Roberto A. Lotufo
Quati: A Brazilian Portuguese Information Retrieval Dataset from Native Speakers
Apr 10, 2024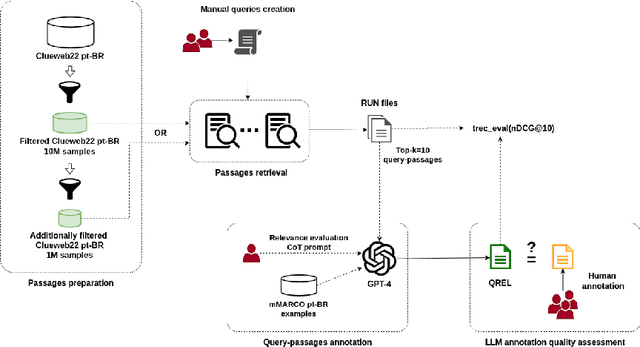
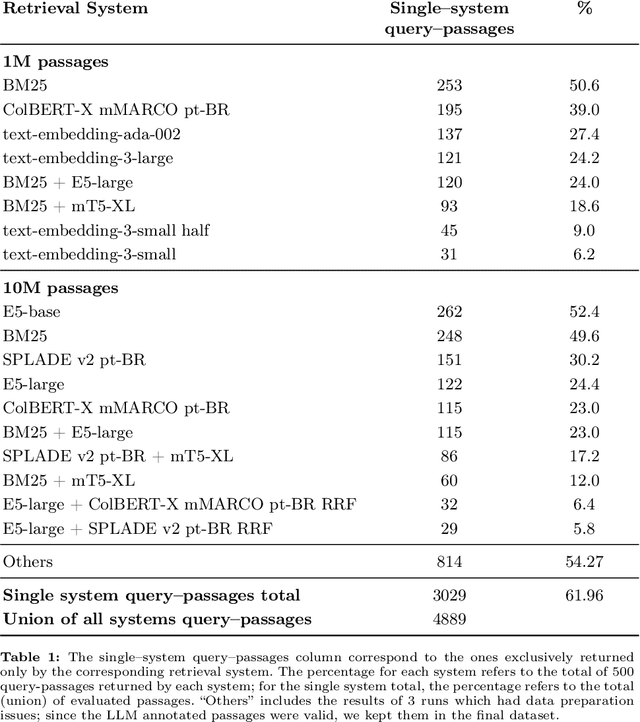
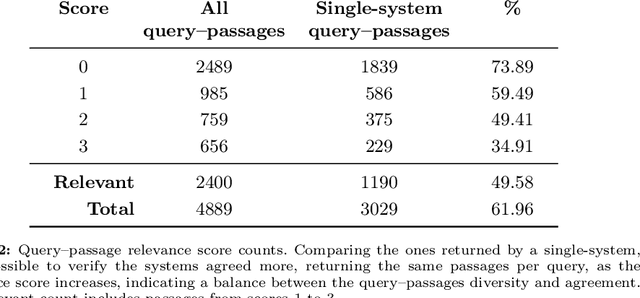
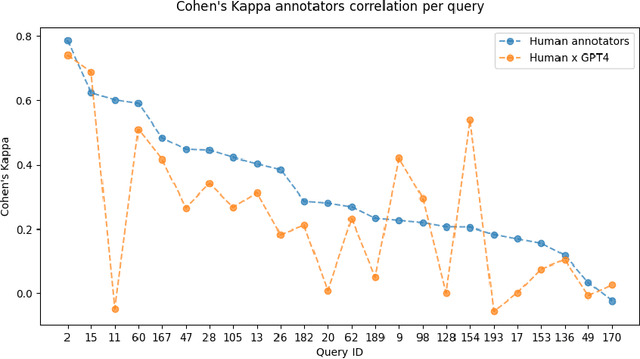
Abstract:Despite Portuguese being one of the most spoken languages in the world, there is a lack of high-quality information retrieval datasets in that language. We present Quati, a dataset specifically designed for the Brazilian Portuguese language. It comprises a collection of queries formulated by native speakers and a curated set of documents sourced from a selection of high-quality Brazilian Portuguese websites. These websites are frequented more likely by real users compared to those randomly scraped, ensuring a more representative and relevant corpus. To label the query-document pairs, we use a state-of-the-art LLM, which shows inter-annotator agreement levels comparable to human performance in our assessments. We provide a detailed description of our annotation methodology to enable others to create similar datasets for other languages, providing a cost-effective way of creating high-quality IR datasets with an arbitrary number of labeled documents per query. Finally, we evaluate a diverse range of open-source and commercial retrievers to serve as baseline systems. Quati is publicly available at https://huggingface.co/datasets/unicamp-dl/quati and all scripts at https://github.com/unicamp-dl/quati .
MEDPSeg: End-to-end segmentation of pulmonary structures and lesions in computed tomography
Dec 04, 2023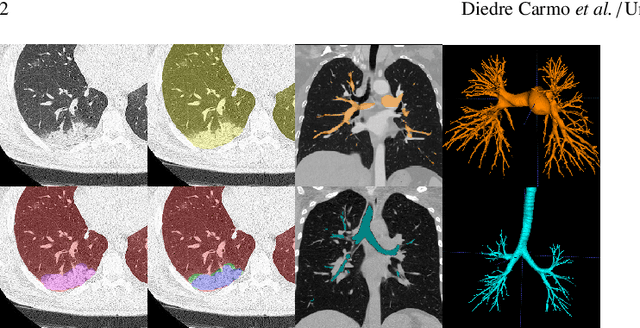
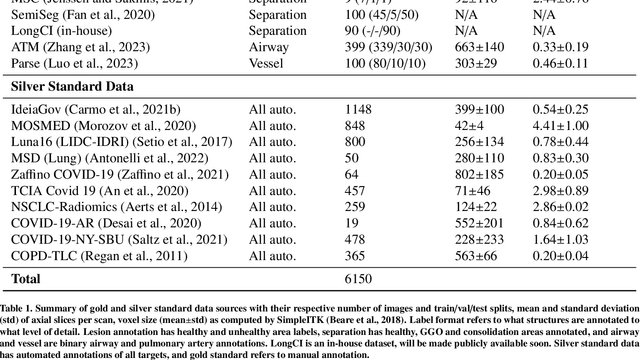
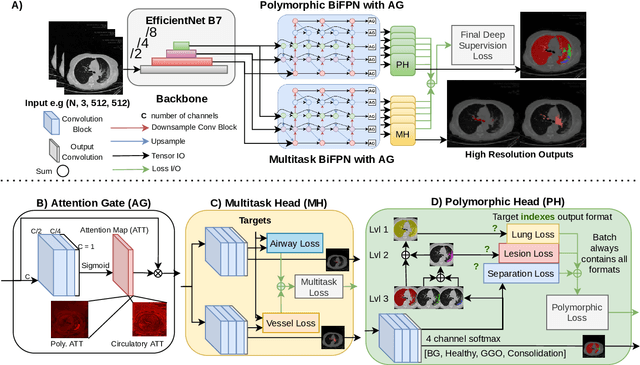

Abstract:The COVID-19 pandemic response highlighted the potential of deep learning methods in facilitating the diagnosis and prognosis of lung diseases through automated segmentation of normal and abnormal tissue in computed tomography (CT). Such methods not only have the potential to aid in clinical decision-making but also contribute to the comprehension of novel diseases. In light of the labor-intensive nature of manual segmentation for large chest CT cohorts, there is a pressing need for reliable automated approaches that enable efficient analysis of chest CT anatomy in vast research databases, especially in more scarcely annotated targets such as pneumonia consolidations. A limiting factor for the development of such methods is that most current models optimize a fixed annotation format per network output. To tackle this problem, polymorphic training is used to optimize a network with a fixed number of output channels to represent multiple hierarchical anatomic structures, indirectly optimizing more complex labels with simpler annotations. We combined over 6000 volumetric CT scans containing varying formats of manual and automated labels from different sources, and used polymorphic training along with multitask learning to develop MEDPSeg, an end-to-end method for the segmentation of lungs, airways, pulmonary artery, and lung lesions with separation of ground glass opacities, and parenchymal consolidations, all in a single forward prediction. We achieve state-of-the-art performance in multiple targets, particularly in the segmentation of ground glass opacities and consolidations, a challenging problem with limited manual annotation availability. In addition, we provide an open-source implementation with a graphical user interface at https://github.com/MICLab-Unicamp/medpseg.
Automatic segmentation of lung findings in CT and application to Long COVID
Oct 13, 2023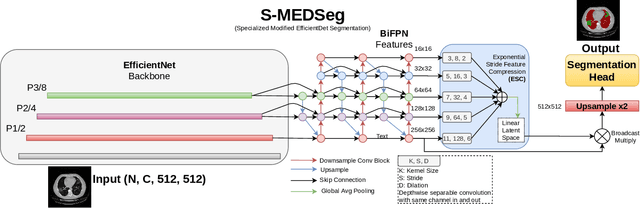
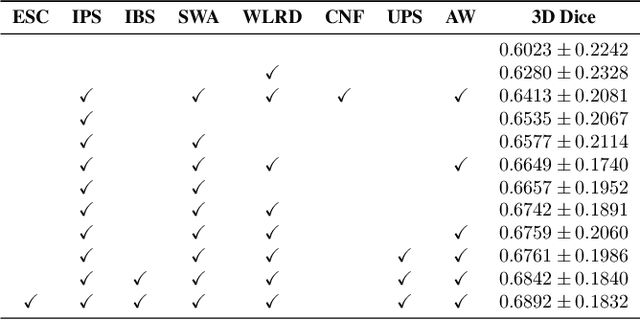
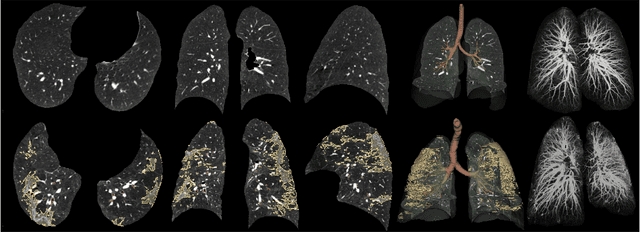
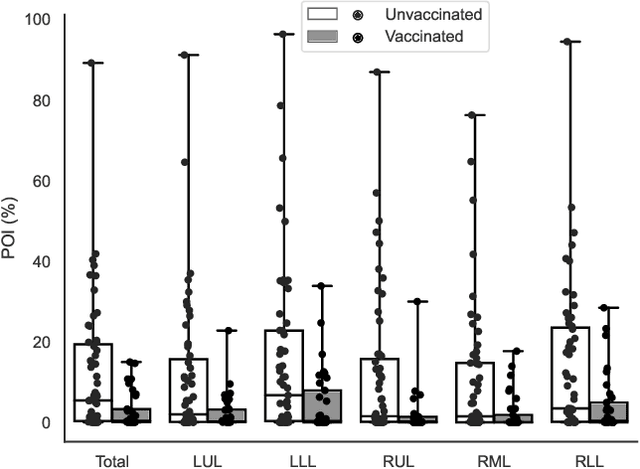
Abstract:Automated segmentation of lung abnormalities in computed tomography is an important step for diagnosing and characterizing lung disease. In this work, we improve upon a previous method and propose S-MEDSeg, a deep learning based approach for accurate segmentation of lung lesions in chest CT images. S-MEDSeg combines a pre-trained EfficientNet backbone, bidirectional feature pyramid network, and modern network advancements to achieve improved segmentation performance. A comprehensive ablation study was performed to evaluate the contribution of the proposed network modifications. The results demonstrate modifications introduced in S-MEDSeg significantly improves segmentation performance compared to the baseline approach. The proposed method is applied to an independent dataset of long COVID inpatients to study the effect of post-acute infection vaccination on extent of lung findings. Open-source code, graphical user interface and pip package are available at https://github.com/MICLab-Unicamp/medseg.
Sequence-to-Sequence Models for Extracting Information from Registration and Legal Documents
Jan 14, 2022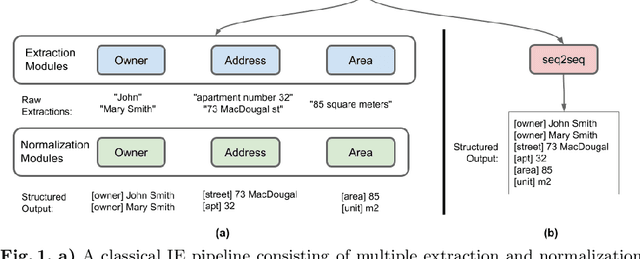



Abstract:A typical information extraction pipeline consists of token- or span-level classification models coupled with a series of pre- and post-processing scripts. In a production pipeline, requirements often change, with classes being added and removed, which leads to nontrivial modifications to the source code and the possible introduction of bugs. In this work, we evaluate sequence-to-sequence models as an alternative to token-level classification methods for information extraction of legal and registration documents. We finetune models that jointly extract the information and generate the output already in a structured format. Post-processing steps are learned during training, thus eliminating the need for rule-based methods and simplifying the pipeline. Furthermore, we propose a novel method to align the output with the input text, thus facilitating system inspection and auditing. Our experiments on four real-world datasets show that the proposed method is an alternative to classical pipelines.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge