Florian Bartenschlager
Dataset on Bi- and Multi-Nucleated Tumor Cells in Canine Cutaneous Mast Cell Tumors
Jan 05, 2021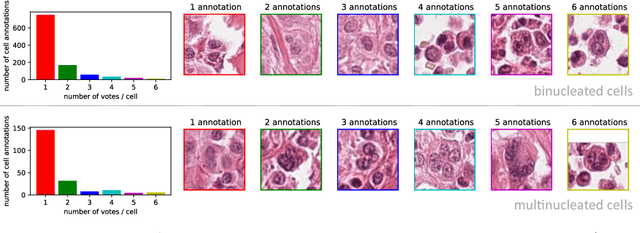
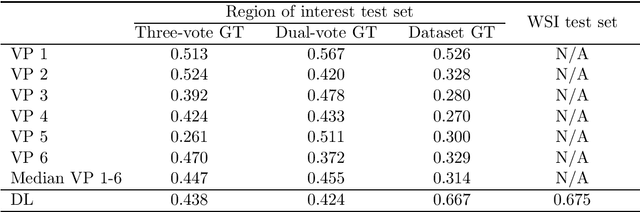
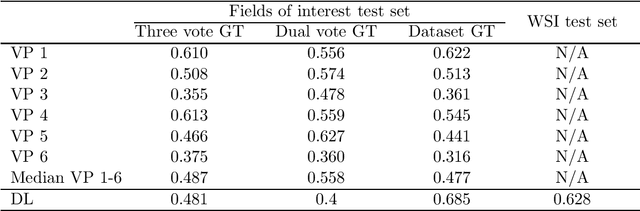
Abstract:Tumor cells with two nuclei (binucleated cells, BiNC) or more nuclei (multinucleated cells, MuNC) indicate an increased amount of cellular genetic material which is thought to facilitate oncogenesis, tumor progression and treatment resistance. In canine cutaneous mast cell tumors (ccMCT), binucleation and multinucleation are parameters used in cytologic and histologic grading schemes (respectively) which correlate with poor patient outcome. For this study, we created the first open source data-set with 19,983 annotations of BiNC and 1,416 annotations of MuNC in 32 histological whole slide images of ccMCT. Labels were created by a pathologist and an algorithmic-aided labeling approach with expert review of each generated candidate. A state-of-the-art deep learning-based model yielded an $F_1$ score of 0.675 for BiNC and 0.623 for MuNC on 11 test whole slide images. In regions of interest ($2.37 mm^2$) extracted from these test images, 6 pathologists had an object detection performance between 0.270 - 0.526 for BiNC and 0.316 - 0.622 for MuNC, while our model archived an $F_1$ score of 0.667 for BiNC and 0.685 for MuNC. This open dataset can facilitate development of automated image analysis for this task and may thereby help to promote standardization of this facet of histologic tumor prognostication.
Deep Learning-Based Quantification of Pulmonary Hemosiderophages in Cytology Slides
Aug 12, 2019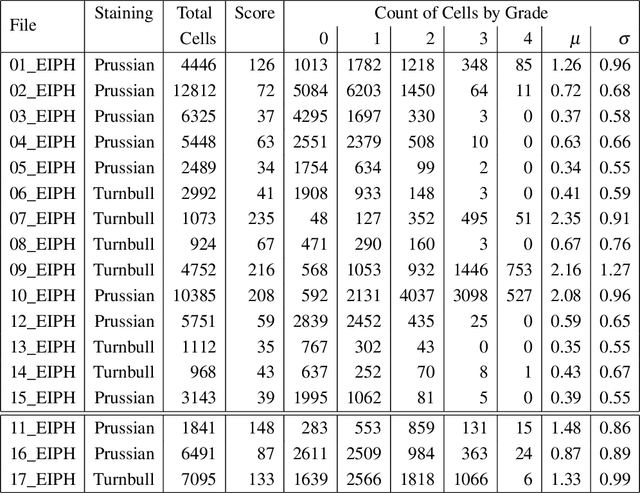
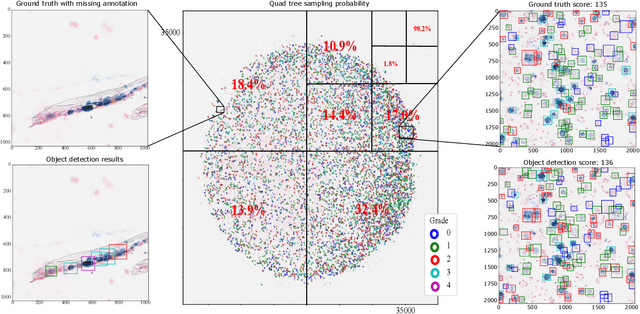
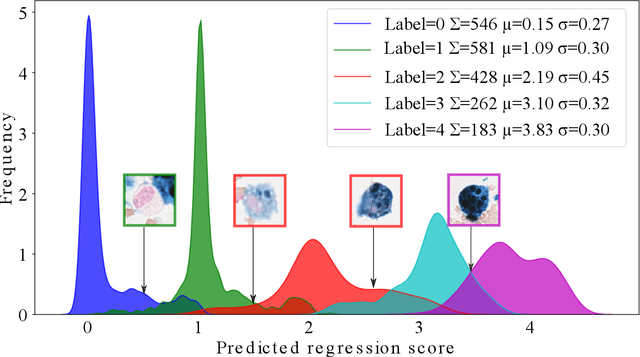
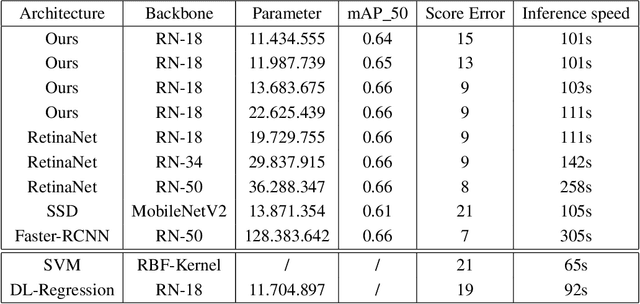
Abstract:Purpose: Exercise-induced pulmonary hemorrhage (EIPH) is a common syndrome in sport horses with negative impact on performance. Cytology of bronchoalveolar lavage fluid by use of a scoring system is considered the most sensitive diagnostic method. Macrophages are classified depending on the degree of cytoplasmic hemosiderin content. The current gold standard is manual grading, which is however monotonous and time-consuming. Methods: We evaluated state-of-the-art deep learning-based methods for single cell macrophage classification and compared them against the performance of nine cytology experts and evaluated inter- and intra-observer variability. Additionally, we evaluated object detection methods on a novel data set of 17 completely annotated cytology whole slide images (WSI) containing 78,047 hemosiderophages. Resultsf: Our deep learning-based approach reached a concordance of 0.85, partially exceeding human expert concordance (0.68 to 0.86, $\mu$=0.73, $\sigma$ =0.04). Intra-observer variability was high (0.68 to 0.88) and inter-observer concordance was moderate (Fleiss kappa = 0.67). Our object detection approach has a mean average precision of 0.66 over the five classes from the whole slide gigapixel image and a computation time of below two minutes. Conclusion: To mitigate the high inter- and intra-rater variability, we propose our automated object detection pipeline, enabling accurate, reproducible and quick EIPH scoring in WSI.
Field of Interest Prediction for Computer-Aided Mitotic Count
Feb 12, 2019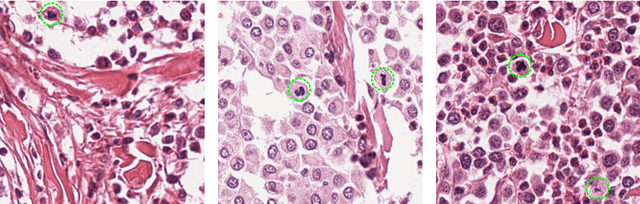
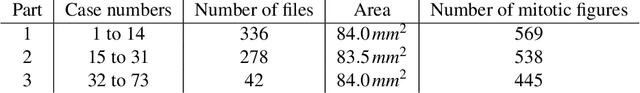
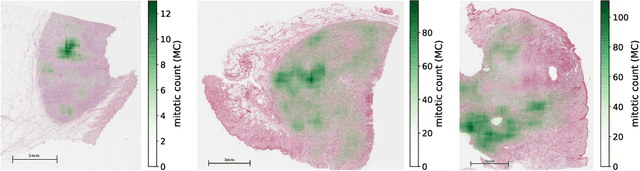
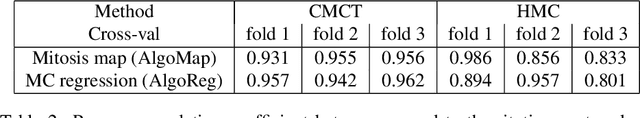
Abstract:Manual counts of mitotic figures, which are determined in the tumor region with the highest mitotic activity, are a key parameter of most tumor grading schemes. It is however strongly dependent on the area selection. To reduce potential variability of prognosis due to this, we propose to use an algorithmic field of interest prediction to assess the area of highest mitotic activity in a whole-slide image. Methods: We evaluated two state-of-the-art methods, all based on the use of deep convolutional neural networks on their ability to predict the mitotic count in digital histopathology slides. We evaluated them on a novel dataset of 32 completely annotated whole slide images from canine cutaneous mast cell tumors (CMCT) and one publicly available human mamma carcinoma (HMC) dataset. We first compared the mitotic counts (MC) predicted by the two models with the ground truth MC on both data sets. Second, for the CMCT data set, we compared the computationally predicted position and MC of the area of highest mitotic activity with size-equivalent areas selected by eight veterinary pathologists. Results: We found a high correlation between the mitotic count as predicted by the models (Pearson's correlation coefficient between 0.931 and 0.962 for the CMCT data set and between 0.801 and 0.986 for the HMC data set) on the slides. For the CMCT data set, this is also reflected in the predicted position representing mitotic counts in mostly the upper quartile of the slide's ground truth MC distribution. Further, we found strong differences between experts in position selection. Conclusion: While the mitotic counts in areas selected by the experts substantially varied, both algorithmic approaches were consistently able to generate a good estimate of the area of highest mitotic count. To achieve better inter-rater agreement, we propose to use computer-based area selection for manual mitotic count.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge