Saima Ben Hadj
Tribvn Healthcare
Interpretable HER2 scoring by evaluating clinical Guidelines through a weakly supervised, constrained Deep Learning Approach
Nov 17, 2022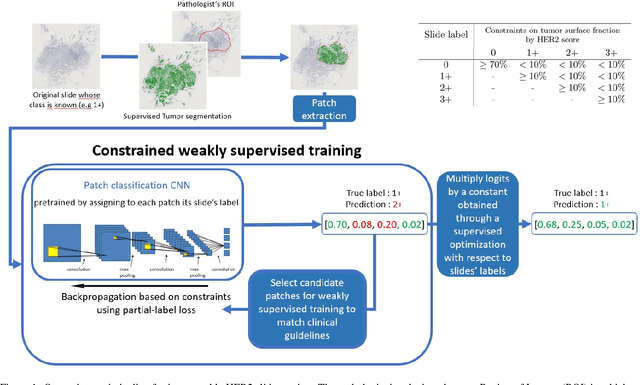
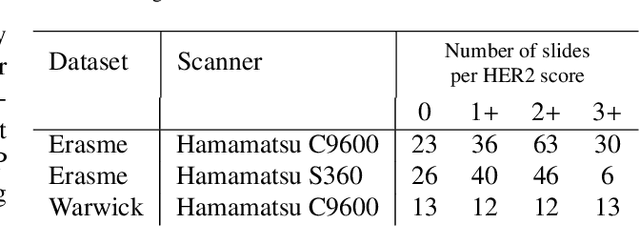

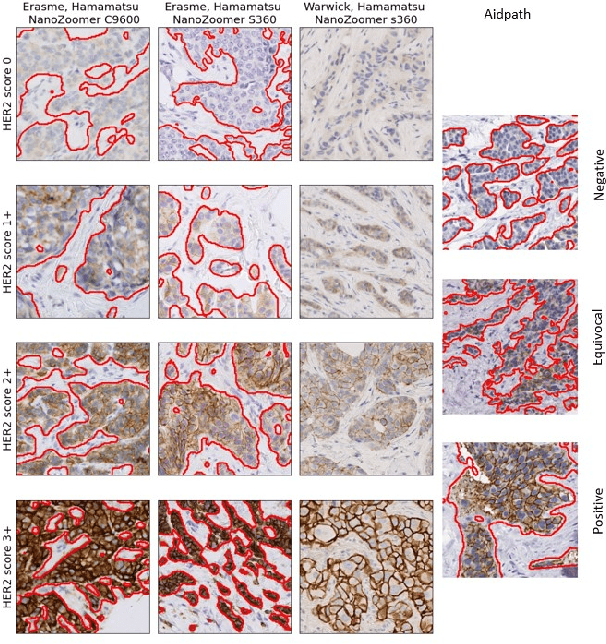
Abstract:The evaluation of the Human Epidermal growth factor Receptor-2 (HER2) expression is an important prognostic biomarker for breast cancer treatment selection. However, HER2 scoring has notoriously high interobserver variability due to stain variations between centers and the need to estimate visually the staining intensity in specific percentages of tumor area. In this paper, focusing on the interpretability of HER2 scoring by a pathologist, we propose a semi-automatic, two-stage deep learning approach that directly evaluates the clinical HER2 guidelines defined by the American Society of Clinical Oncology/ College of American Pathologists (ASCO/CAP). In the first stage, we segment the invasive tumor over the user-indicated Region of Interest (ROI). Then, in the second stage, we classify the tumor tissue into four HER2 classes. For the classification stage, we use weakly supervised, constrained optimization to find a model that classifies cancerous patches such that the tumor surface percentage meets the guidelines specification of each HER2 class. We end the second stage by freezing the model and refining its output logits in a supervised way to all slide labels in the training set. To ensure the quality of our dataset's labels, we conducted a multi-pathologist HER2 scoring consensus. For the assessment of doubtful cases where no consensus was found, our model can help by interpreting its HER2 class percentages output. We achieve a performance of 0.78 in F1-score on the test set while keeping our model interpretable for the pathologist, hopefully contributing to interpretable AI models in digital pathology.
Enabling Collagen Quantification on HE-stained Slides Through Stain Deconvolution and Restained HE-HES
Nov 17, 2022



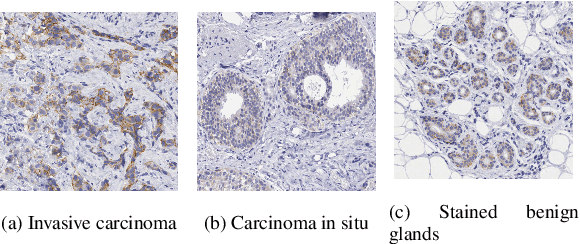
Abstract:In histology, the presence of collagen in the extra-cellular matrix has both diagnostic and prognostic value for cancer malignancy, and can be highlighted by adding Saffron (S) to a routine Hematoxylin and Eosin (HE) staining. However, Saffron is not usually added because of the additional cost and because pathologists are accustomed to HE, with the exception of France-based laboratories. In this paper, we show that it is possible to quantify the collagen content from the HE image alone and to digitally create an HES image. To do so, we trained a UNet to predict the Saffron densities from HE images. We created a dataset of registered, restained HE-HES slides and we extracted the Saffron concentrations as ground truth using stain deconvolution on the HES images. Our model reached a Mean Absolute Error of 0.0668 $\pm$ 0.0002 (Saffron values between 0 and 1) on a 3-fold testing set. We hope our approach can aid in improving the clinical workflow while reducing reagent costs for laboratories.
Mitosis domain generalization in histopathology images -- The MIDOG challenge
Apr 06, 2022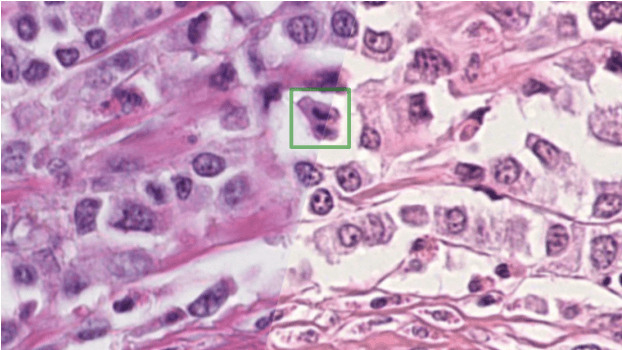
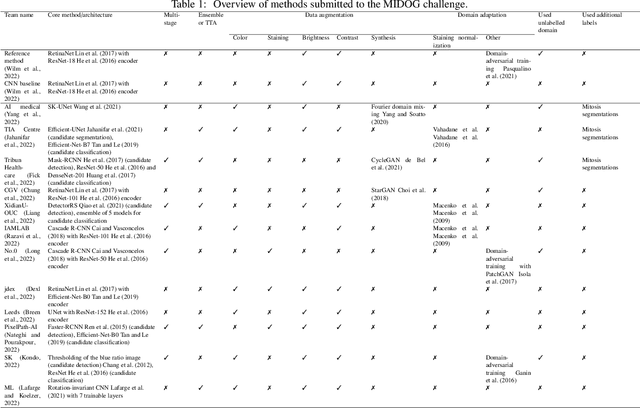
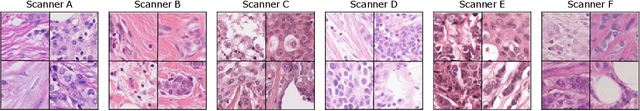
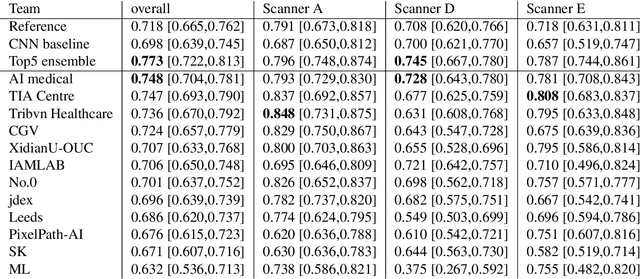
Abstract:The density of mitotic figures within tumor tissue is known to be highly correlated with tumor proliferation and thus is an important marker in tumor grading. Recognition of mitotic figures by pathologists is known to be subject to a strong inter-rater bias, which limits the prognostic value. State-of-the-art deep learning methods can support the expert in this assessment but are known to strongly deteriorate when applied in a different clinical environment than was used for training. One decisive component in the underlying domain shift has been identified as the variability caused by using different whole slide scanners. The goal of the MICCAI MIDOG 2021 challenge has been to propose and evaluate methods that counter this domain shift and derive scanner-agnostic mitosis detection algorithms. The challenge used a training set of 200 cases, split across four scanning systems. As a test set, an additional 100 cases split across four scanning systems, including two previously unseen scanners, were given. The best approaches performed on an expert level, with the winning algorithm yielding an F_1 score of 0.748 (CI95: 0.704-0.781). In this paper, we evaluate and compare the approaches that were submitted to the challenge and identify methodological factors contributing to better performance.
Robust Mitosis Detection Using a Cascade Mask-RCNN Approach With Domain-Specific Residual Cycle-GAN Data Augmentation
Sep 28, 2021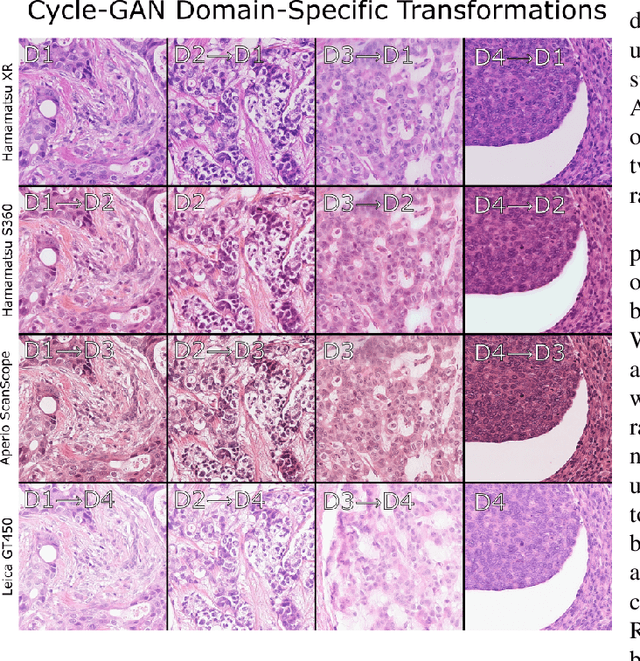
Abstract:For the MIDOG mitosis detection challenge, we created a cascade algorithm consisting of a Mask-RCNN detector, followed by a classification ensemble consisting of ResNet50 and DenseNet201 to refine detected mitotic candidates. The MIDOG training data consists of 200 frames originating from four scanners, three of which are annotated for mitotic instances with centroid annotations. Our main algorithmic choices are as follows: first, to enhance the generalizability of our detector and classification networks, we use a state-of-the-art residual Cycle-GAN to transform each scanner domain to every other scanner domain. During training, we then randomly load, for each image, one of the four domains. In this way, our networks can learn from the fourth non-annotated scanner domain even if we don't have annotations for it. Second, for training the detector network, rather than using centroid-based fixed-size bounding boxes, we create mitosis-specific bounding boxes. We do this by manually annotating a small selection of mitoses, training a Mask-RCNN on this small dataset, and applying it to the rest of the data to obtain full annotations. We trained the follow-up classification ensemble using only the challenge-provided positive and hard-negative examples. On the preliminary test set, the algorithm scores an F1 score of 0.7578, putting us as the second-place team on the leaderboard.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge