Dietrich Rebholz-Schuhmann
From Large Language Models to Knowledge Graphs for Biomarker Discovery in Cancer
Oct 12, 2023Abstract:Domain experts often rely on up-to-date knowledge for apprehending and disseminating specific biological processes that help them design strategies to develop prevention and therapeutic decision-making. A challenging scenario for artificial intelligence (AI) is using biomedical data (e.g., texts, imaging, omics, and clinical) to provide diagnosis and treatment recommendations for cancerous conditions. Data and knowledge about cancer, drugs, genes, proteins, and their mechanism is spread across structured (knowledge bases (KBs)) and unstructured (e.g., scientific articles) sources. A large-scale knowledge graph (KG) can be constructed by integrating these data, followed by extracting facts about semantically interrelated entities and relations. Such KGs not only allow exploration and question answering (QA) but also allow domain experts to deduce new knowledge. However, exploring and querying large-scale KGs is tedious for non-domain users due to a lack of understanding of the underlying data assets and semantic technologies. In this paper, we develop a domain KG to leverage cancer-specific biomarker discovery and interactive QA. For this, a domain ontology called OncoNet Ontology (ONO) is developed to enable semantic reasoning for validating gene-disease relations. The KG is then enriched by harmonizing the ONO, controlled vocabularies, and additional biomedical concepts from scientific articles by employing BioBERT- and SciBERT-based information extraction (IE) methods. Further, since the biomedical domain is evolving, where new findings often replace old ones, without employing up-to-date findings, there is a high chance an AI system exhibits concept drift while providing diagnosis and treatment. Therefore, we finetuned the KG using large language models (LLMs) based on more recent articles and KBs that might not have been seen by the named entity recognition models.
A Biomedical Knowledge Graph for Biomarker Discovery in Cancer
Feb 23, 2023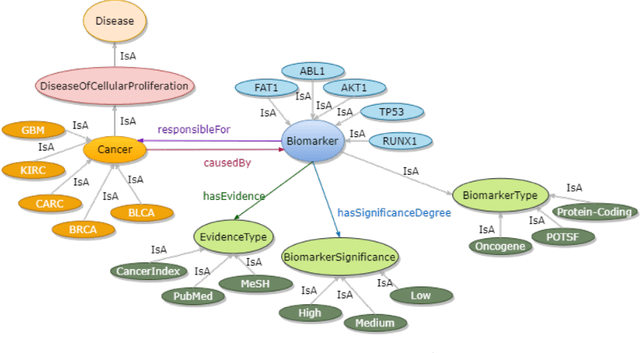
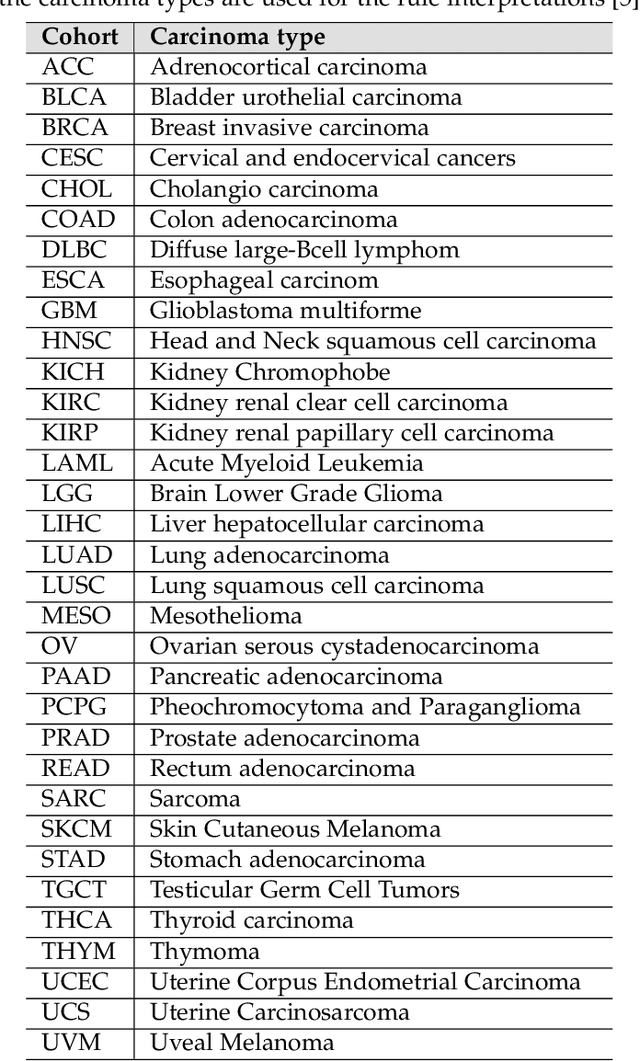
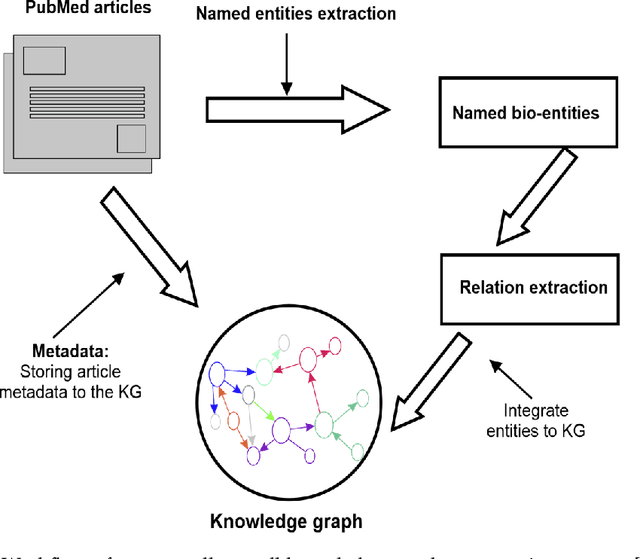

Abstract:Structured and unstructured data and facts about drugs, genes, protein, viruses, and their mechanism are spread across a huge number of scientific articles. These articles are a large-scale knowledge source and can have a huge impact on disseminating knowledge about the mechanisms of certain biological processes. A domain-specific knowledge graph~(KG) is an explicit conceptualization of a specific subject-matter domain represented w.r.t semantically interrelated entities and relations. A KG can be constructed by integrating such facts and data and be used for data integration, exploration, and federated queries. However, exploration and querying large-scale KGs is tedious for certain groups of users due to a lack of knowledge about underlying data assets or semantic technologies. Such a KG will not only allow deducing new knowledge and question answering(QA) but also allows domain experts to explore. Since cross-disciplinary explanations are important for accurate diagnosis, it is important to query the KG to provide interactive explanations about learned biomarkers. Inspired by these, we construct a domain-specific KG, particularly for cancer-specific biomarker discovery. The KG is constructed by integrating cancer-related knowledge and facts from multiple sources. First, we construct a domain-specific ontology, which we call OncoNet Ontology (ONO). The ONO ontology is developed to enable semantic reasoning for verification of the predictions for relations between diseases and genes. The KG is then developed and enriched by harmonizing the ONO, additional metadata schemas, ontologies, controlled vocabularies, and additional concepts from external sources using a BERT-based information extraction method. BioBERT and SciBERT are finetuned with the selected articles crawled from PubMed. We listed down some queries and some examples of QA and deducing knowledge based on the KG.
Explainable AI for Bioinformatics: Methods, Tools, and Applications
Dec 25, 2022Abstract:Artificial intelligence(AI) systems based on deep neural networks (DNNs) and machine learning (ML) algorithms are increasingly used to solve critical problems in bioinformatics, biomedical informatics, and precision medicine. However, complex DNN or ML models that are unavoidably opaque and perceived as black-box methods, may not be able to explain why and how they make certain decisions. Such black-box models are difficult to comprehend not only for targeted users and decision-makers but also for AI developers. Besides, in sensitive areas like healthcare, explainability and accountability are not only desirable properties of AI but also legal requirements -- especially when AI may have significant impacts on human lives. Explainable artificial intelligence (XAI) is an emerging field that aims to mitigate the opaqueness of black-box models and make it possible to interpret how AI systems make their decisions with transparency. An interpretable ML model can explain how it makes predictions and which factors affect the model's outcomes. The majority of state-of-the-art interpretable ML methods have been developed in a domain-agnostic way and originate from computer vision, automated reasoning, or even statistics. Many of these methods cannot be directly applied to bioinformatics problems, without prior customization, extension, and domain adoption. In this paper, we discuss the importance of explainability with a focus on bioinformatics. We analyse and comprehensively overview of model-specific and model-agnostic interpretable ML methods and tools. Via several case studies covering bioimaging, cancer genomics, and biomedical text mining, we show how bioinformatics research could benefit from XAI methods and how they could help improve decision fairness.
DeepCOVIDExplainer: Explainable COVID-19 Predictions Based on Chest X-ray Images
Apr 10, 2020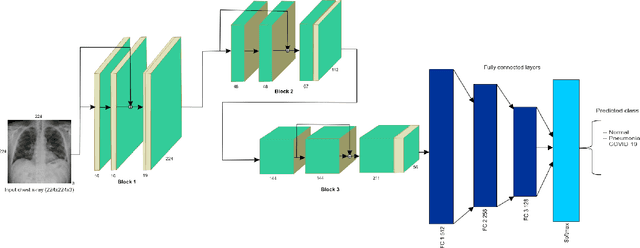
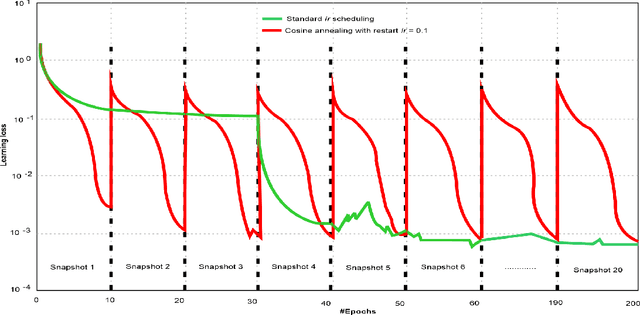
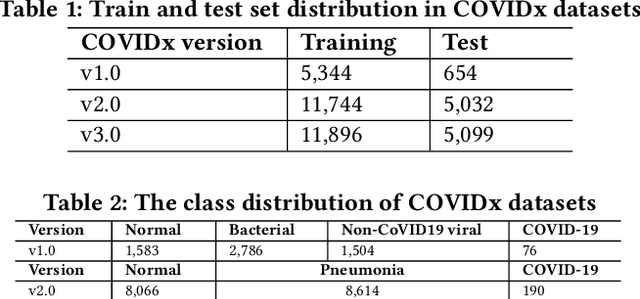
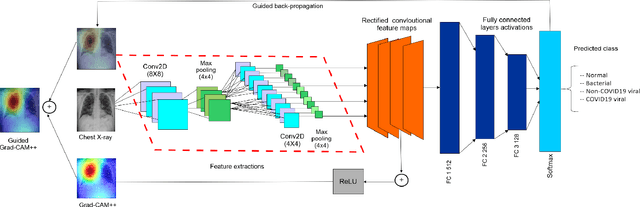
Abstract:Amid the coronavirus disease(COVID-19) pandemic, humanity experiences a rapid increase in infection numbers across the world. Challenge hospitals are faced with, in the fight against the virus, is the effective screening of incoming patients. One methodology is the assessment of chest radiography(CXR) images, which usually requires expert radiologists' knowledge. In this paper, we propose an explainable deep neural networks(DNN)-based method for automatic detection of COVID-19 symptoms from CXR images, which we call 'DeepCOVIDExplainer'. We used 16,995 CXR images across 13,808 patients, covering normal, pneumonia, and COVID-19 cases. CXR images are first comprehensively preprocessed, before being augmented and classified with a neural ensemble method, followed by highlighting class-discriminating regions using gradient-guided class activation maps(Grad-CAM++) and layer-wise relevance propagation(LRP). Further, we provide human-interpretable explanations of the predictions. Evaluation results based on hold-out data show that our approach can identify COVID-19 confidently with a positive predictive value(PPV) of 89.61% and recall of 83%, improving over recent comparable approaches. We hope that our findings will be a useful contribution to the fight against COVID-19 and, in more general, towards an increasing acceptance and adoption of AI-assisted applications in the clinical practice.
Predicting knee osteoarthritis severity: comparative modeling based on patient's data and plain X-ray images
Aug 23, 2019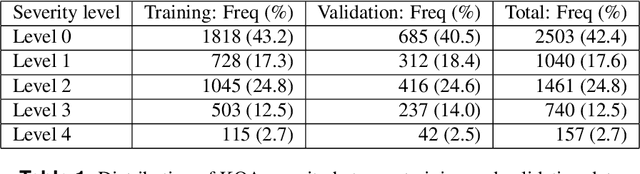
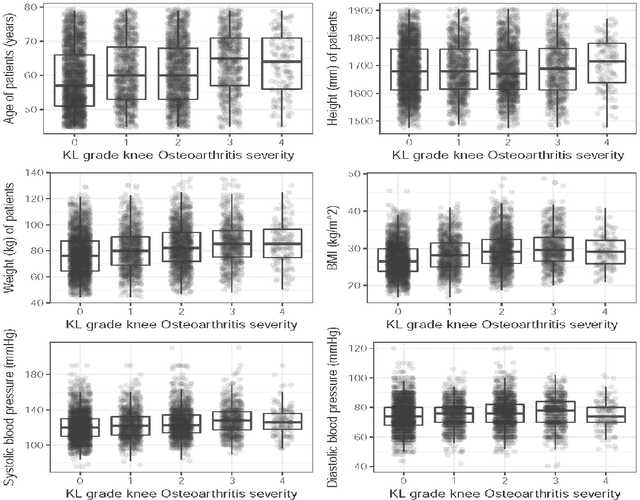
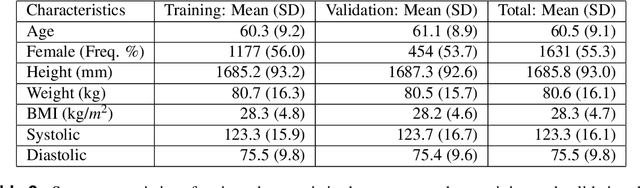

Abstract:Knee osteoarthritis (KOA) is a disease that impairs knee function and causes pain. A radiologist reviews knee X-ray images and grades the severity level of the impairments according to the Kellgren and Lawrence grading scheme; a five-point ordinal scale (0--4). In this study, we used Elastic Net (EN) and Random Forests (RF) to build predictive models using patient assessment data (i.e. signs and symptoms of both knees and medication use) and a convolution neural network (CNN) trained using X-ray images only. Linear mixed effect models (LMM) were used to model the within subject correlation between the two knees. The root mean squared error for the CNN, EN, and RF models was 0.77, 0.97, and 0.94 respectively. The LMM shows similar overall prediction accuracy as the EN regression but correctly accounted for the hierarchical structure of the data resulting in more reliable inference. Useful explanatory variables were identified that could be used for patient monitoring before X-ray imaging. Our analyses suggest that the models trained for predicting the KOA severity levels achieve comparable results when modeling X-ray images and patient data. The subjectivity in the KL grade is still a primary concern.
* Published in Nature Scientific Reports, 2019
First steps in the logic-based assessment of post-composed phenotypic descriptions
Dec 08, 2010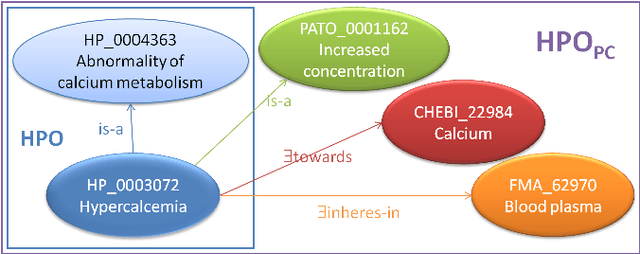
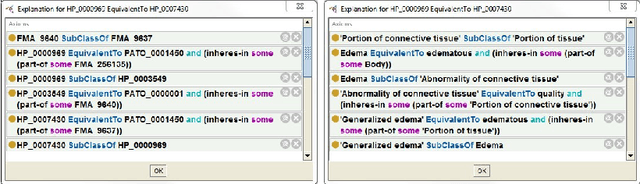
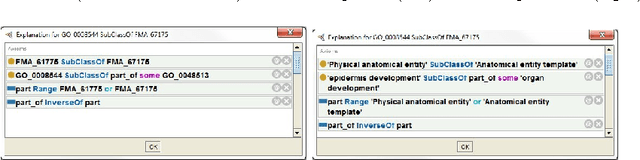
Abstract:In this paper we present a preliminary logic-based evaluation of the integration of post-composed phenotypic descriptions with domain ontologies. The evaluation has been performed using a description logic reasoner together with scalable techniques: ontology modularization and approximations of the logical difference between ontologies.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge