Yili Shen
Driving Reaction Trajectories via Latent Flow Matching
Feb 11, 2026Abstract:Recent advances in reaction prediction have achieved near-saturated accuracy on standard benchmarks (e.g., USPTO), yet most state-of-the-art models formulate the task as a one-shot mapping from reactants to products, offering limited insight into the underlying reaction process. Procedural alternatives introduce stepwise generation but often rely on mechanism-specific supervision, discrete symbolic edits, and computationally expensive inference. In this work, we propose LatentRxnFlow, a new reaction prediction paradigm that models reactions as continuous latent trajectories anchored at the thermodynamic product state. Built on Conditional Flow Matching, our approach learns time-dependent latent dynamics directly from standard reactant-product pairs, without requiring mechanistic annotations or curated intermediate labels. While LatentRxnFlow achieves state-of-the-art performance on USPTO benchmarks, more importantly, the continuous formulation exposes the full generative trajectory, enabling trajectory-level diagnostics that are difficult to realize with discrete or one-shot models. We show that latent trajectory analysis allows us to localize and characterize failure modes and to mitigate certain errors via gated inference. Furthermore, geometric properties of the learned trajectories provide an intrinsic signal of epistemic uncertainty, helping prioritize reliably predictable reaction outcomes and flag ambiguous cases for additional validation. Overall, LatentRxnFlow combines strong predictive accuracy with improved transparency, diagnosability, and uncertainty awareness, moving reaction prediction toward more trustworthy deployment in high-throughput discovery workflows.
Integrating Graph Neural Networks and Many-Body Expansion Theory for Potential Energy Surfaces
Nov 03, 2024



Abstract:Rational design of next-generation functional materials relied on quantitative predictions of their electronic structures beyond single building blocks. First-principles quantum mechanical (QM) modeling became infeasible as the size of a material grew beyond hundreds of atoms. In this study, we developed a new computational tool integrating fragment-based graph neural networks (FBGNN) into the fragment-based many-body expansion (MBE) theory, referred to as FBGNN-MBE, and demonstrated its capacity to reproduce full-dimensional potential energy surfaces (FD-PES) for hierarchic chemical systems with manageable accuracy, complexity, and interpretability. In particular, we divided the entire system into basic building blocks (fragments), evaluated their single-fragment energies using a first-principles QM model and attacked many-fragment interactions using the structure-property relationships trained by FBGNNs. Our development of FBGNN-MBE demonstrated the potential of a new framework integrating deep learning models into fragment-based QM methods, and marked a significant step towards computationally aided design of large functional materials.
TCR: A Transformer Based Deep Network for Predicting Cancer Drugs Response
Jul 10, 2022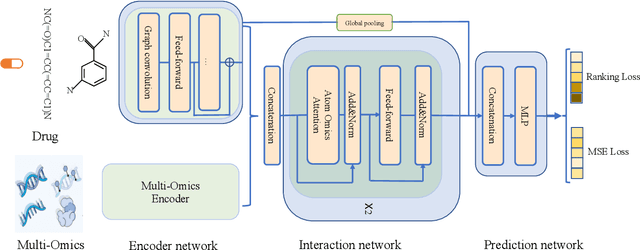
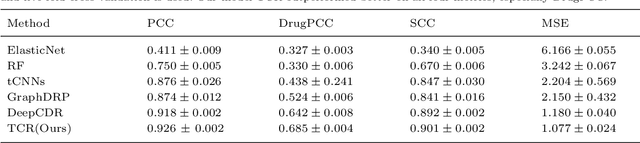
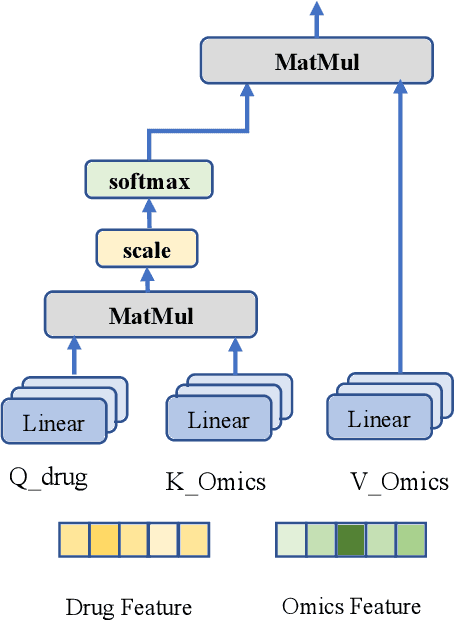

Abstract:Predicting clinical outcomes to anti-cancer drugs on a personalized basis is challenging in cancer treatment due to the heterogeneity of tumors. Traditional computational efforts have been made to model the effect of drug response on individual samples depicted by their molecular profile, yet overfitting occurs because of the high dimension for omics data, hindering models from clinical application. Recent research shows that deep learning is a promising approach to build drug response models by learning alignment patterns between drugs and samples. However, existing studies employed the simple feature fusion strategy and only considered the drug features as a whole representation while ignoring the substructure information that may play a vital role when aligning drugs and genes. Hereby in this paper, we propose TCR (Transformer based network for Cancer drug Response) to predict anti-cancer drug response. By utilizing an attention mechanism, TCR is able to learn the interactions between drug atom/sub-structure and molecular signatures efficiently in our study. Furthermore, a dual loss function and cross sampling strategy were designed to improve the prediction power of TCR. We show that TCR outperformed all other methods under various data splitting strategies on all evaluation matrices (some with significant improvement). Extensive experiments demonstrate that TCR shows significantly improved generalization ability on independent in-vitro experiments and in-vivo real patient data. Our study highlights the prediction power of TCR and its potential value for cancer drug repurpose and precision oncology treatment.
Improving Subgraph Representation Learning via Multi-View Augmentation
May 25, 2022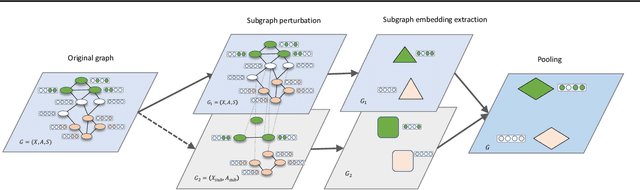
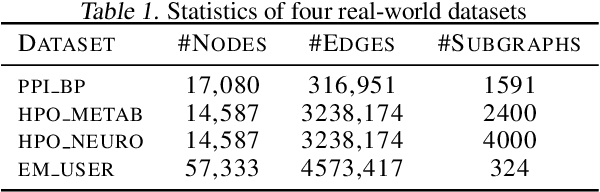
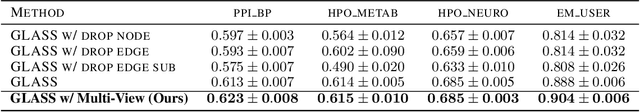
Abstract:Subgraph representation learning based on Graph Neural Network (GNN) has broad applications in chemistry and biology, such as molecule property prediction and gene collaborative function prediction. On the other hand, graph augmentation techniques have shown promising results in improving graph-based and node-based classification tasks but are rarely explored in the GNN-based subgraph representation learning literature. In this work, we developed a novel multiview augmentation mechanism to improve subgraph representation learning and thus the accuracy of downstream prediction tasks. The augmentation technique creates multiple variants of subgraphs and embeds these variants into the original graph to achieve both high training efficiency, scalability, and improved accuracy. Experiments on several real-world subgraph benchmarks demonstrate the superiority of our proposed multi-view augmentation techniques.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge