IID) data and corrupted-client scenarios. By maintaining diagnostic accuracy and performance stability under varying conditions, FedAgain represents a practical advance toward reliable, privacy-preserving, and clinically deployable federated AI for medical imaging.
A comprehensive multimodal dataset and benchmark for ulcerative colitis scoring in endoscopy
Mar 15, 2026Abstract:Ulcerative colitis (UC) is a chronic mucosal inflammatory condition that places patients at increased risk of colorectal cancer. Colonoscopic surveillance remains the gold standard for assessing disease activity, and reporting typically relies on standardised endoscopic scoring metrics. The most widely used is the Mayo Endoscopic Score (MES), with some centres also adopting the Ulcerative Colitis Endoscopic Index of Severity (UCEIS). Both are descriptive assessments of mucosal inflammation (MES: 0 to 3; UCEIS: 0 to 8), where higher values indicate more severe disease. However, computational methods for automatically predicting these scores remain limited, largely due to the lack of publicly available expert-annotated datasets and the absence of robust benchmarking. There is also a significant research gap in generating clinically meaningful descriptions of UC images, despite image captioning being a well-established computer vision task. Variability in endoscopic systems and procedural workflows across centres further highlights the need for multi-centre datasets to ensure algorithmic robustness and generalisability. In this work, we introduce a curated multi-centre, multi-resolution dataset that includes expert-validated MES and UCEIS labels, alongside detailed clinical descriptions. To our knowledge, this is the first comprehensive dataset that combines dual scoring metrics for classification tasks with expert-generated captions describing mucosal appearance and clinically accepted reasoning for image captioning. This resource opens new opportunities for developing clinically meaningful multimodal algorithms. In addition to the dataset, we also provide benchmarking using convolutional neural networks, vision transformers, hybrid models, and widely used multimodal vision-language captioning algorithms.
Evaluation of Few-Shot Learning Methods for Kidney Stone Type Recognition in Ureteroscopy
May 23, 2025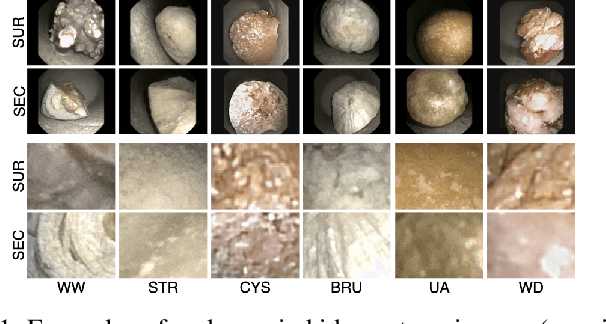
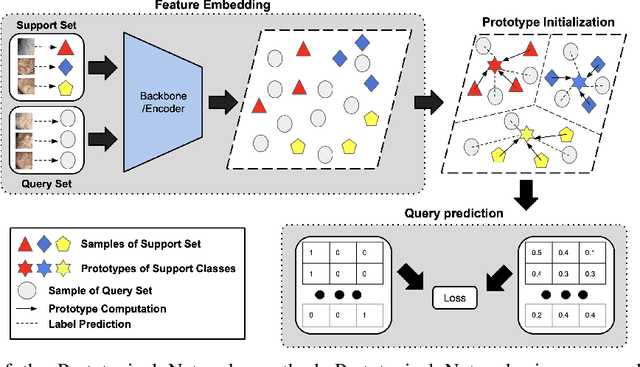
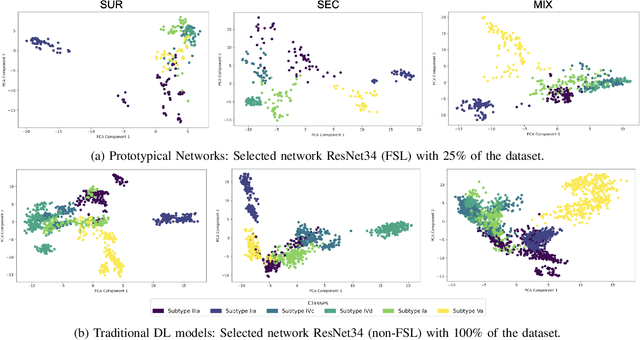
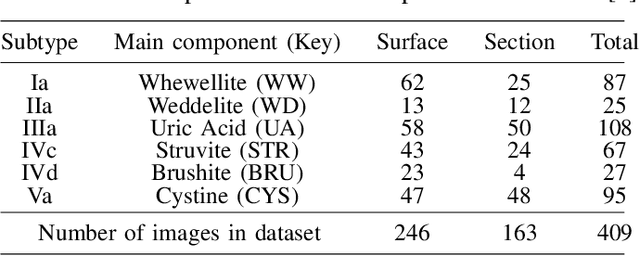
Abstract:Determining the type of kidney stones is crucial for prescribing appropriate treatments to prevent recurrence. Currently, various approaches exist to identify the type of kidney stones. However, obtaining results through the reference ex vivo identification procedure can take several weeks, while in vivo visual recognition requires highly trained specialists. For this reason, deep learning models have been developed to provide urologists with an automated classification of kidney stones during ureteroscopies. Nevertheless, a common issue with these models is the lack of training data. This contribution presents a deep learning method based on few-shot learning, aimed at producing sufficiently discriminative features for identifying kidney stone types in endoscopic images, even with a very limited number of samples. This approach was specifically designed for scenarios where endoscopic images are scarce or where uncommon classes are present, enabling classification even with a limited training dataset. The results demonstrate that Prototypical Networks, using up to 25% of the training data, can achieve performance equal to or better than traditional deep learning models trained with the complete dataset.
Assessing the generalization performance of SAM for ureteroscopy scene understanding
May 22, 2025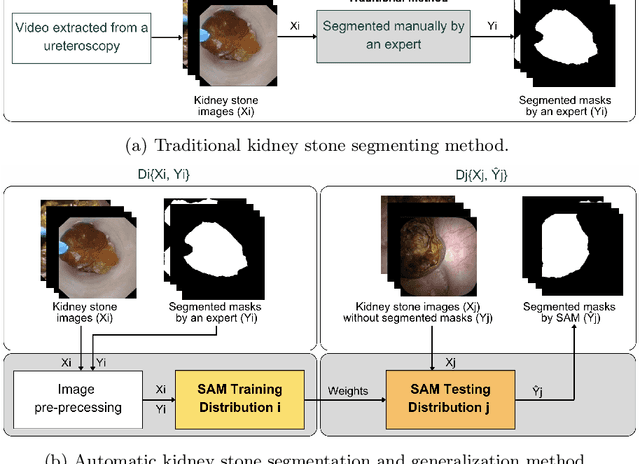
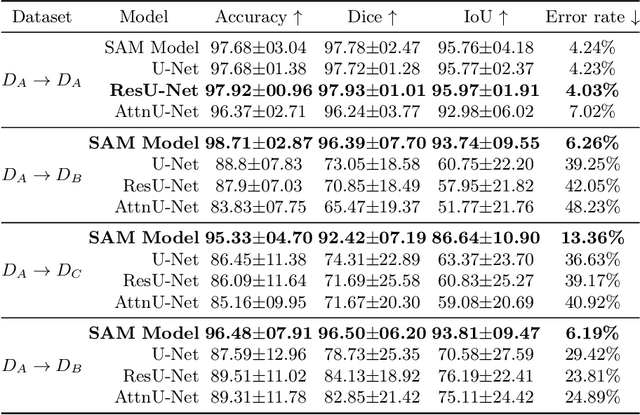
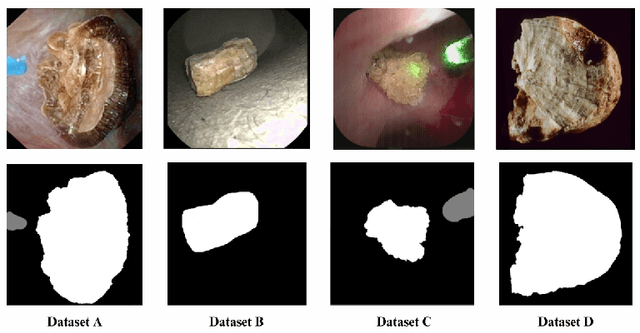
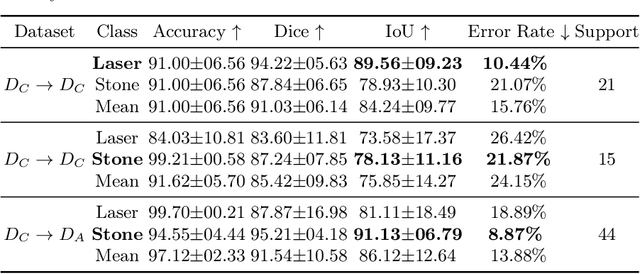
Abstract:The segmentation of kidney stones is regarded as a critical preliminary step to enable the identification of urinary stone types through machine- or deep-learning-based approaches. In urology, manual segmentation is considered tedious and impractical due to the typically large scale of image databases and the continuous generation of new data. In this study, the potential of the Segment Anything Model (SAM) -- a state-of-the-art deep learning framework -- is investigated for the automation of kidney stone segmentation. The performance of SAM is evaluated in comparison to traditional models, including U-Net, Residual U-Net, and Attention U-Net, which, despite their efficiency, frequently exhibit limitations in generalizing to unseen datasets. The findings highlight SAM's superior adaptability and efficiency. While SAM achieves comparable performance to U-Net on in-distribution data (Accuracy: 97.68 + 3.04; Dice: 97.78 + 2.47; IoU: 95.76 + 4.18), it demonstrates significantly enhanced generalization capabilities on out-of-distribution data, surpassing all U-Net variants by margins of up to 23 percent.
Tackling domain generalization for out-of-distribution endoscopic imaging
Oct 18, 2024Abstract:While recent advances in deep learning (DL) for surgical scene segmentation have yielded promising results on single-center and single-imaging modality data, these methods usually do not generalize well to unseen distributions or modalities. Even though human experts can identify visual appearances, DL methods often fail to do so when data samples do not follow a similar distribution. Current literature addressing domain gaps in modality changes has focused primarily on natural scene data. However, these methods cannot be directly applied to endoscopic data, as visual cues in such data are more limited compared to natural scenes. In this work, we exploit both style and content information in images by performing instance normalization and feature covariance mapping techniques to preserve robust and generalizable feature representations. Additionally, to avoid the risk of removing salient feature representations associated with objects of interest, we introduce a restitution module within the feature-learning ResNet backbone that retains useful task-relevant features. Our proposed method shows a 13.7% improvement over the baseline DeepLabv3+ and nearly an 8% improvement over recent state-of-the-art (SOTA) methods for the target (different modality) set of the EndoUDA polyp dataset. Similarly, our method achieved a 19% improvement over the baseline and 6% over the best-performing SOTA method on the EndoUDA Barrett's esophagus (BE) dataset.
EndoDepth: A Benchmark for Assessing Robustness in Endoscopic Depth Prediction
Sep 30, 2024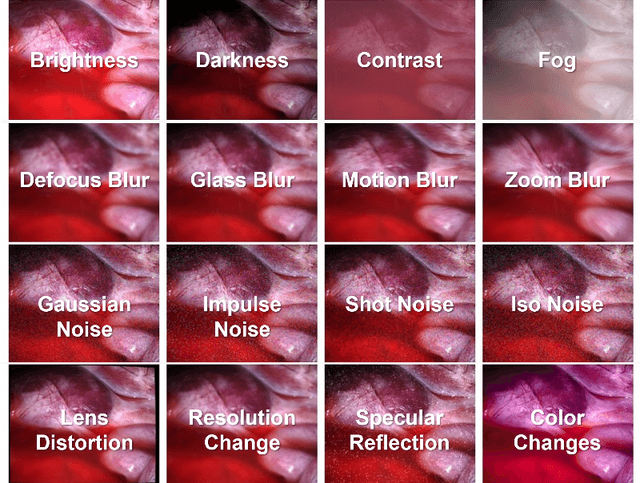
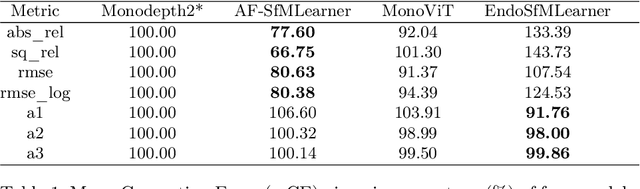
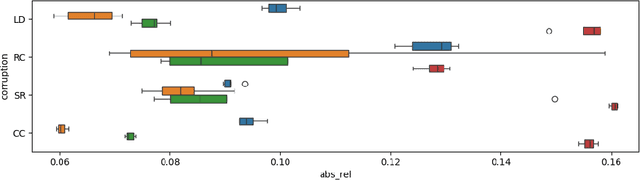

Abstract:Accurate depth estimation in endoscopy is vital for successfully implementing computer vision pipelines for various medical procedures and CAD tools. In this paper, we present the EndoDepth benchmark, an evaluation framework designed to assess the robustness of monocular depth prediction models in endoscopic scenarios. Unlike traditional datasets, the EndoDepth benchmark incorporates common challenges encountered during endoscopic procedures. We present an evaluation approach that is consistent and specifically designed to evaluate the robustness performance of the model in endoscopic scenarios. Among these is a novel composite metric called the mean Depth Estimation Robustness Score (mDERS), which offers an in-depth evaluation of a model's accuracy against errors brought on by endoscopic image corruptions. Moreover, we present SCARED-C, a new dataset designed specifically to assess endoscopy robustness. Through extensive experimentation, we evaluate state-of-the-art depth prediction architectures on the EndoDepth benchmark, revealing their strengths and weaknesses in handling endoscopic challenging imaging artifacts. Our results demonstrate the importance of specialized techniques for accurate depth estimation in endoscopy and provide valuable insights for future research directions.
Leveraging Pre-trained Models for Robust Federated Learning for Kidney Stone Type Recognition
Sep 30, 2024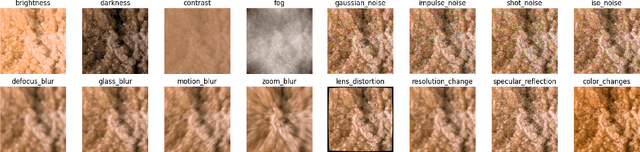
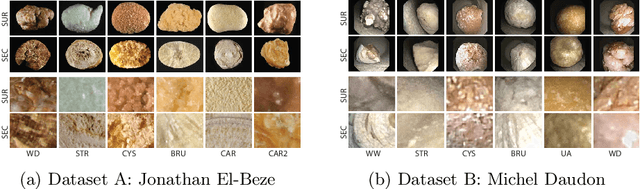
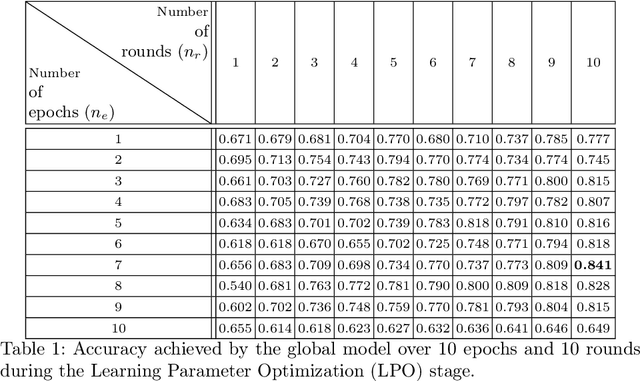
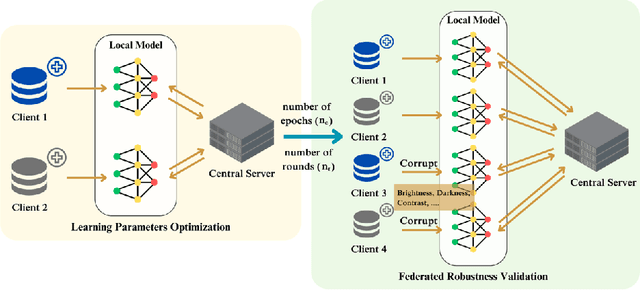

Abstract:Deep learning developments have improved medical imaging diagnoses dramatically, increasing accuracy in several domains. Nonetheless, obstacles continue to exist because of the requirement for huge datasets and legal limitations on data exchange. A solution is provided by Federated Learning (FL), which permits decentralized model training while maintaining data privacy. However, FL models are susceptible to data corruption, which may result in performance degradation. Using pre-trained models, this research suggests a strong FL framework to improve kidney stone diagnosis. Two different kidney stone datasets, each with six different categories of images, are used in our experimental setting. Our method involves two stages: Learning Parameter Optimization (LPO) and Federated Robustness Validation (FRV). We achieved a peak accuracy of 84.1% with seven epochs and 10 rounds during LPO stage, and 77.2% during FRV stage, showing enhanced diagnostic accuracy and robustness against image corruption. This highlights the potential of merging pre-trained models with FL to address privacy and performance concerns in medical diagnostics, and guarantees improved patient care and enhanced trust in FL-based medical systems.
An FPGA smart camera implementation of segmentation models for drone wildfire imagery
Sep 04, 2023Abstract:Wildfires represent one of the most relevant natural disasters worldwide, due to their impact on various societal and environmental levels. Thus, a significant amount of research has been carried out to investigate and apply computer vision techniques to address this problem. One of the most promising approaches for wildfire fighting is the use of drones equipped with visible and infrared cameras for the detection, monitoring, and fire spread assessment in a remote manner but in close proximity to the affected areas. However, implementing effective computer vision algorithms on board is often prohibitive since deploying full-precision deep learning models running on GPU is not a viable option, due to their high power consumption and the limited payload a drone can handle. Thus, in this work, we posit that smart cameras, based on low-power consumption field-programmable gate arrays (FPGAs), in tandem with binarized neural networks (BNNs), represent a cost-effective alternative for implementing onboard computing on the edge. Herein we present the implementation of a segmentation model applied to the Corsican Fire Database. We optimized an existing U-Net model for such a task and ported the model to an edge device (a Xilinx Ultra96-v2 FPGA). By pruning and quantizing the original model, we reduce the number of parameters by 90%. Furthermore, additional optimizations enabled us to increase the throughput of the original model from 8 frames per second (FPS) to 33.63 FPS without loss in the segmentation performance: our model obtained 0.912 in Matthews correlation coefficient (MCC),0.915 in F1 score and 0.870 in Hafiane quality index (HAF), and comparable qualitative segmentation results when contrasted to the original full-precision model. The final model was integrated into a low-cost FPGA, which was used to implement a neural network accelerator.
FAU-Net: An Attention U-Net Extension with Feature Pyramid Attention for Prostate Cancer Segmentation
Sep 04, 2023

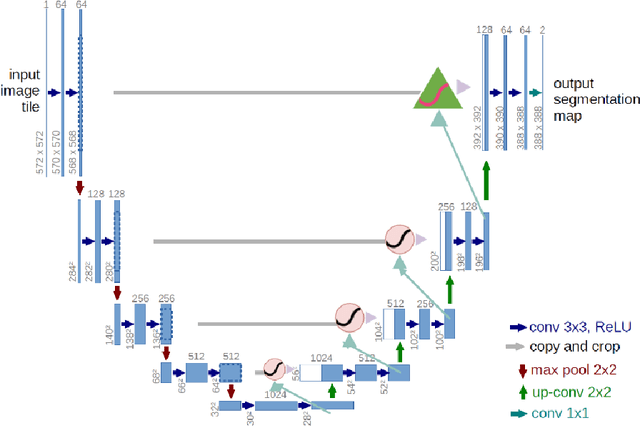

Abstract:This contribution presents a deep learning method for the segmentation of prostate zones in MRI images based on U-Net using additive and feature pyramid attention modules, which can improve the workflow of prostate cancer detection and diagnosis. The proposed model is compared to seven different U-Net-based architectures. The automatic segmentation performance of each model of the central zone (CZ), peripheral zone (PZ), transition zone (TZ) and Tumor were evaluated using Dice Score (DSC), and the Intersection over Union (IoU) metrics. The proposed alternative achieved a mean DSC of 84.15% and IoU of 76.9% in the test set, outperforming most of the studied models in this work except from R2U-Net and attention R2U-Net architectures.
Assessing the performance of deep learning-based models for prostate cancer segmentation using uncertainty scores
Aug 09, 2023



Abstract:This study focuses on comparing deep learning methods for the segmentation and quantification of uncertainty in prostate segmentation from MRI images. The aim is to improve the workflow of prostate cancer detection and diagnosis. Seven different U-Net-based architectures, augmented with Monte-Carlo dropout, are evaluated for automatic segmentation of the central zone, peripheral zone, transition zone, and tumor, with uncertainty estimation. The top-performing model in this study is the Attention R2U-Net, achieving a mean Intersection over Union (IoU) of 76.3% and Dice Similarity Coefficient (DSC) of 85% for segmenting all zones. Additionally, Attention R2U-Net exhibits the lowest uncertainty values, particularly in the boundaries of the transition zone and tumor, when compared to the other models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge