Pengpeng Zhang
Retrieval Augmented Comic Image Generation
Jun 14, 2025



Abstract:We present RaCig, a novel system for generating comic-style image sequences with consistent characters and expressive gestures. RaCig addresses two key challenges: (1) maintaining character identity and costume consistency across frames, and (2) producing diverse and vivid character gestures. Our approach integrates a retrieval-based character assignment module, which aligns characters in textual prompts with reference images, and a regional character injection mechanism that embeds character features into specified image regions. Experimental results demonstrate that RaCig effectively generates engaging comic narratives with coherent characters and dynamic interactions. The source code will be publicly available to support further research in this area.
Deformation Driven Seq2Seq Longitudinal Tumor and Organs-at-Risk Prediction for Radiotherapy
Jun 18, 2021



Abstract:Purpose: Radiotherapy presents unique challenges and clinical requirements for longitudinal tumor and organ-at-risk (OAR) prediction during treatment. The challenges include tumor inflammation/edema and radiation-induced changes in organ geometry, whereas the clinical requirements demand flexibility in input/output sequence timepoints to update the predictions on rolling basis and the grounding of all predictions in relationship to the pre-treatment imaging information for response and toxicity assessment in adaptive radiotherapy. Methods: To deal with the aforementioned challenges and to comply with the clinical requirements, we present a novel 3D sequence-to-sequence model based on Convolution Long Short Term Memory (ConvLSTM) that makes use of series of deformation vector fields (DVF) between individual timepoints and reference pre-treatment/planning CTs to predict future anatomical deformations and changes in gross tumor volume as well as critical OARs. High-quality DVF training data is created by employing hyper-parameter optimization on the subset of the training data with DICE coefficient and mutual information metric. We validated our model on two radiotherapy datasets: a publicly available head-and-neck dataset (28 patients with manually contoured pre-, mid-, and post-treatment CTs), and an internal non-small cell lung cancer dataset (63 patients with manually contoured planning CT and 6 weekly CBCTs). Results: The use of DVF representation and skip connections overcomes the blurring issue of ConvLSTM prediction with the traditional image representation. The mean and standard deviation of DICE for predictions of lung GTV at week 4, 5, and 6 were 0.83$\pm$0.09, 0.82$\pm$0.08, and 0.81$\pm$0.10, respectively, and for post-treatment ipsilateral and contralateral parotids, were 0.81$\pm$0.06 and 0.85$\pm$0.02.
Multitask 3D CBCT-to-CT Translation and Organs-at-Risk Segmentation Using Physics-Based Data Augmentation
Mar 09, 2021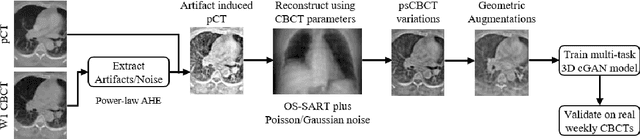
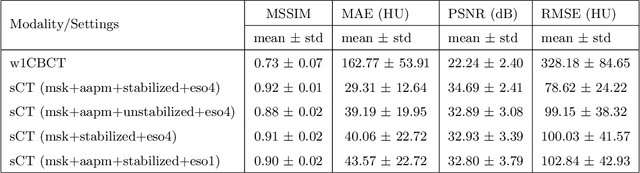
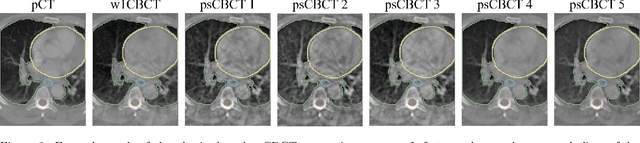
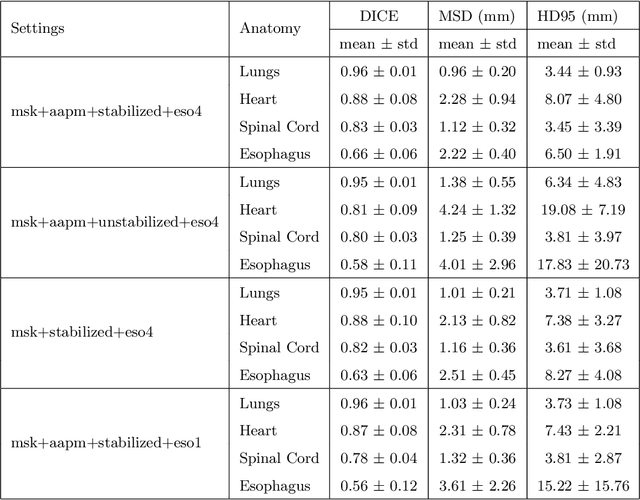
Abstract:Purpose: In current clinical practice, noisy and artifact-ridden weekly cone-beam computed tomography (CBCT) images are only used for patient setup during radiotherapy. Treatment planning is done once at the beginning of the treatment using high-quality planning CT (pCT) images and manual contours for organs-at-risk (OARs) structures. If the quality of the weekly CBCT images can be improved while simultaneously segmenting OAR structures, this can provide critical information for adapting radiotherapy mid-treatment as well as for deriving biomarkers for treatment response. Methods: Using a novel physics-based data augmentation strategy, we synthesize a large dataset of perfectly/inherently registered planning CT and synthetic-CBCT pairs for locally advanced lung cancer patient cohort, which are then used in a multitask 3D deep learning framework to simultaneously segment and translate real weekly CBCT images to high-quality planning CT-like images. Results: We compared the synthetic CT and OAR segmentations generated by the model to real planning CT and manual OAR segmentations and showed promising results. The real week 1 (baseline) CBCT images which had an average MAE of 162.77 HU compared to pCT images are translated to synthetic CT images that exhibit a drastically improved average MAE of 29.31 HU and average structural similarity of 92% with the pCT images. The average DICE scores of the 3D organs-at-risk segmentations are: lungs 0.96, heart 0.88, spinal cord 0.83 and esophagus 0.66. Conclusions: We demonstrate an approach to translate artifact-ridden CBCT images to high quality synthetic CT images while simultaneously generating good quality segmentation masks for different organs-at-risk. This approach could allow clinicians to adjust treatment plans using only the routine low-quality CBCT images, potentially improving patient outcomes.
Generalizable Cone Beam CT Esophagus Segmentation Using In Silico Data Augmentation
Jun 28, 2020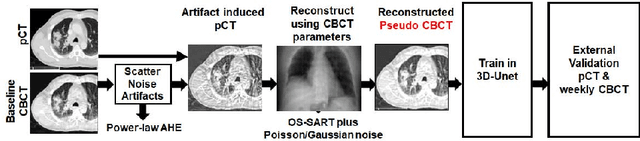
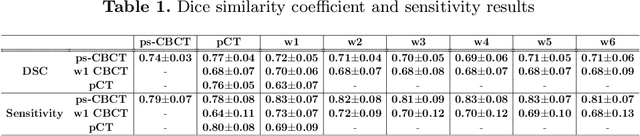
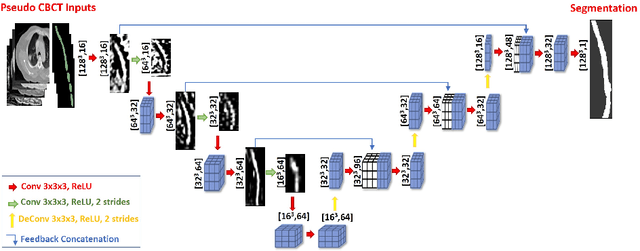

Abstract:Lung cancer radiotherapy entails high quality planning computed tomography (pCT) imaging of the patient with radiation oncologist contouring of the tumor and the organs at risk (OARs) at the start of the treatment. This is followed by weekly low-quality cone beam CT (CBCT) imaging for treatment setup and qualitative visual assessment of tumor and critical OARs. In this work, we aim to make the weekly CBCT assessment quantitative by automatically segmenting the most critical OAR, esophagus, using deep learning and in silico (image-driven simulation) artifact induction to convert pCTs to pseudo-CBCTs (pCTs$+$artifacts). Specifically, for the in silico data augmentation, we make use of the critical insight that CT and CBCT have the same underlying physics and that it is easier to deteriorate the pCT to look more like CBCT (and use the accompanying high quality manual contours for segmentation) than to synthesize CT from CBCT where the critical anatomical information may have already been lost (which leads to anatomical hallucination with the prevalent generative adversarial networks for example). Given these pseudo-CBCTs and the high quality manual contours, we introduce a modified 3D-Unet architecture and a multi-objective loss function specifically designed for segmenting soft-tissue organs such as esophagus on real weekly CBCTs. The model achieved 0.74 dice overlap (against manual contours of an experienced radiation oncologist) on weekly CBCTs and was robust and generalizable enough to also produce state-of-the-art results on pCTs, achieving 0.77 dice overlap against the previous best of 0.72. This shows that our in silico data augmentation spans the realistic noise/artifact spectrum across patient CBCT/pCT data and can generalize well across modalities (without requiring retraining or domain adaptation), eventually improving the accuracy of treatment setup and response analysis.
Cross-modality (CT-MRI) prior augmented deep learning for robust lung tumor segmentation from small MR datasets
Feb 27, 2019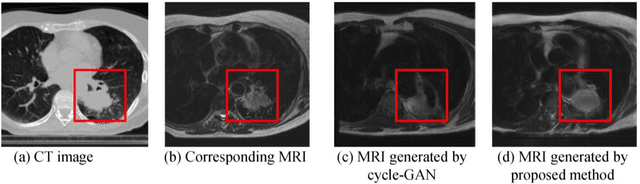
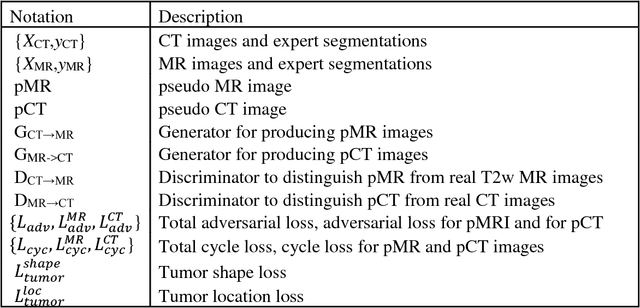

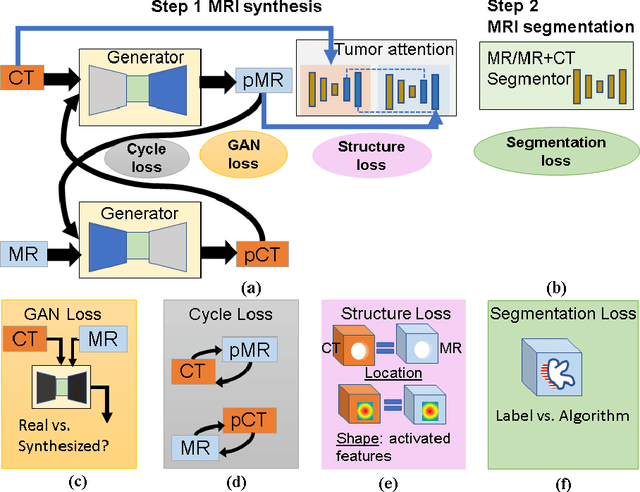
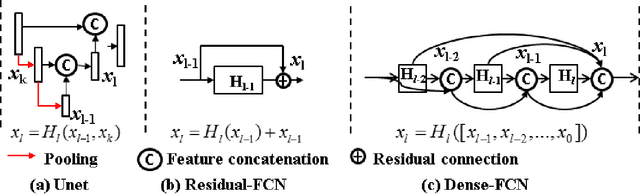
Abstract:Lack of large expert annotated MR datasets makes training deep learning models difficult. Therefore, a cross-modality (MR-CT) deep learning segmentation approach that augments training data using pseudo MR images produced by transforming expert-segmented CT images was developed. Eighty-One T2-weighted MRI scans from 28 patients with non-small cell lung cancers were analyzed. Cross-modality prior encoding the transformation of CT to pseudo MR images resembling T2w MRI was learned as a generative adversarial deep learning model. This model augmented training data arising from 6 expert-segmented T2w MR patient scans with 377 pseudo MRI from non-small cell lung cancer CT patient scans with obtained from the Cancer Imaging Archive. A two-dimensional Unet implemented with batch normalization was trained to segment the tumors from T2w MRI. This method was benchmarked against (a) standard data augmentation and two state-of-the art cross-modality pseudo MR-based augmentation and (b) two segmentation networks. Segmentation accuracy was computed using Dice similarity coefficient (DSC), Hausdroff distance metrics, and volume ratio. The proposed approach produced the lowest statistical variability in the intensity distribution between pseudo and T2w MR images measured as Kullback-Leibler divergence of 0.069. This method produced the highest segmentation accuracy with a DSC of 0.75 and the lowest Hausdroff distance on the test dataset. This approach produced highly similar estimations of tumor growth as an expert (P = 0.37). A novel deep learning MR segmentation was developed that overcomes the limitation of learning robust models from small datasets by leveraging learned cross-modality priors to augment training. The results show the feasibility of the approach and the corresponding improvement over the state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge