Steven E. Williams
ECG Feature Importance Rankings: Cardiologists vs. Algorithms
Apr 05, 2023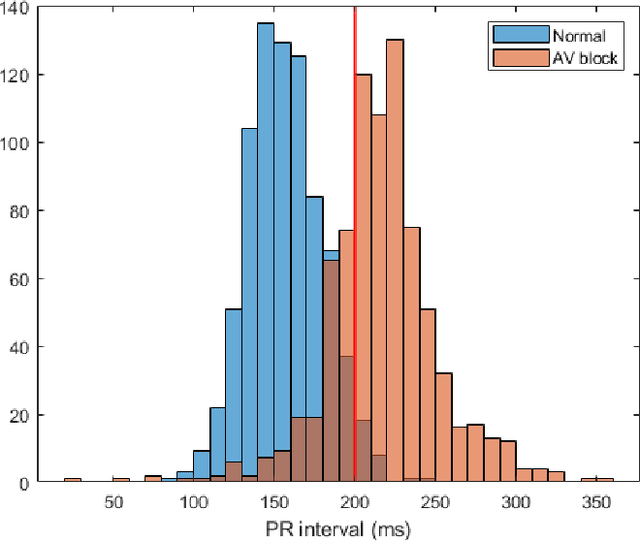
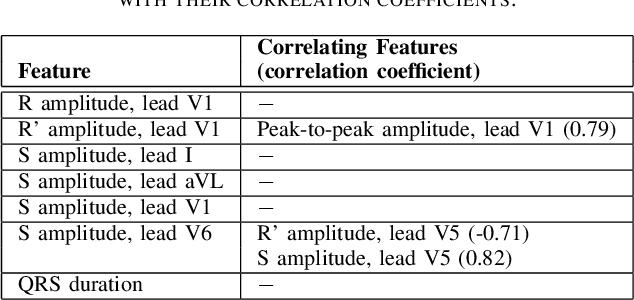
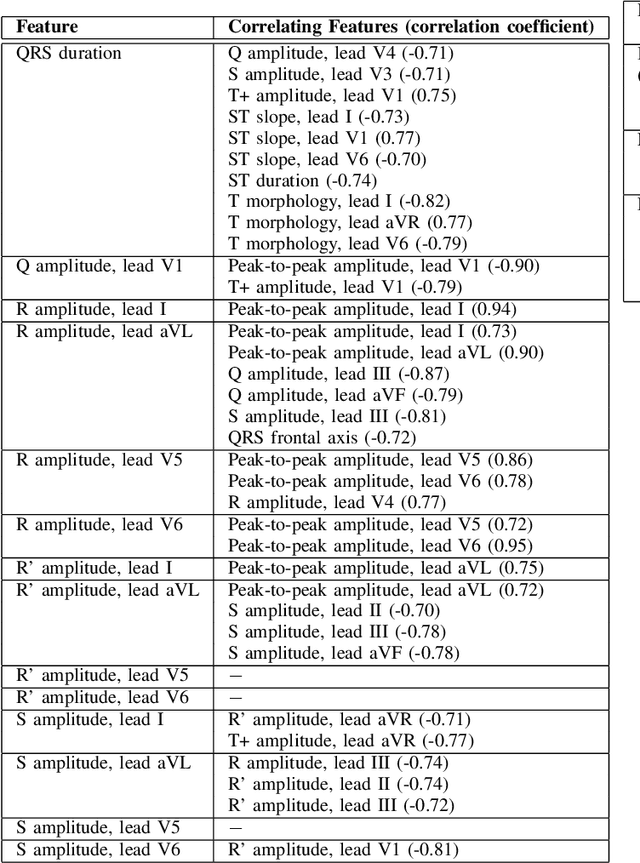

Abstract:Feature importance methods promise to provide a ranking of features according to importance for a given classification task. A wide range of methods exist but their rankings often disagree and they are inherently difficult to evaluate due to a lack of ground truth beyond synthetic datasets. In this work, we put feature importance methods to the test on real-world data in the domain of cardiology, where we try to distinguish three specific pathologies from healthy subjects based on ECG features comparing to features used in cardiologists' decision rules as ground truth. Some methods generally performed well and others performed poorly, while some methods did well on some but not all of the problems considered.
MedalCare-XL: 16,900 healthy and pathological 12 lead ECGs obtained through electrophysiological simulations
Nov 29, 2022Abstract:Mechanistic cardiac electrophysiology models allow for personalized simulations of the electrical activity in the heart and the ensuing electrocardiogram (ECG) on the body surface. As such, synthetic signals possess known ground truth labels of the underlying disease and can be employed for validation of machine learning ECG analysis tools in addition to clinical signals. Recently, synthetic ECGs were used to enrich sparse clinical data or even replace them completely during training leading to improved performance on real-world clinical test data. We thus generated a novel synthetic database comprising a total of 16,900 12 lead ECGs based on electrophysiological simulations equally distributed into healthy control and 7 pathology classes. The pathological case of myocardial infraction had 6 sub-classes. A comparison of extracted features between the virtual cohort and a publicly available clinical ECG database demonstrated that the synthetic signals represent clinical ECGs for healthy and pathological subpopulations with high fidelity. The ECG database is split into training, validation, and test folds for development and objective assessment of novel machine learning algorithms.
Whole Heart Anatomical Refinement from CCTA using Extrapolation and Parcellation
Nov 18, 2021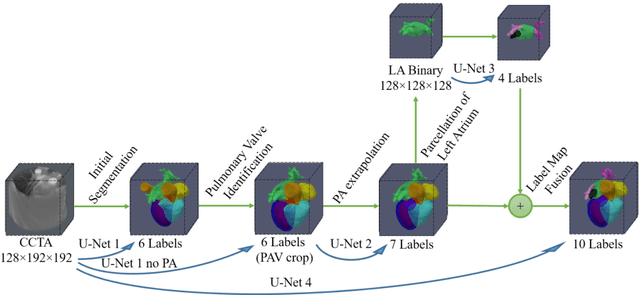
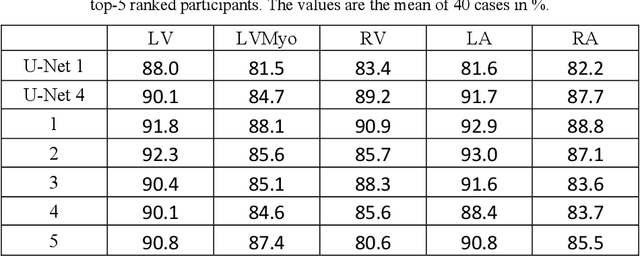
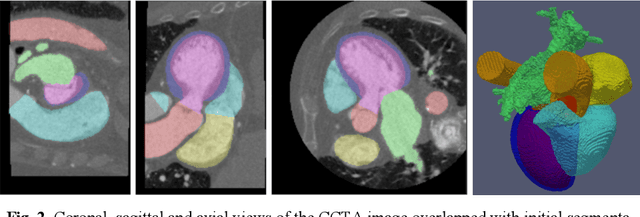
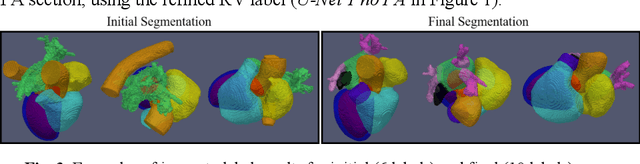
Abstract:Coronary computed tomography angiography (CCTA) provides detailed an-atomical information on all chambers of the heart. Existing segmentation tools can label the gross anatomy, but addition of application-specific labels can require detailed and often manual refinement. We developed a U-Net based framework to i) extrapolate a new label from existing labels, and ii) parcellate one label into multiple labels, both using label-to-label mapping, to create a desired segmentation that could then be learnt directly from the image (image- to-label mapping). This approach only required manual correction in a small subset of cases (80 for extrapolation, 50 for parcella-tion, compared with 260 for initial labels). An initial 6-label segmentation (left ventricle, left ventricular myocardium, right ventricle, left atrium, right atrium and aorta) was refined to a 10-label segmentation that added a label for the pulmonary artery and divided the left atrium label into body, left and right veins and appendage components. The final method was tested using 30 cases, 10 each from Philips, Siemens and Toshiba scanners. In addition to the new labels, the median Dice scores were improved for all the initial 6 labels to be above 95% in the 10-label segmentation, e.g. from 91% to 97% for the left atrium body and from 92% to 96% for the right ventricle. This method provides a simple framework for flexible refinement of anatomical labels. The code and executables are available at cemrg.com.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge