Tianfang Li
Spatiotemporal Gaussian Optimization for 4D Cone Beam CT Reconstruction from Sparse Projections
Jan 07, 2025
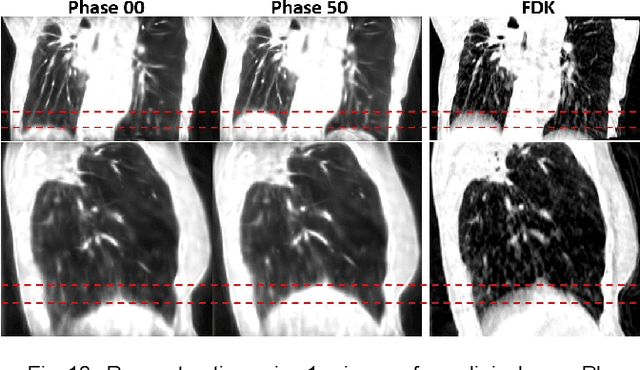
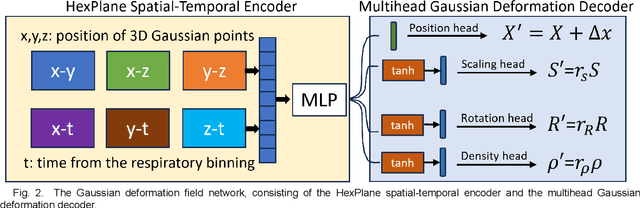
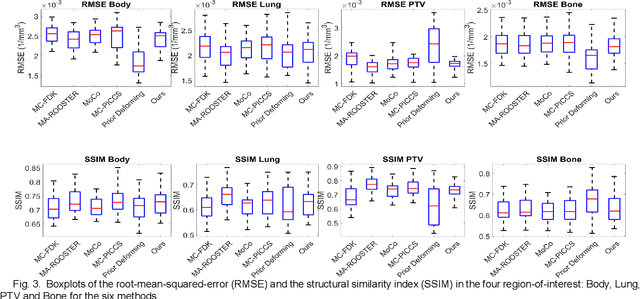
Abstract:In image-guided radiotherapy (IGRT), four-dimensional cone-beam computed tomography (4D-CBCT) is critical for assessing tumor motion during a patients breathing cycle prior to beam delivery. However, generating 4D-CBCT images with sufficient quality requires significantly more projection images than a standard 3D-CBCT scan, leading to extended scanning times and increased imaging dose to the patient. To address these limitations, there is a strong demand for methods capable of reconstructing high-quality 4D-CBCT images from a 1-minute 3D-CBCT acquisition. The challenge lies in the sparse sampling of projections, which introduces severe streaking artifacts and compromises image quality. This paper introduces a novel framework leveraging spatiotemporal Gaussian representation for 4D-CBCT reconstruction from sparse projections, achieving a balance between streak artifact reduction, dynamic motion preservation, and fine detail restoration. Each Gaussian is characterized by its 3D position, covariance, rotation, and density. Two-dimensional X-ray projection images can be rendered from the Gaussian point cloud representation via X-ray rasterization. The properties of each Gaussian were optimized by minimizing the discrepancy between the measured projections and the rendered X-ray projections. A Gaussian deformation network is jointly optimized to deform these Gaussian properties to obtain a 4D Gaussian representation for dynamic CBCT scene modeling. The final 4D-CBCT images are reconstructed by voxelizing the 4D Gaussians, achieving a high-quality representation that preserves both motion dynamics and spatial detail. The code and reconstruction results can be found at https://github.com/fuyabo/4DGS_for_4DCBCT/tree/main
Generalizable Cone Beam CT Esophagus Segmentation Using In Silico Data Augmentation
Jun 28, 2020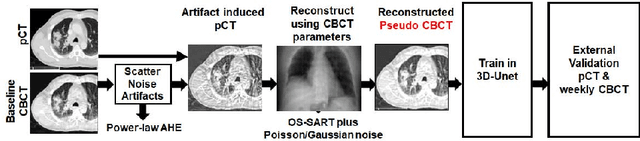
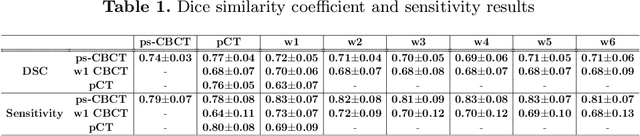
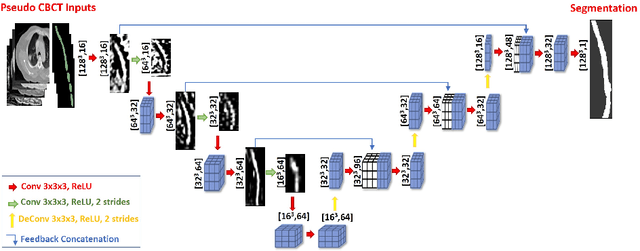

Abstract:Lung cancer radiotherapy entails high quality planning computed tomography (pCT) imaging of the patient with radiation oncologist contouring of the tumor and the organs at risk (OARs) at the start of the treatment. This is followed by weekly low-quality cone beam CT (CBCT) imaging for treatment setup and qualitative visual assessment of tumor and critical OARs. In this work, we aim to make the weekly CBCT assessment quantitative by automatically segmenting the most critical OAR, esophagus, using deep learning and in silico (image-driven simulation) artifact induction to convert pCTs to pseudo-CBCTs (pCTs$+$artifacts). Specifically, for the in silico data augmentation, we make use of the critical insight that CT and CBCT have the same underlying physics and that it is easier to deteriorate the pCT to look more like CBCT (and use the accompanying high quality manual contours for segmentation) than to synthesize CT from CBCT where the critical anatomical information may have already been lost (which leads to anatomical hallucination with the prevalent generative adversarial networks for example). Given these pseudo-CBCTs and the high quality manual contours, we introduce a modified 3D-Unet architecture and a multi-objective loss function specifically designed for segmenting soft-tissue organs such as esophagus on real weekly CBCTs. The model achieved 0.74 dice overlap (against manual contours of an experienced radiation oncologist) on weekly CBCTs and was robust and generalizable enough to also produce state-of-the-art results on pCTs, achieving 0.77 dice overlap against the previous best of 0.72. This shows that our in silico data augmentation spans the realistic noise/artifact spectrum across patient CBCT/pCT data and can generalize well across modalities (without requiring retraining or domain adaptation), eventually improving the accuracy of treatment setup and response analysis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge