Florian Jug
MamaDino: A Hybrid Vision Model for Breast Cancer 3-Year Risk Prediction
Feb 14, 2026Abstract:Breast cancer screening programmes increasingly seek to move from one-size-fits-all interval to risk-adapted and personalized strategies. Deep learning (DL) has enabled image-based risk models with stronger 1- to 5-year prediction than traditional clinical models, but leading systems (e.g., Mirai) typically use convolutional backbones, very high-resolution inputs (>1M pixels) and simple multi-view fusion, with limited explicit modelling of contralateral asymmetry. We hypothesised that combining complementary inductive biases (convolutional and transformer-based) with explicit contralateral asymmetry modelling would allow us to match state-of-the-art 3-year risk prediction performance even when operating on substantially lower-resolution mammograms, indicating that using less detailed images in a more structured way can recover state-of-the-art accuracy. We present MamaDino, a mammography-aware multi-view attentional DINO model. MamaDino fuses frozen self-supervised DINOv3 ViT-S features with a trainable CNN encoder at 512x512 resolution, and aggregates bilateral breast information via a BilateralMixer to output a 3-year breast cancer risk score. We train on 53,883 women from OPTIMAM (UK) and evaluate on matched 3-year case-control cohorts: an in-distribution test set from four screening sites and an external out-of-distribution cohort from an unseen site. At breast-level, MamaDino matches Mirai on both internal and external tests while using ~13x fewer input pixels. Adding the BilateralMixer improves discrimination to AUC 0.736 (vs 0.713) in-distribution and 0.677 (vs 0.666) out-of-distribution, with consistent performance across age, ethnicity, scanner, tumour type and grade. These findings demonstrate that explicit contralateral modelling and complementary inductive biases enable predictions that match Mirai, despite operating on substantially lower-resolution mammograms.
ResMatching: Noise-Resilient Computational Super-Resolution via Guided Conditional Flow Matching
Oct 30, 2025Abstract:Computational Super-Resolution (CSR) in fluorescence microscopy has, despite being an ill-posed problem, a long history. At its very core, CSR is about finding a prior that can be used to extrapolate frequencies in a micrograph that have never been imaged by the image-generating microscope. It stands to reason that, with the advent of better data-driven machine learning techniques, stronger prior can be learned and hence CSR can lead to better results. Here, we present ResMatching, a novel CSR method that uses guided conditional flow matching to learn such improved data-priors. We evaluate ResMatching on 4 diverse biological structures from the BioSR dataset and compare its results against 7 baselines. ResMatching consistently achieves competitive results, demonstrating in all cases the best trade-off between data fidelity and perceptual realism. We observe that CSR using ResMatching is particularly effective in cases where a strong prior is hard to learn, e.g. when the given low-resolution images contain a lot of noise. Additionally, we show that ResMatching can be used to sample from an implicitly learned posterior distribution and that this distribution is calibrated for all tested use-cases, enabling our method to deliver a pixel-wise data-uncertainty term that can guide future users to reject uncertain predictions.
indiSplit: Bringing Severity Cognizance to Image Decomposition in Fluorescence Microscopy
Mar 29, 2025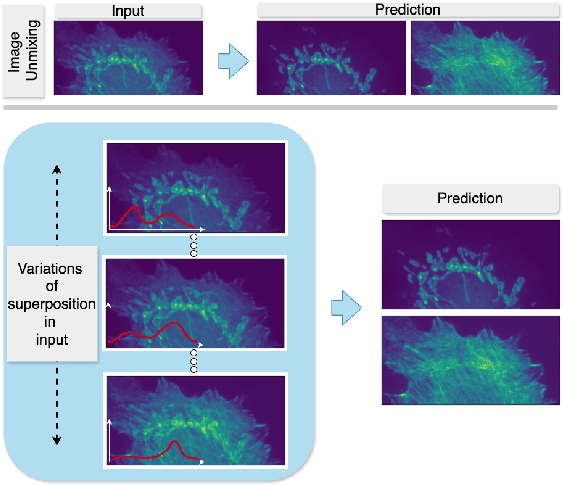
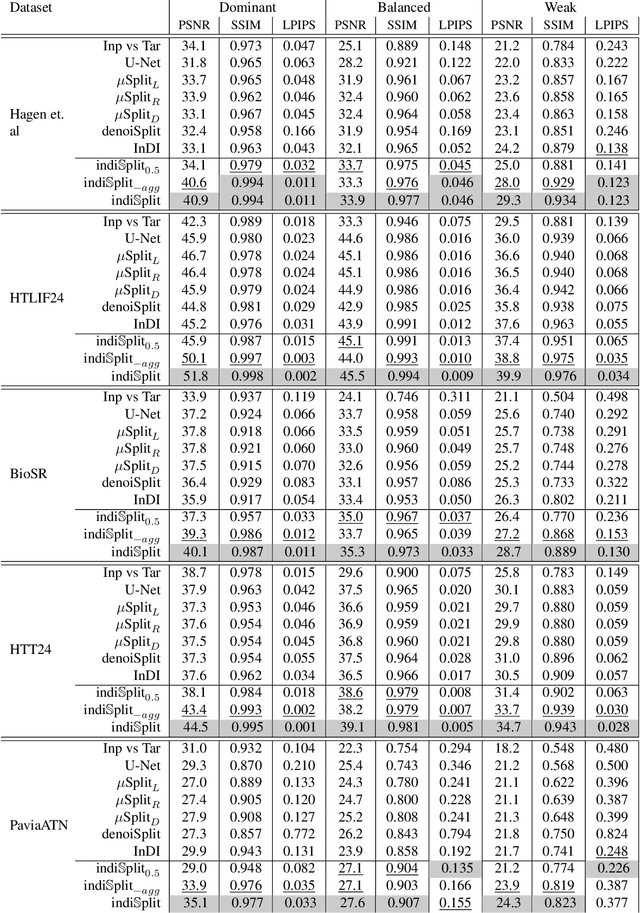
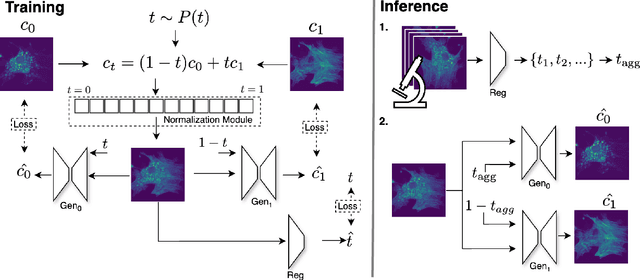
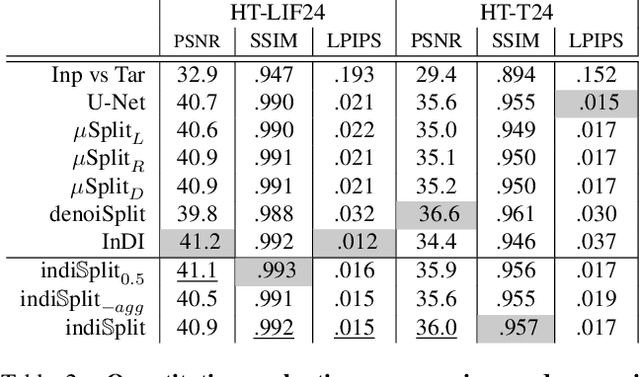
Abstract:Fluorescence microscopy, while being a key driver for progress in the life sciences, is also subject to technical limitations. To overcome them, computational multiplexing techniques have recently been proposed, which allow multiple cellular structures to be captured in a single image and later be unmixed. Existing image decomposition methods are trained on a set of superimposed input images and the respective unmixed target images. It is critical to note that the relative strength (mixing ratio) of the superimposed images for a given input is a priori unknown. However, existing methods are trained on a fixed intensity ratio of superimposed inputs, making them not cognizant to the range of relative intensities that can occur in fluorescence microscopy. In this work, we propose a novel method called indiSplit that is cognizant of the severity of the above mentioned mixing ratio. Our idea is based on InDI, a popular iterative method for image restoration, and an ideal starting point to embrace the unknown mixing ratio in any given input. We introduce (i) a suitably trained regressor network that predicts the degradation level (mixing asymmetry) of a given input image and (ii) a degradation-specific normalization module, enabling degradation-aware inference across all mixing ratios. We show that this method solves two relevant tasks in fluorescence microscopy, namely image splitting and bleedthrough removal, and empirically demonstrate the applicability of indiSplit on $5$ public datasets. We will release all sources under a permissive license.
MicroSSIM: Improved Structural Similarity for Comparing Microscopy Data
Aug 16, 2024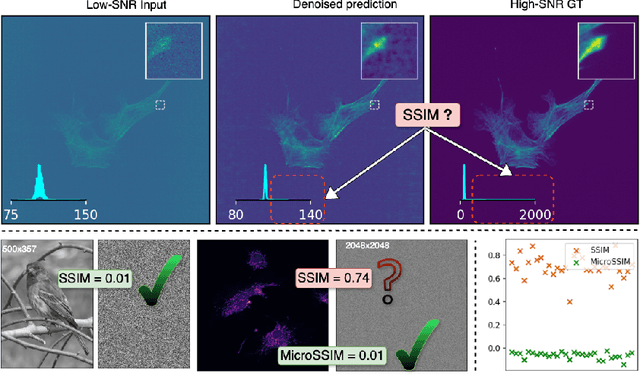

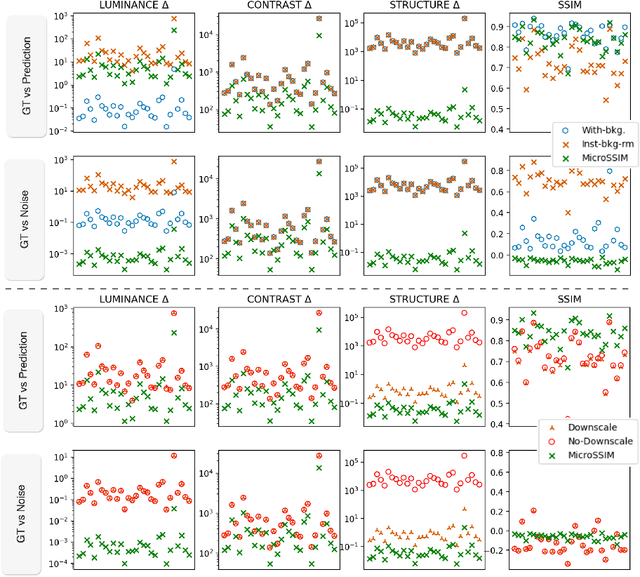
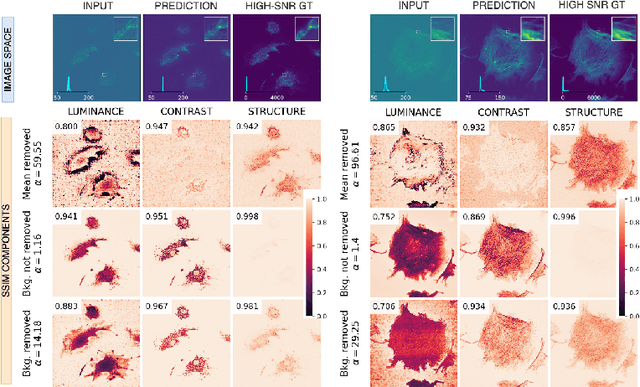
Abstract:Microscopy is routinely used to image biological structures of interest. Due to imaging constraints, acquired images are typically low-SNR and contain noise. Over the last few years, regression-based tasks like unsupervised denoising and splitting have found utility in working with such noisy micrographs. For evaluation, Structural Similarity (SSIM) is one of the most popular measures used in the field. For such tasks, the best evaluation would be when both low-SNR noisy images and corresponding high-SNR clean images are obtained directly from a microscope. However, due to the following three peculiar properties of the microscopy data, we observe that SSIM is not well suited to this data regime: (a) high-SNR micrographs have higher intensity pixels as compared to low SNR micrographs, (b) high-SNR micrographs have higher intensity pixels than found in natural images, images for which SSIM was developed, and (c) a digitally configurable offset is added by the detector present inside the microscope. We show that SSIM components behave unexpectedly when the prediction generated from low-SNR input is compared with the corresponding high-SNR data. We explain this behavior by introducing the phenomenon of saturation, where the value of SSIM components becomes less sensitive to (dis)similarity between the images. We introduce microSSIM, a variant of SSIM, which overcomes the above-discussed issues. We justify the soundness and utility of microSSIM using theoretical and empirical arguments and show the utility of microSSIM on two tasks: unsupervised denoising and joint image splitting with unsupervised denoising. Since our formulation can be applied to a broad family of SSIM-based measures, we also introduce MicroMS3IM, a microscopy-specific variation of MS-SSIM. The source code and python package is available at https://github.com/juglab/MicroSSIM.
denoiSplit: a method for joint image splitting and unsupervised denoising
Mar 25, 2024Abstract:In this work we present denoiSplit, a method to tackle a new analysis task, i.e. the challenge of joint semantic image splitting and unsupervised denoising. This dual approach has important applications in fluorescence microscopy, where semantic image splitting has important applications but noise does generally hinder the downstream analysis of image content. Image splitting involves dissecting an image into its distinguishable semantic structures. We show that the current state-of-the-art method for this task struggles in the presence of image noise, inadvertently also distributing the noise across the predicted outputs. The method we present here can deal with image noise by integrating an unsupervised denoising sub-task. This integration results in improved semantic image unmixing, even in the presence of notable and realistic levels of imaging noise. A key innovation in denoiSplit is the use of specifically formulated noise models and the suitable adjustment of KL-divergence loss for the high-dimensional hierarchical latent space we are training. We showcase the performance of denoiSplit across 4 tasks on real-world microscopy images. Additionally, we perform qualitative and quantitative evaluations and compare results to existing benchmarks, demonstrating the effectiveness of using denoiSplit: a single Variational Splitting Encoder-Decoder (VSE) Network using two suitable noise models to jointly perform semantic splitting and denoising.
MIFA: Metadata, Incentives, Formats, and Accessibility guidelines to improve the reuse of AI datasets for bioimage analysis
Nov 22, 2023



Abstract:Artificial Intelligence methods are powerful tools for biological image analysis and processing. High-quality annotated images are key to training and developing new methods, but access to such data is often hindered by the lack of standards for sharing datasets. We brought together community experts in a workshop to develop guidelines to improve the reuse of bioimages and annotations for AI applications. These include standards on data formats, metadata, data presentation and sharing, and incentives to generate new datasets. We are positive that the MIFA (Metadata, Incentives, Formats, and Accessibility) recommendations will accelerate the development of AI tools for bioimage analysis by facilitating access to high quality training data.
DeepContrast: Deep Tissue Contrast Enhancement using Synthetic Data Degradations and OOD Model Predictions
Aug 16, 2023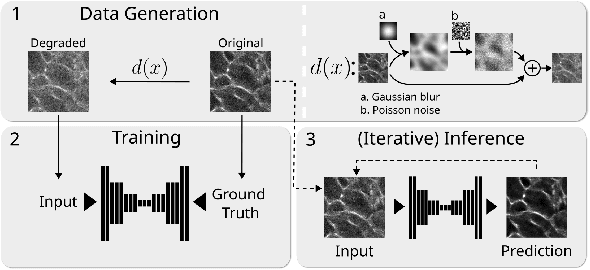
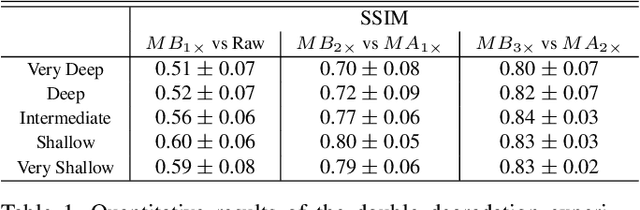
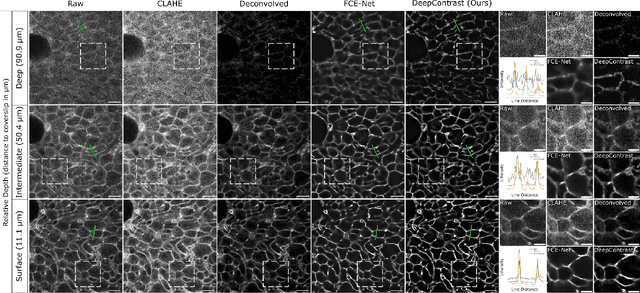
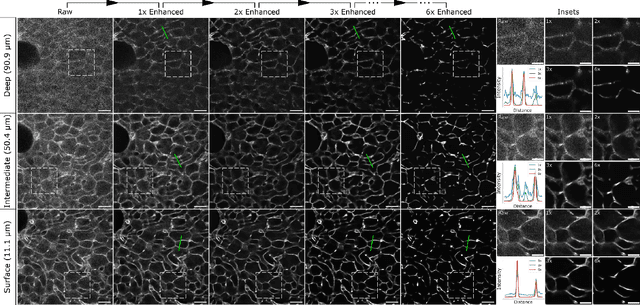
Abstract:Microscopy images are crucial for life science research, allowing detailed inspection and characterization of cellular and tissue-level structures and functions. However, microscopy data are unavoidably affected by image degradations, such as noise, blur, or others. Many such degradations also contribute to a loss of image contrast, which becomes especially pronounced in deeper regions of thick samples. Today, best performing methods to increase the quality of images are based on Deep Learning approaches, which typically require ground truth (GT) data during training. Our inability to counteract blurring and contrast loss when imaging deep into samples prevents the acquisition of such clean GT data. The fact that the forward process of blurring and contrast loss deep into tissue can be modeled, allowed us to propose a new method that can circumvent the problem of unobtainable GT data. To this end, we first synthetically degraded the quality of microscopy images even further by using an approximate forward model for deep tissue image degradations. Then we trained a neural network that learned the inverse of this degradation function from our generated pairs of raw and degraded images. We demonstrated that networks trained in this way can be used out-of-distribution (OOD) to improve the quality of less severely degraded images, e.g. the raw data imaged in a microscope. Since the absolute level of degradation in such microscopy images can be stronger than the additional degradation introduced by our forward model, we also explored the effect of iterative predictions. Here, we observed that in each iteration the measured image contrast kept improving while detailed structures in the images got increasingly removed. Therefore, dependent on the desired downstream analysis, a balance between contrast improvement and retention of image details has to be found.
bAIoimage analysis: elevating the rate of scientific discovery -- as a community
Mar 29, 2023Abstract:The future of bioimage analysis is increasingly defined by the development and use of tools that rely on deep learning and artificial intelligence (AI). For this trend to continue in a way most useful for stimulating scientific progress, it will require our multidisciplinary community to work together, establish FAIR data sharing and deliver usable, reproducible analytical tools.
μSplit: efficient image decomposition for microscopy data
Nov 23, 2022



Abstract:Light microscopy is routinely used to look at living cells and biological tissues at sub-cellular resolution. Components of the imaged cells can be highlighted using fluorescent labels, allowing biologists to investigate individual structures of interest. Given the complexity of biological processes, it is typically necessary to look at multiple structures simultaneously, typically via a temporal multiplexing scheme. Still, imaging more than 3 or 4 structures in this way is difficult for technical reasons and limits the rate of scientific progress in the life sciences. Hence, a computational method to split apart (decompose) superimposed biological structures acquired in a single image channel, i.e. without temporal multiplexing, would have tremendous impact. Here we present {\mu}Split, a dedicated approach for trained image decomposition. We find that best results using regular deep architectures is achieved when large image patches are used during training, making memory consumption the limiting factor to further improving performance. We therefore introduce lateral contextualization (LC), a memory efficient way to train deep networks that operate well on small input patches. In later layers, additional image context is fed at adequately lowered resolution. We integrate LC with Hierarchical Autoencoders and Hierarchical VAEs.For the latter, we also present a modified ELBO loss and show that it enables sound VAE training. We apply {\mu}Split to five decomposition tasks, one on a synthetic dataset, four others derived from two real microscopy datasets. LC consistently achieves SOTA results, while simultaneously requiring considerably less GPU memory than competing architectures not using LC. When introducing LC, results obtained with the above-mentioned vanilla architectures do on average improve by 2.36 dB (PSNR decibel), with individual improvements ranging from 0.9 to 3.4 dB.
N2V2 -- Fixing Noise2Void Checkerboard Artifacts with Modified Sampling Strategies and a Tweaked Network Architecture
Nov 21, 2022



Abstract:In recent years, neural network based image denoising approaches have revolutionized the analysis of biomedical microscopy data. Self-supervised methods, such as Noise2Void (N2V), are applicable to virtually all noisy datasets, even without dedicated training data being available. Arguably, this facilitated the fast and widespread adoption of N2V throughout the life sciences. Unfortunately, the blind-spot training underlying N2V can lead to rather visible checkerboard artifacts, thereby reducing the quality of final predictions considerably. In this work, we present two modifications to the vanilla N2V setup that both help to reduce the unwanted artifacts considerably. Firstly, we propose a modified network architecture, i.e., using BlurPool instead of MaxPool layers throughout the used U-Net, rolling back the residual U-Net to a non-residual U-Net, and eliminating the skip connections at the uppermost U-Net level. Additionally, we propose new replacement strategies to determine the pixel intensity values that fill in the elected blind-spot pixels. We validate our modifications on a range of microscopy and natural image data. Based on added synthetic noise from multiple noise types and at varying amplitudes, we show that both proposed modifications push the current state-of-the-art for fully self-supervised image denoising.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge