Kevin George
Pathological Pulmonary Lobe Segmentation from CT Images using Progressive Holistically Nested Neural Networks and Random Walker
Aug 15, 2017



Abstract:Automatic pathological pulmonary lobe segmentation(PPLS) enables regional analyses of lung disease, a clinically important capability. Due to often incomplete lobe boundaries, PPLS is difficult even for experts, and most prior art requires inference from contextual information. To address this, we propose a novel PPLS method that couples deep learning with the random walker (RW) algorithm. We first employ the recent progressive holistically-nested network (P-HNN) model to identify potential lobar boundaries, then generate final segmentations using a RW that is seeded and weighted by the P-HNN output. We are the first to apply deep learning to PPLS. The advantages are independence from prior airway/vessel segmentations, increased robustness in diseased lungs, and methodological simplicity that does not sacrifice accuracy. Our method posts a high mean Jaccard score of 0.888$\pm$0.164 on a held-out set of 154 CT scans from lung-disease patients, while also significantly (p < 0.001) outperforming a state-of-the-art method.
Progressive and Multi-Path Holistically Nested Neural Networks for Pathological Lung Segmentation from CT Images
Jun 12, 2017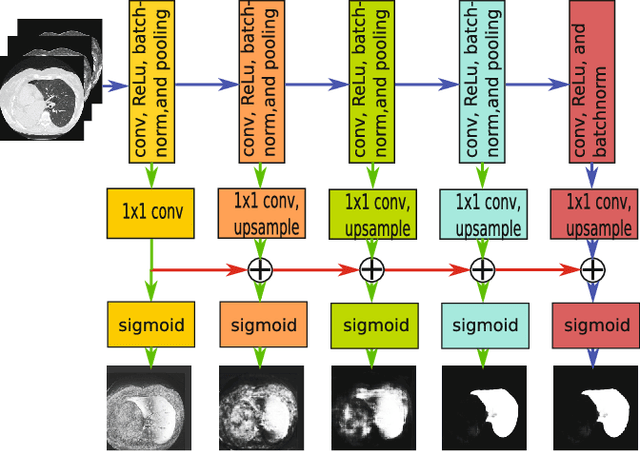

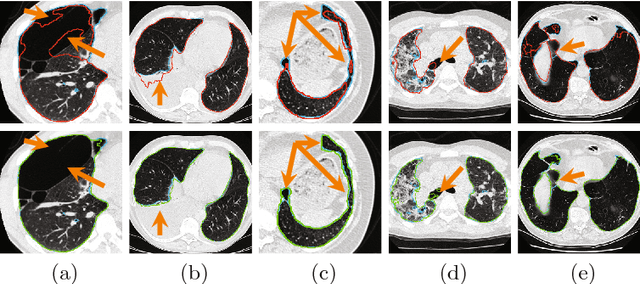
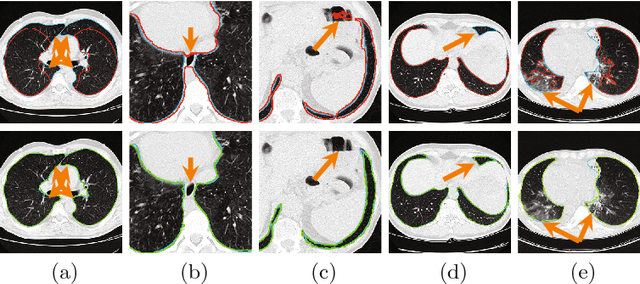
Abstract:Pathological lung segmentation (PLS) is an important, yet challenging, medical image application due to the wide variability of pathological lung appearance and shape. Because PLS is often a pre-requisite for other imaging analytics, methodological simplicity and generality are key factors in usability. Along those lines, we present a bottom-up deep-learning based approach that is expressive enough to handle variations in appearance, while remaining unaffected by any variations in shape. We incorporate the deeply supervised learning framework, but enhance it with a simple, yet effective, progressive multi-path scheme, which more reliably merges outputs from different network stages. The result is a deep model able to produce finer detailed masks, which we call progressive holistically-nested networks (P-HNNs). Using extensive cross-validation, our method is tested on multi-institutional datasets comprising 929 CT scans (848 publicly available), of pathological lungs, reporting mean dice scores of 0.985 and demonstrating significant qualitative and quantitative improvements over state-of-the art approaches.
* 8 Pages, 4 figures, MICCAI 2007
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge