Sophie Hanna Langbein
Machine Learning in Epidemiology
Feb 18, 2026Abstract:In the age of digital epidemiology, epidemiologists are faced by an increasing amount of data of growing complexity and dimensionality. Machine learning is a set of powerful tools that can help to analyze such enormous amounts of data. This chapter lays the methodological foundations for successfully applying machine learning in epidemiology. It covers the principles of supervised and unsupervised learning and discusses the most important machine learning methods. Strategies for model evaluation and hyperparameter optimization are developed and interpretable machine learning is introduced. All these theoretical parts are accompanied by code examples in R, where an example dataset on heart disease is used throughout the chapter.
Functional Decomposition and Shapley Interactions for Interpreting Survival Models
Feb 18, 2026Abstract:Hazard and survival functions are natural, interpretable targets in time-to-event prediction, but their inherent non-additivity fundamentally limits standard additive explanation methods. We introduce Survival Functional Decomposition (SurvFD), a principled approach for analyzing feature interactions in machine learning survival models. By decomposing higher-order effects into time-dependent and time-independent components, SurvFD offers a previously unrecognized perspective on survival explanations, explicitly characterizing when and why additive explanations fail. Building on this theoretical decomposition, we propose SurvSHAP-IQ, which extends Shapley interactions to time-indexed functions, providing a practical estimator for higher-order, time-dependent interactions. Together, SurvFD and SurvSHAP-IQ establish an interaction- and time-aware interpretability approach for survival modeling, with broad applicability across time-to-event prediction tasks.
Gradient-based Explanations for Deep Learning Survival Models
Feb 07, 2025Abstract:Deep learning survival models often outperform classical methods in time-to-event predictions, particularly in personalized medicine, but their "black box" nature hinders broader adoption. We propose a framework for gradient-based explanation methods tailored to survival neural networks, extending their use beyond regression and classification. We analyze the implications of their theoretical assumptions for time-dependent explanations in the survival setting and propose effective visualizations incorporating the temporal dimension. Experiments on synthetic data show that gradient-based methods capture the magnitude and direction of local and global feature effects, including time dependencies. We introduce GradSHAP(t), a gradient-based counterpart to SurvSHAP(t), which outperforms SurvSHAP(t) and SurvLIME in a computational speed vs. accuracy trade-off. Finally, we apply these methods to medical data with multi-modal inputs, revealing relevant tabular features and visual patterns, as well as their temporal dynamics.
Interpretable Machine Learning for Survival Analysis
Mar 15, 2024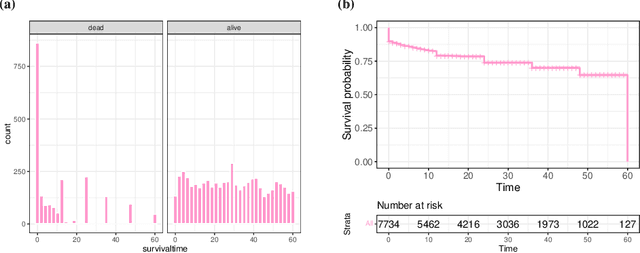
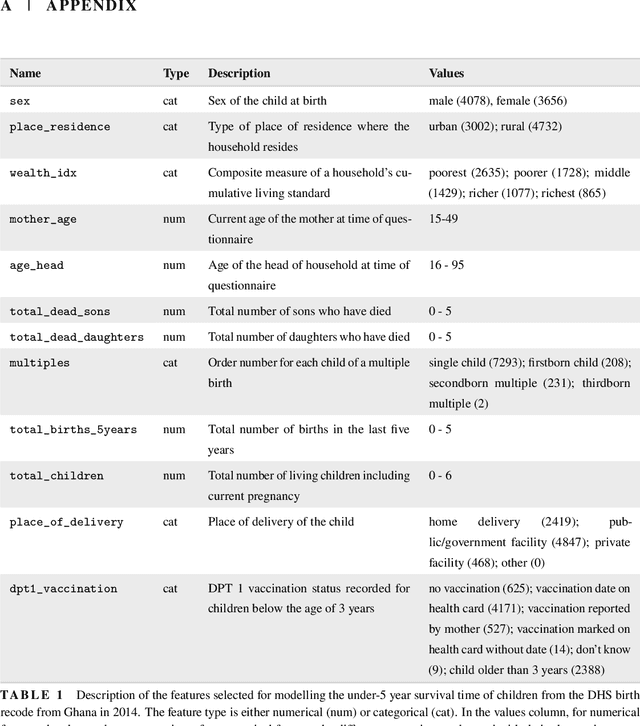
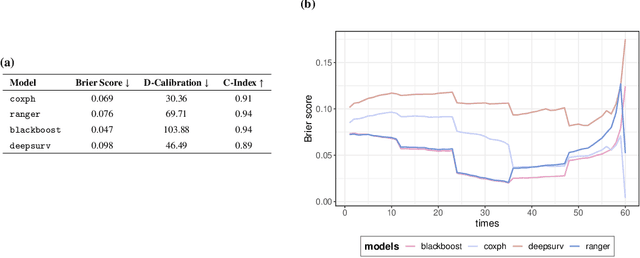
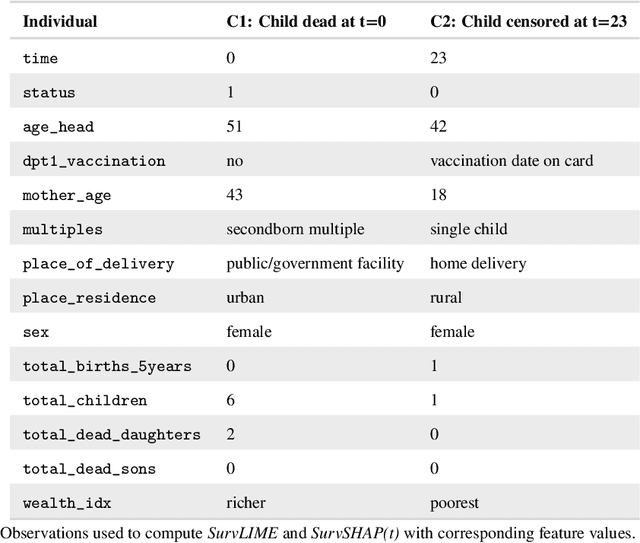
Abstract:With the spread and rapid advancement of black box machine learning models, the field of interpretable machine learning (IML) or explainable artificial intelligence (XAI) has become increasingly important over the last decade. This is particularly relevant for survival analysis, where the adoption of IML techniques promotes transparency, accountability and fairness in sensitive areas, such as clinical decision making processes, the development of targeted therapies, interventions or in other medical or healthcare related contexts. More specifically, explainability can uncover a survival model's potential biases and limitations and provide more mathematically sound ways to understand how and which features are influential for prediction or constitute risk factors. However, the lack of readily available IML methods may have deterred medical practitioners and policy makers in public health from leveraging the full potential of machine learning for predicting time-to-event data. We present a comprehensive review of the limited existing amount of work on IML methods for survival analysis within the context of the general IML taxonomy. In addition, we formally detail how commonly used IML methods, such as such as individual conditional expectation (ICE), partial dependence plots (PDP), accumulated local effects (ALE), different feature importance measures or Friedman's H-interaction statistics can be adapted to survival outcomes. An application of several IML methods to real data on data on under-5 year mortality of Ghanaian children from the Demographic and Health Surveys (DHS) Program serves as a tutorial or guide for researchers, on how to utilize the techniques in practice to facilitate understanding of model decisions or predictions.
survex: an R package for explaining machine learning survival models
Aug 30, 2023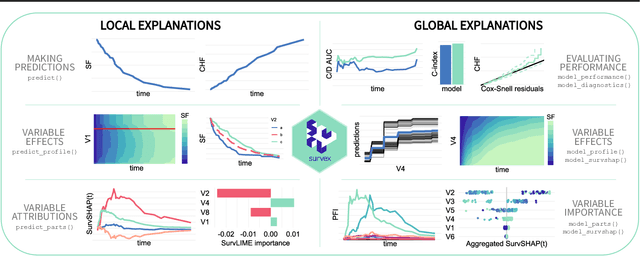
Abstract:Due to their flexibility and superior performance, machine learning models frequently complement and outperform traditional statistical survival models. However, their widespread adoption is hindered by a lack of user-friendly tools to explain their internal operations and prediction rationales. To tackle this issue, we introduce the survex R package, which provides a cohesive framework for explaining any survival model by applying explainable artificial intelligence techniques. The capabilities of the proposed software encompass understanding and diagnosing survival models, which can lead to their improvement. By revealing insights into the decision-making process, such as variable effects and importances, survex enables the assessment of model reliability and the detection of biases. Thus, transparency and responsibility may be promoted in sensitive areas, such as biomedical research and healthcare applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge