Tonia Vincent
Deformation-Recovery Diffusion Model (DRDM): Instance Deformation for Image Manipulation and Synthesis
Jul 10, 2024Abstract:In medical imaging, the diffusion models have shown great potential in synthetic image generation tasks. However, these models often struggle with the interpretable connections between the generated and existing images and could create illusions. To address these challenges, our research proposes a novel diffusion-based generative model based on deformation diffusion and recovery. This model, named Deformation-Recovery Diffusion Model (DRDM), diverges from traditional score/intensity and latent feature-based approaches, emphasizing morphological changes through deformation fields rather than direct image synthesis. This is achieved by introducing a topological-preserving deformation field generation method, which randomly samples and integrates a set of multi-scale Deformation Vector Fields (DVF). DRDM is trained to learn to recover unreasonable deformation components, thereby restoring each randomly deformed image to a realistic distribution. These innovations facilitate the generation of diverse and anatomically plausible deformations, enhancing data augmentation and synthesis for further analysis in downstream tasks, such as few-shot learning and image registration. Experimental results in cardiac MRI and pulmonary CT show DRDM is capable of creating diverse, large (over 10% image size deformation scale), and high-quality (negative ratio of folding rate is lower than 1%) deformation fields. The further experimental results in downstream tasks, 2D image segmentation and 3D image registration, indicate significant improvements resulting from DRDM, showcasing the potential of our model to advance image manipulation and synthesis in medical imaging and beyond. Our implementation will be available at https://github.com/jianqingzheng/def_diff_rec.
Recurrent Image Registration using Mutual Attention based Network
Jun 04, 2022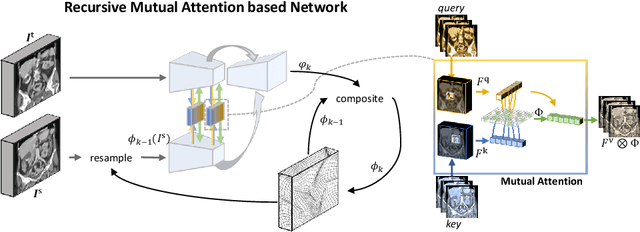

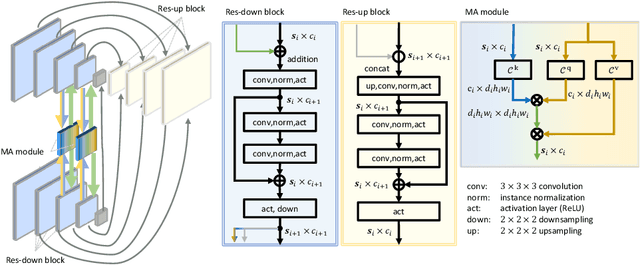

Abstract:Image registration is an important task in medical imaging which estimates the spatial transformation between different images. Many previous studies have used learning-based methods for multi-stage registration to perform 3D image registration to improve performance. The performance of the multi-stage approach, however, is limited by the size of the receptive field where complex motion does not occur at a single spatial scale. We propose a new registration network combining recursive network architecture and mutual attention mechanism to overcome these limitations. Compared with the previous deep learning methods, our network based on the recursive structure achieves the highest accuracy in lung Computed Tomography (CT) data set (Dice score of 92\% and average surface distance of 3.8mm for lungs) and one of the most accurate results in abdominal CT data set with 9 organs of various sizes (Dice score of 55\% and average surface distance of 7.8mm). We also showed that adding 3 recursive networks is sufficient to achieve the state-of-the-art results without a significant increase in the inference time.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge