Tom Syer
Combiner and HyperCombiner Networks: Rules to Combine Multimodality MR Images for Prostate Cancer Localisation
Jul 17, 2023



Abstract:One of the distinct characteristics in radiologists' reading of multiparametric prostate MR scans, using reporting systems such as PI-RADS v2.1, is to score individual types of MR modalities, T2-weighted, diffusion-weighted, and dynamic contrast-enhanced, and then combine these image-modality-specific scores using standardised decision rules to predict the likelihood of clinically significant cancer. This work aims to demonstrate that it is feasible for low-dimensional parametric models to model such decision rules in the proposed Combiner networks, without compromising the accuracy of predicting radiologic labels: First, it is shown that either a linear mixture model or a nonlinear stacking model is sufficient to model PI-RADS decision rules for localising prostate cancer. Second, parameters of these (generalised) linear models are proposed as hyperparameters, to weigh multiple networks that independently represent individual image modalities in the Combiner network training, as opposed to end-to-end modality ensemble. A HyperCombiner network is developed to train a single image segmentation network that can be conditioned on these hyperparameters during inference, for much improved efficiency. Experimental results based on data from 850 patients, for the application of automating radiologist labelling multi-parametric MR, compare the proposed combiner networks with other commonly-adopted end-to-end networks. Using the added advantages of obtaining and interpreting the modality combining rules, in terms of the linear weights or odds-ratios on individual image modalities, three clinical applications are presented for prostate cancer segmentation, including modality availability assessment, importance quantification and rule discovery.
Bi-parametric prostate MR image synthesis using pathology and sequence-conditioned stable diffusion
Mar 03, 2023Abstract:We propose an image synthesis mechanism for multi-sequence prostate MR images conditioned on text, to control lesion presence and sequence, as well as to generate paired bi-parametric images conditioned on images e.g. for generating diffusion-weighted MR from T2-weighted MR for paired data, which are two challenging tasks in pathological image synthesis. Our proposed mechanism utilises and builds upon the recent stable diffusion model by proposing image-based conditioning for paired data generation. We validate our method using 2D image slices from real suspected prostate cancer patients. The realism of the synthesised images is validated by means of a blind expert evaluation for identifying real versus fake images, where a radiologist with 4 years experience reading urological MR only achieves 59.4% accuracy across all tested sequences (where chance is 50%). For the first time, we evaluate the realism of the generated pathology by blind expert identification of the presence of suspected lesions, where we find that the clinician performs similarly for both real and synthesised images, with a 2.9 percentage point difference in lesion identification accuracy between real and synthesised images, demonstrating the potentials in radiological training purposes. Furthermore, we also show that a machine learning model, trained for lesion identification, shows better performance (76.2% vs 70.4%, statistically significant improvement) when trained with real data augmented by synthesised data as opposed to training with only real images, demonstrating usefulness for model training.
Cross-Modality Image Registration using a Training-Time Privileged Third Modality
Jul 26, 2022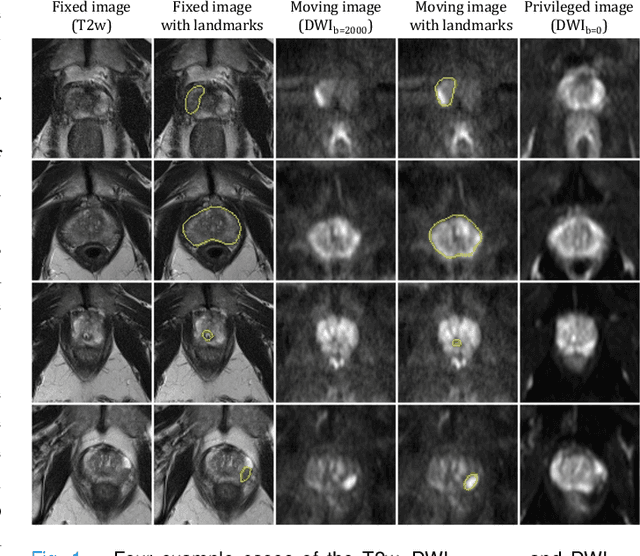
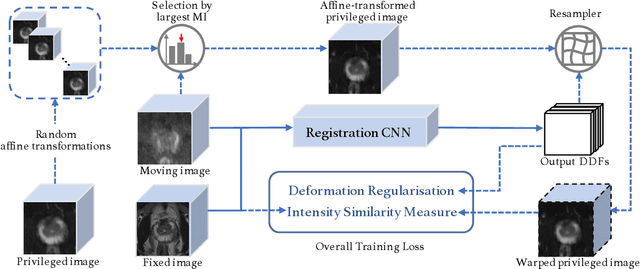
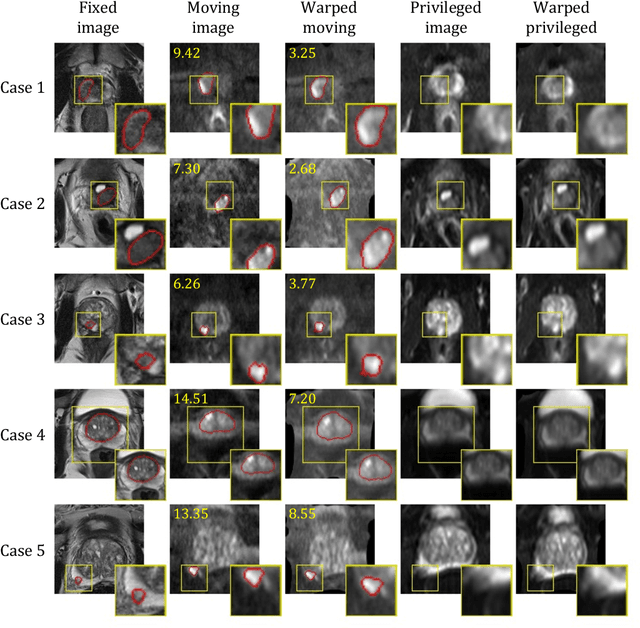
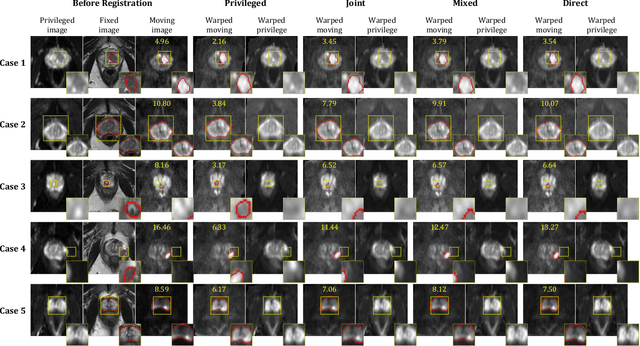
Abstract:In this work, we consider the task of pairwise cross-modality image registration, which may benefit from exploiting additional images available only at training time from an additional modality that is different to those being registered. As an example, we focus on aligning intra-subject multiparametric Magnetic Resonance (mpMR) images, between T2-weighted (T2w) scans and diffusion-weighted scans with high b-value (DWI$_{high-b}$). For the application of localising tumours in mpMR images, diffusion scans with zero b-value (DWI$_{b=0}$) are considered easier to register to T2w due to the availability of corresponding features. We propose a learning from privileged modality algorithm, using a training-only imaging modality DWI$_{b=0}$, to support the challenging multi-modality registration problems. We present experimental results based on 369 sets of 3D multiparametric MRI images from 356 prostate cancer patients and report, with statistical significance, a lowered median target registration error of 4.34 mm, when registering the holdout DWI$_{high-b}$ and T2w image pairs, compared with that of 7.96 mm before registration. Results also show that the proposed learning-based registration networks enabled efficient registration with comparable or better accuracy, compared with a classical iterative algorithm and other tested learning-based methods with/without the additional modality. These compared algorithms also failed to produce any significantly improved alignment between DWI$_{high-b}$ and T2w in this challenging application.
The impact of using voxel-level segmentation metrics on evaluating multifocal prostate cancer localisation
Mar 31, 2022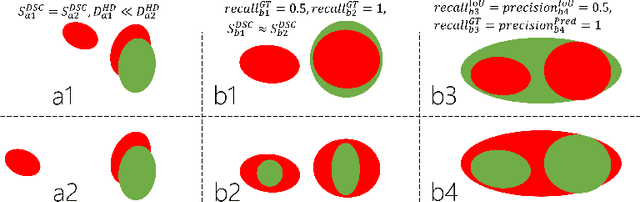
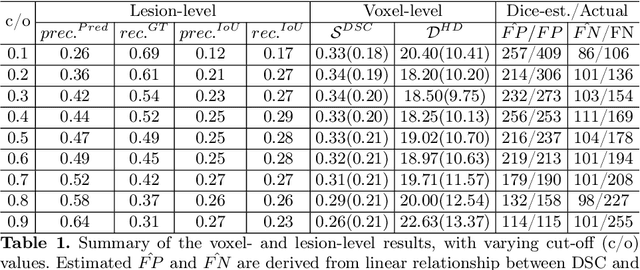
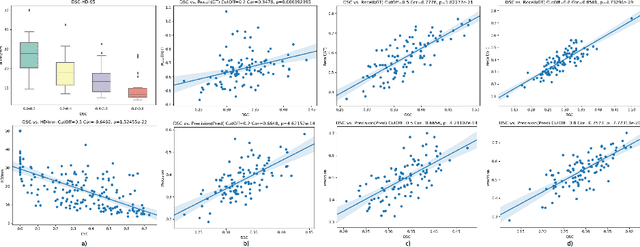

Abstract:Dice similarity coefficient (DSC) and Hausdorff distance (HD) are widely used for evaluating medical image segmentation. They have also been criticised, when reported alone, for their unclear or even misleading clinical interpretation. DSCs may also differ substantially from HDs, due to boundary smoothness or multiple regions of interest (ROIs) within a subject. More importantly, either metric can also have a nonlinear, non-monotonic relationship with outcomes based on Type 1 and 2 errors, designed for specific clinical decisions that use the resulting segmentation. Whilst cases causing disagreement between these metrics are not difficult to postulate. This work first proposes a new asymmetric detection metric, adapting those used in object detection, for planning prostate cancer procedures. The lesion-level metrics is then compared with the voxel-level DSC and HD, whereas a 3D UNet is used for segmenting lesions from multiparametric MR (mpMR) images. Based on experimental results we report pairwise agreement and correlation 1) between DSC and HD, and 2) between voxel-level DSC and recall-controlled precision at lesion-level, with Cohen's [0.49, 0.61] and Pearson's [0.66, 0.76] (p-values}<0.001) at varying cut-offs. However, the differences in false-positives and false-negatives, between the actual errors and the perceived counterparts if DSC is used, can be as high as 152 and 154, respectively, out of the 357 test set lesions. We therefore carefully conclude that, despite of the significant correlations, voxel-level metrics such as DSC can misrepresent lesion-level detection accuracy for evaluating localisation of multifocal prostate cancer and should be interpreted with caution.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge