Daniel J. Mollura
CT-Realistic Lung Nodule Simulation from 3D Conditional Generative Adversarial Networks for Robust Lung Segmentation
Jun 11, 2018
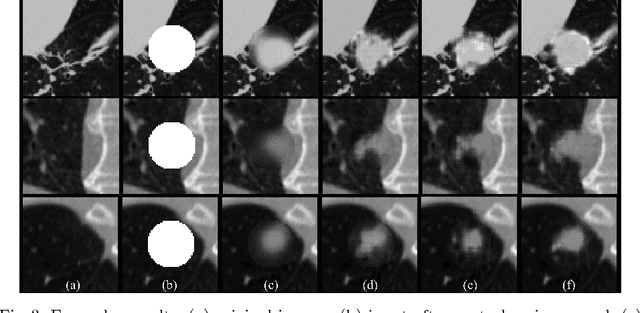
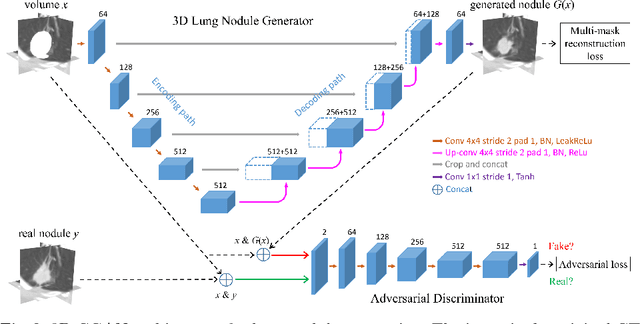


Abstract:Data availability plays a critical role for the performance of deep learning systems. This challenge is especially acute within the medical image domain, particularly when pathologies are involved, due to two factors: 1) limited number of cases, and 2) large variations in location, scale, and appearance. In this work, we investigate whether augmenting a dataset with artificially generated lung nodules can improve the robustness of the progressive holistically nested network (P-HNN) model for pathological lung segmentation of CT scans. To achieve this goal, we develop a 3D generative adversarial network (GAN) that effectively learns lung nodule property distributions in 3D space. In order to embed the nodules within their background context, we condition the GAN based on a volume of interest whose central part containing the nodule has been erased. To further improve realism and blending with the background, we propose a novel multi-mask reconstruction loss. We train our method on over 1000 nodules from the LIDC dataset. Qualitative results demonstrate the effectiveness of our method compared to the state-of-art. We then use our GAN to generate simulated training images where nodules lie on the lung border, which are cases where the published P-HNN model struggles. Qualitative and quantitative results demonstrate that armed with these simulated images, the P-HNN model learns to better segment lung regions under these challenging situations. As a result, our system provides a promising means to help overcome the data paucity that commonly afflicts medical imaging.
White matter hyperintensity segmentation from T1 and FLAIR images using fully convolutional neural networks enhanced with residual connections
Mar 19, 2018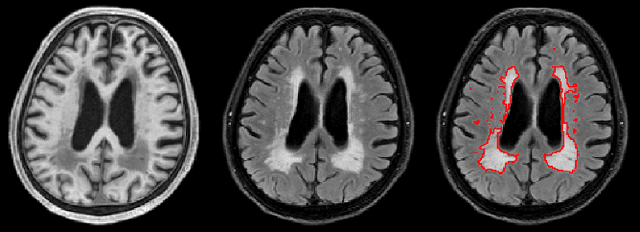
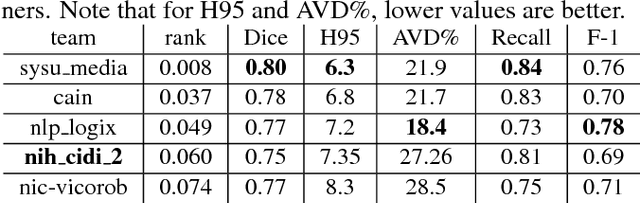

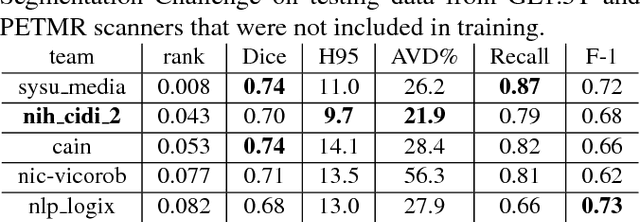
Abstract:Segmentation and quantification of white matter hyperintensities (WMHs) are of great importance in studying and understanding various neurological and geriatric disorders. Although automatic methods have been proposed for WMH segmentation on magnetic resonance imaging (MRI), manual corrections are often necessary to achieve clinically practical results. Major challenges for WMH segmentation stem from their inhomogeneous MRI intensities, random location and size distributions, and MRI noise. The presence of other brain anatomies or diseases with enhanced intensities adds further difficulties. To cope with these challenges, we present a specifically designed fully convolutional neural network (FCN) with residual connections to segment WMHs by using combined T1 and fluid-attenuated inversion recovery (FLAIR) images. Our customized FCN is designed to be straightforward and generalizable, providing efficient end-to-end training due to its enhanced information propagation. We tested our method on the open WMH Segmentation Challenge MICCAI2017 dataset, and, despite our method's relative simplicity, results show that it performs amongst the leading techniques across five metrics. More importantly, our method achieves the best score for hausdorff distance and average volume difference in testing datasets from two MRI scanners that were not included in training, demonstrating better generalization ability of our proposed method over its competitors.
Pathological Pulmonary Lobe Segmentation from CT Images using Progressive Holistically Nested Neural Networks and Random Walker
Aug 15, 2017



Abstract:Automatic pathological pulmonary lobe segmentation(PPLS) enables regional analyses of lung disease, a clinically important capability. Due to often incomplete lobe boundaries, PPLS is difficult even for experts, and most prior art requires inference from contextual information. To address this, we propose a novel PPLS method that couples deep learning with the random walker (RW) algorithm. We first employ the recent progressive holistically-nested network (P-HNN) model to identify potential lobar boundaries, then generate final segmentations using a RW that is seeded and weighted by the P-HNN output. We are the first to apply deep learning to PPLS. The advantages are independence from prior airway/vessel segmentations, increased robustness in diseased lungs, and methodological simplicity that does not sacrifice accuracy. Our method posts a high mean Jaccard score of 0.888$\pm$0.164 on a held-out set of 154 CT scans from lung-disease patients, while also significantly (p < 0.001) outperforming a state-of-the-art method.
Progressive and Multi-Path Holistically Nested Neural Networks for Pathological Lung Segmentation from CT Images
Jun 12, 2017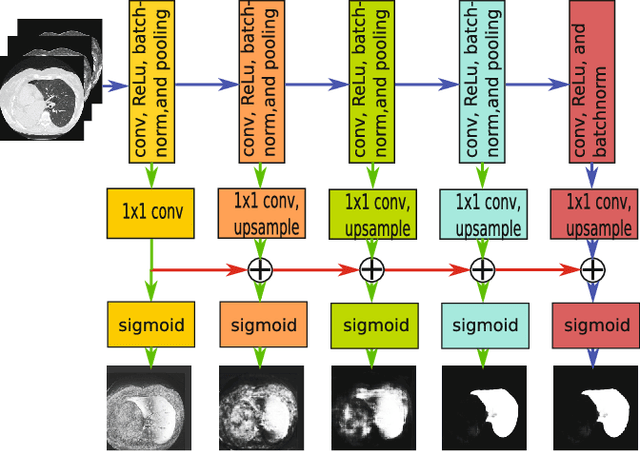

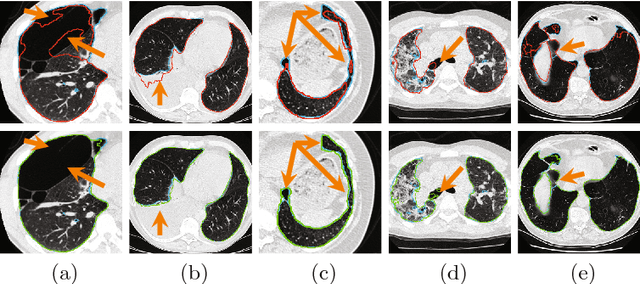
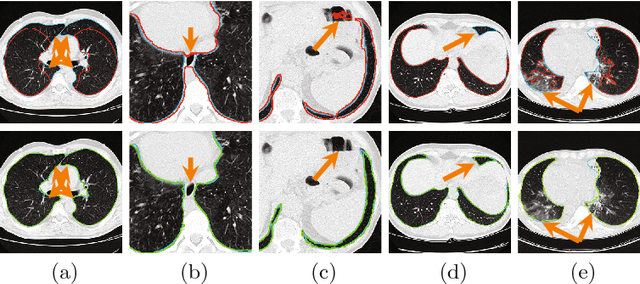
Abstract:Pathological lung segmentation (PLS) is an important, yet challenging, medical image application due to the wide variability of pathological lung appearance and shape. Because PLS is often a pre-requisite for other imaging analytics, methodological simplicity and generality are key factors in usability. Along those lines, we present a bottom-up deep-learning based approach that is expressive enough to handle variations in appearance, while remaining unaffected by any variations in shape. We incorporate the deeply supervised learning framework, but enhance it with a simple, yet effective, progressive multi-path scheme, which more reliably merges outputs from different network stages. The result is a deep model able to produce finer detailed masks, which we call progressive holistically-nested networks (P-HNNs). Using extensive cross-validation, our method is tested on multi-institutional datasets comprising 929 CT scans (848 publicly available), of pathological lungs, reporting mean dice scores of 0.985 and demonstrating significant qualitative and quantitative improvements over state-of-the art approaches.
* 8 Pages, 4 figures, MICCAI 2007
Holistic Interstitial Lung Disease Detection using Deep Convolutional Neural Networks: Multi-label Learning and Unordered Pooling
Jan 19, 2017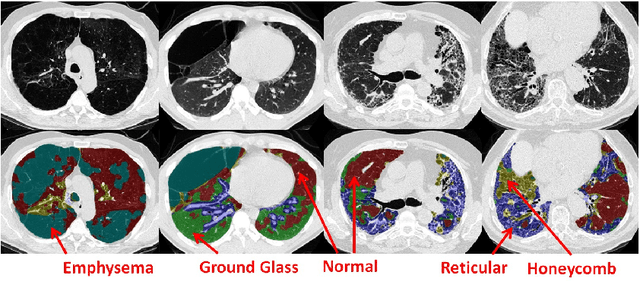
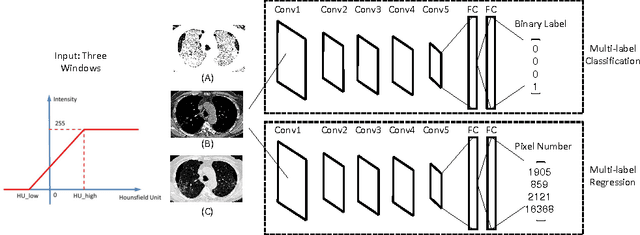
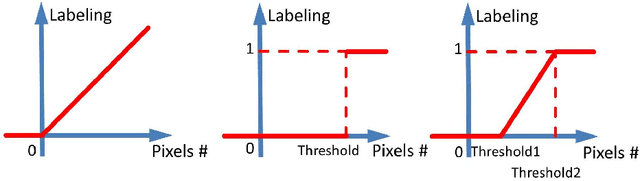
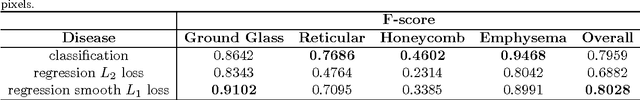
Abstract:Accurately predicting and detecting interstitial lung disease (ILD) patterns given any computed tomography (CT) slice without any pre-processing prerequisites, such as manually delineated regions of interest (ROIs), is a clinically desirable, yet challenging goal. The majority of existing work relies on manually-provided ILD ROIs to extract sampled 2D image patches from CT slices and, from there, performs patch-based ILD categorization. Acquiring manual ROIs is labor intensive and serves as a bottleneck towards fully-automated CT imaging ILD screening over large-scale populations. Furthermore, despite the considerable high frequency of more than one ILD pattern on a single CT slice, previous works are only designed to detect one ILD pattern per slice or patch. To tackle these two critical challenges, we present multi-label deep convolutional neural networks (CNNs) for detecting ILDs from holistic CT slices (instead of ROIs or sub-images). Conventional single-labeled CNN models can be augmented to cope with the possible presence of multiple ILD pattern labels, via 1) continuous-valued deep regression based robust norm loss functions or 2) a categorical objective as the sum of element-wise binary logistic losses. Our methods are evaluated and validated using a publicly available database of 658 patient CT scans under five-fold cross-validation, achieving promising performance on detecting four major ILD patterns: Ground Glass, Reticular, Honeycomb, and Emphysema. We also investigate the effectiveness of a CNN activation-based deep-feature encoding scheme using Fisher vector encoding, which treats ILD detection as spatially-unordered deep texture classification.
Characterization of Lung Nodule Malignancy using Hybrid Shape and Appearance Features
Sep 21, 2016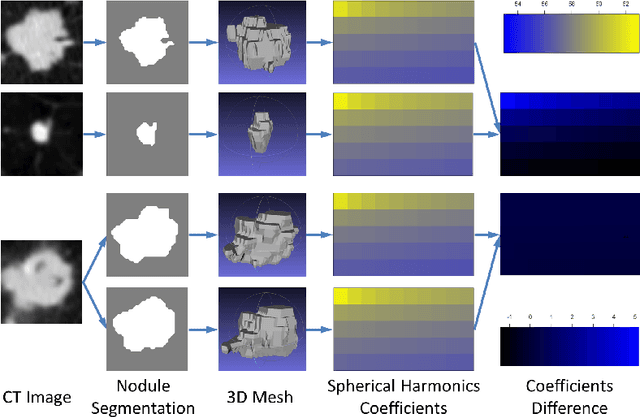


Abstract:Computed tomography imaging is a standard modality for detecting and assessing lung cancer. In order to evaluate the malignancy of lung nodules, clinical practice often involves expert qualitative ratings on several criteria describing a nodule's appearance and shape. Translating these features for computer-aided diagnostics is challenging due to their subjective nature and the difficulties in gaining a complete description. In this paper, we propose a computerized approach to quantitatively evaluate both appearance distinctions and 3D surface variations. Nodule shape was modeled and parameterized using spherical harmonics, and appearance features were extracted using deep convolutional neural networks. Both sets of features were combined to estimate the nodule malignancy using a random forest classifier. The proposed algorithm was tested on the publicly available Lung Image Database Consortium dataset, achieving high accuracy. By providing lung nodule characterization, this method can provide a robust alternative reference opinion for lung cancer diagnosis.
Optimally Stabilized PET Image Denoising Using Trilateral Filtering
Jul 11, 2014

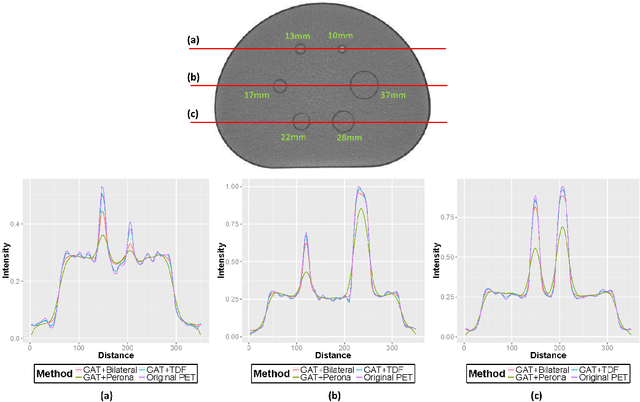

Abstract:Low-resolution and signal-dependent noise distribution in positron emission tomography (PET) images makes denoising process an inevitable step prior to qualitative and quantitative image analysis tasks. Conventional PET denoising methods either over-smooth small-sized structures due to resolution limitation or make incorrect assumptions about the noise characteristics. Therefore, clinically important quantitative information may be corrupted. To address these challenges, we introduced a novel approach to remove signal-dependent noise in the PET images where the noise distribution was considered as Poisson-Gaussian mixed. Meanwhile, the generalized Anscombe's transformation (GAT) was used to stabilize varying nature of the PET noise. Other than noise stabilization, it is also desirable for the noise removal filter to preserve the boundaries of the structures while smoothing the noisy regions. Indeed, it is important to avoid significant loss of quantitative information such as standard uptake value (SUV)-based metrics as well as metabolic lesion volume. To satisfy all these properties, we extended bilateral filtering method into trilateral filtering through multiscaling and optimal Gaussianization process. The proposed method was tested on more than 50 PET-CT images from various patients having different cancers and achieved the superior performance compared to the widely used denoising techniques in the literature.
Near-optimal Keypoint Sampling for Fast Pathological Lung Segmentation
Jul 11, 2014
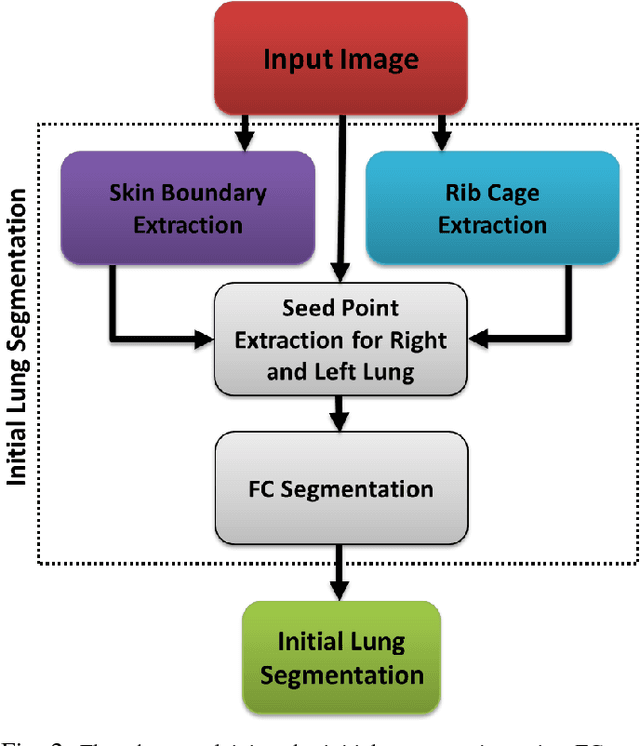
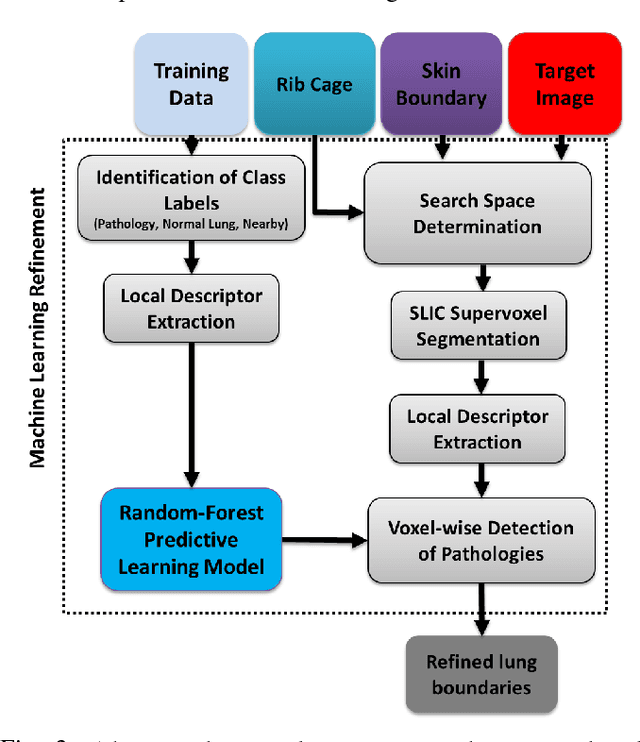
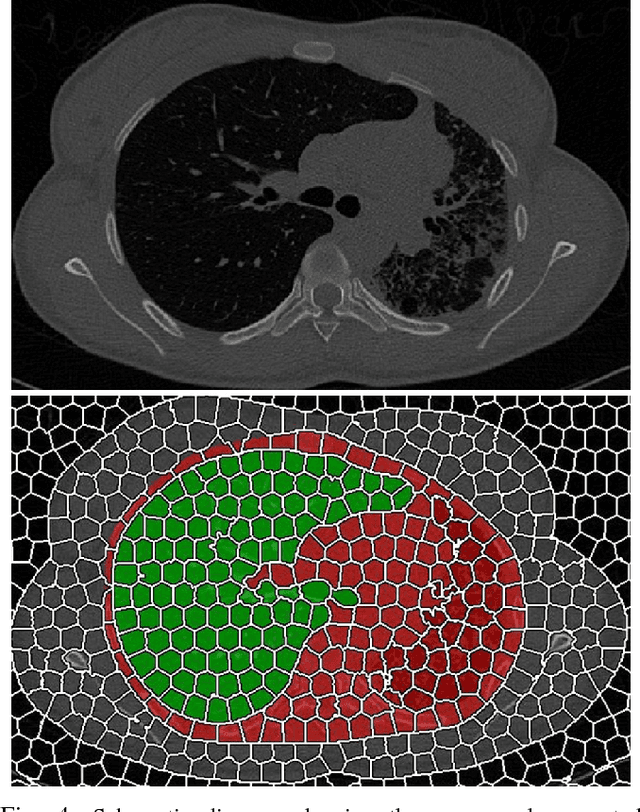
Abstract:Accurate delineation of pathological lungs from computed tomography (CT) images remains mostly unsolved because available methods fail to provide a reliable generic solution due to high variability of abnormality appearance. Local descriptor-based classification methods have shown to work well in annotating pathologies; however, these methods are usually computationally intensive which restricts their widespread use in real-time or near-real-time clinical applications. In this paper, we present a novel approach for fast, accurate, reliable segmentation of pathological lungs from CT scans by combining region-based segmentation method with local descriptor classification that is performed on an optimized sampling grid. Our method works in two stages; during stage one, we adapted the fuzzy connectedness (FC) image segmentation algorithm to perform initial lung parenchyma extraction. In the second stage, texture-based local descriptors are utilized to segment abnormal imaging patterns using a near optimal keypoint analysis by employing centroid of supervoxel as grid points. The quantitative results show that our pathological lung segmentation method is fast, robust, and improves on current standards and has potential to enhance the performance of routine clinical tasks.
CIDI-Lung-Seg: A Single-Click Annotation Tool for Automatic Delineation of Lungs from CT Scans
Jul 11, 2014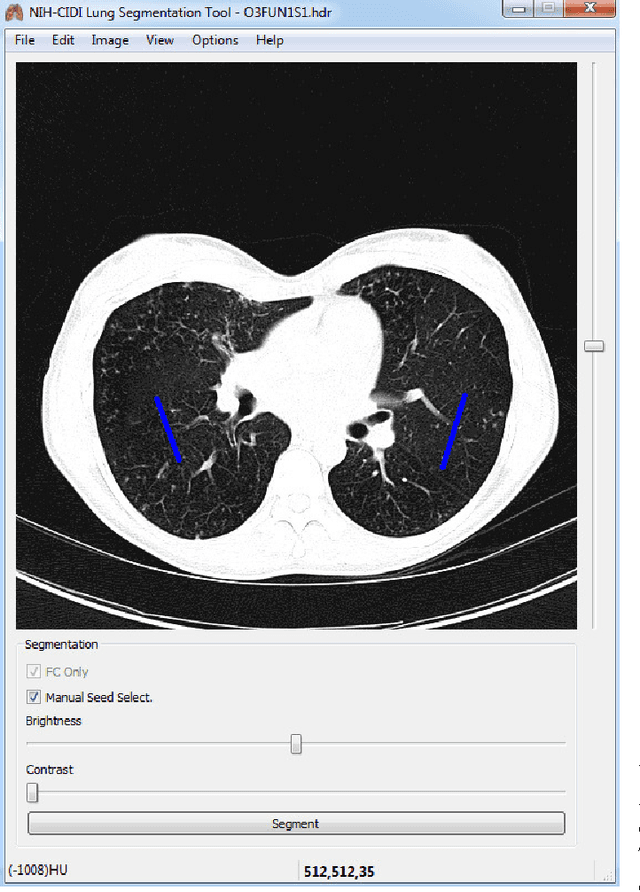
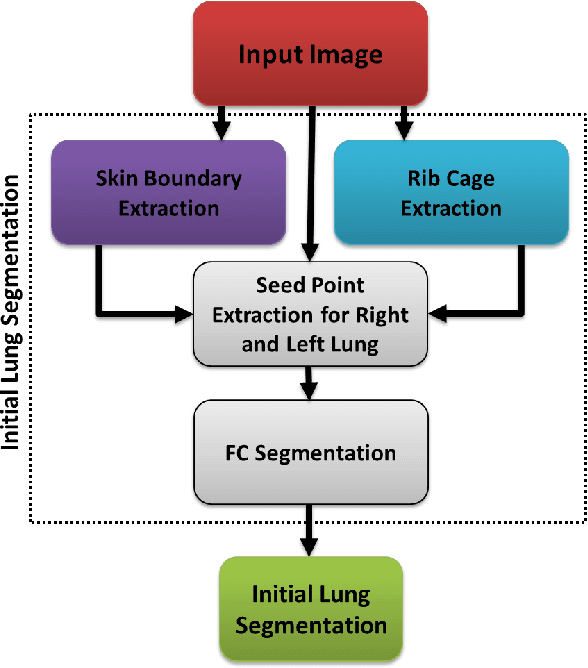

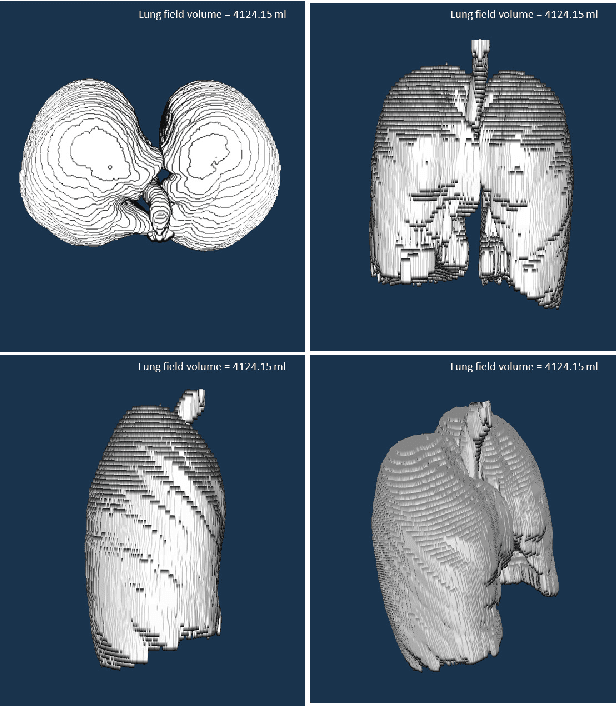
Abstract:Accurate and fast extraction of lung volumes from computed tomography (CT) scans remains in a great demand in the clinical environment because the available methods fail to provide a generic solution due to wide anatomical variations of lungs and existence of pathologies. Manual annotation, current gold standard, is time consuming and often subject to human bias. On the other hand, current state-of-the-art fully automated lung segmentation methods fail to make their way into the clinical practice due to their inability to efficiently incorporate human input for handling misclassifications and praxis. This paper presents a lung annotation tool for CT images that is interactive, efficient, and robust. The proposed annotation tool produces an "as accurate as possible" initial annotation based on the fuzzy-connectedness image segmentation, followed by efficient manual fixation of the initial extraction if deemed necessary by the practitioner. To provide maximum flexibility to the users, our annotation tool is supported in three major operating systems (Windows, Linux, and the Mac OS X). The quantitative results comparing our free software with commercially available lung segmentation tools show higher degree of consistency and precision of our software with a considerable potential to enhance the performance of routine clinical tasks.
Learning Shape and Texture Characteristics of CT Tree-in-Bud Opacities for CAD Systems
Jun 26, 2011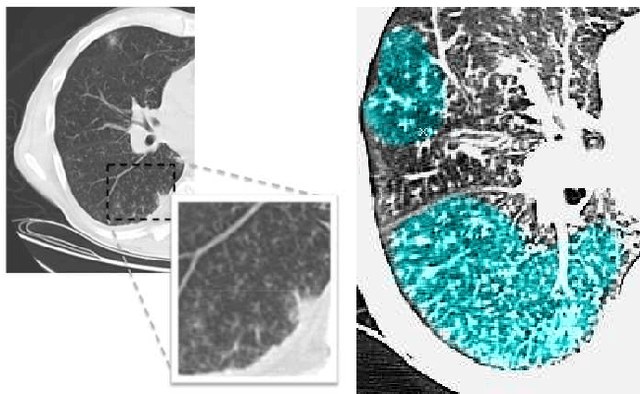
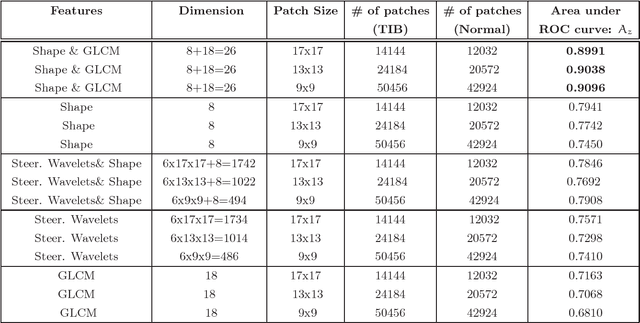
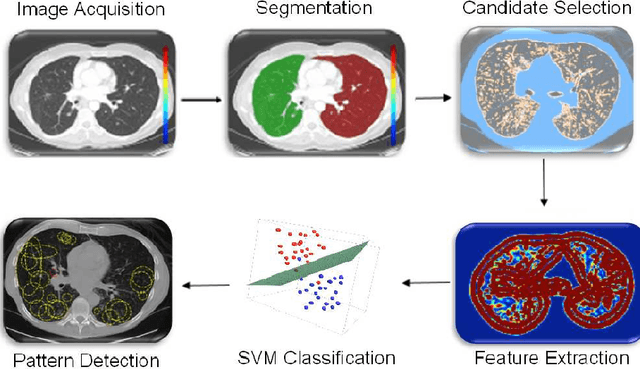
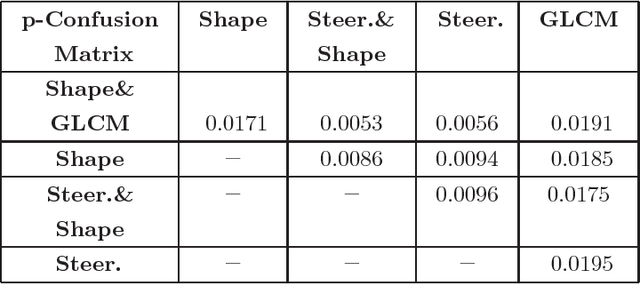
Abstract:Although radiologists can employ CAD systems to characterize malignancies, pulmonary fibrosis and other chronic diseases; the design of imaging techniques to quantify infectious diseases continue to lag behind. There exists a need to create more CAD systems capable of detecting and quantifying characteristic patterns often seen in respiratory tract infections such as influenza, bacterial pneumonia, or tuborculosis. One of such patterns is Tree-in-bud (TIB) which presents \textit{thickened} bronchial structures surrounding by clusters of \textit{micro-nodules}. Automatic detection of TIB patterns is a challenging task because of their weak boundary, noisy appearance, and small lesion size. In this paper, we present two novel methods for automatically detecting TIB patterns: (1) a fast localization of candidate patterns using information from local scale of the images, and (2) a M\"{o}bius invariant feature extraction method based on learned local shape and texture properties. A comparative evaluation of the proposed methods is presented with a dataset of 39 laboratory confirmed viral bronchiolitis human parainfluenza (HPIV) CTs and 21 normal lung CTs. Experimental results demonstrate that the proposed CAD system can achieve high detection rate with an overall accuracy of 90.96%.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge