Oskar Wysocki
Department of Computer Science, The University of Manchester, digital Experimental Cancer Medicine Team, Cancer Biomarker Centre, CRUK Manchester Institute, University of Manchester
Integrating Expert Knowledge into Logical Programs via LLMs
Feb 17, 2025Abstract:This paper introduces ExKLoP, a novel framework designed to evaluate how effectively Large Language Models (LLMs) integrate expert knowledge into logical reasoning systems. This capability is especially valuable in engineering, where expert knowledge-such as manufacturer-recommended operational ranges-can be directly embedded into automated monitoring systems. By mirroring expert verification steps, tasks like range checking and constraint validation help ensure system safety and reliability. Our approach systematically evaluates LLM-generated logical rules, assessing both syntactic fluency and logical correctness in these critical validation tasks. We also explore the models capacity for self-correction via an iterative feedback loop based on code execution outcomes. ExKLoP presents an extensible dataset comprising 130 engineering premises, 950 prompts, and corresponding validation points. It enables comprehensive benchmarking while allowing control over task complexity and scalability of experiments. We leverage the synthetic data creation methodology to conduct extensive empirical evaluation on a diverse set of LLMs including Llama3, Gemma, Mixtral, Mistral, and Qwen. Results reveal that while models generate nearly perfect syntactically correct code, they frequently exhibit logical errors in translating expert knowledge. Furthermore, iterative self-correction yields only marginal improvements (up to 3%). Overall, ExKLoP serves as a robust evaluation platform that streamlines the selection of effective models for self-correcting systems while clearly delineating the types of errors encountered. The complete implementation, along with all relevant data, is available at GitHub.
SylloBio-NLI: Evaluating Large Language Models on Biomedical Syllogistic Reasoning
Oct 18, 2024



Abstract:Syllogistic reasoning is crucial for Natural Language Inference (NLI). This capability is particularly significant in specialized domains such as biomedicine, where it can support automatic evidence interpretation and scientific discovery. This paper presents SylloBio-NLI, a novel framework that leverages external ontologies to systematically instantiate diverse syllogistic arguments for biomedical NLI. We employ SylloBio-NLI to evaluate Large Language Models (LLMs) on identifying valid conclusions and extracting supporting evidence across 28 syllogistic schemes instantiated with human genome pathways. Extensive experiments reveal that biomedical syllogistic reasoning is particularly challenging for zero-shot LLMs, which achieve an average accuracy between 70% on generalized modus ponens and 23% on disjunctive syllogism. At the same time, we found that few-shot prompting can boost the performance of different LLMs, including Gemma (+14%) and LLama-3 (+43%). However, a deeper analysis shows that both techniques exhibit high sensitivity to superficial lexical variations, highlighting a dependency between reliability, models' architecture, and pre-training regime. Overall, our results indicate that, while in-context examples have the potential to elicit syllogistic reasoning in LLMs, existing models are still far from achieving the robustness and consistency required for safe biomedical NLI applications.
An LLM-based Knowledge Synthesis and Scientific Reasoning Framework for Biomedical Discovery
Jun 26, 2024
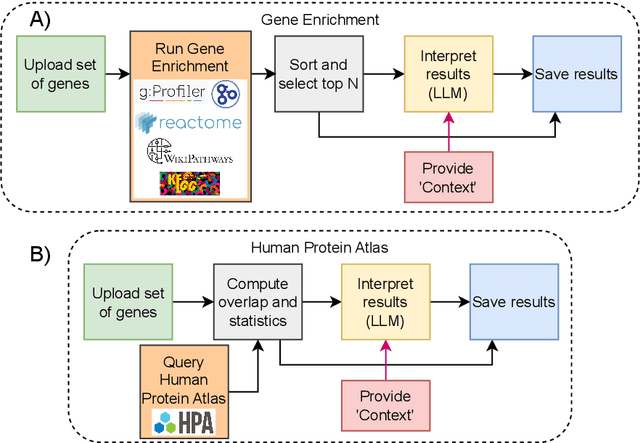
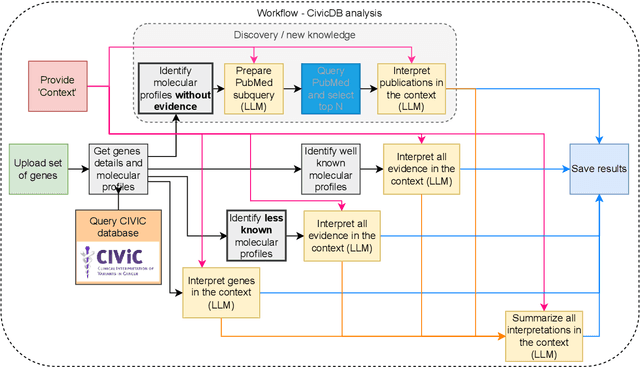
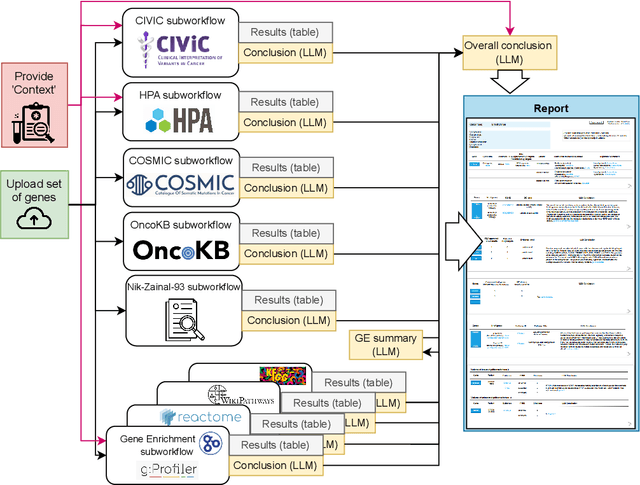
Abstract:We present BioLunar, developed using the Lunar framework, as a tool for supporting biological analyses, with a particular emphasis on molecular-level evidence enrichment for biomarker discovery in oncology. The platform integrates Large Language Models (LLMs) to facilitate complex scientific reasoning across distributed evidence spaces, enhancing the capability for harmonizing and reasoning over heterogeneous data sources. Demonstrating its utility in cancer research, BioLunar leverages modular design, reusable data access and data analysis components, and a low-code user interface, enabling researchers of all programming levels to construct LLM-enabled scientific workflows. By facilitating automatic scientific discovery and inference from heterogeneous evidence, BioLunar exemplifies the potential of the integration between LLMs, specialised databases and biomedical tools to support expert-level knowledge synthesis and discovery.
Large Language Models, scientific knowledge and factuality: A systematic analysis in antibiotic discovery
May 28, 2023Abstract:Inferring over and extracting information from Large Language Models (LLMs) trained on a large corpus of scientific literature can potentially drive a new era in biomedical research, reducing the barriers for accessing existing medical evidence. This work examines the potential of LLMs for dialoguing with biomedical background knowledge, using the context of antibiotic discovery as an exemplar motivational scenario. The context of biomedical discovery from natural products entails understanding the relational evidence between an organism, an associated chemical and its associated antibiotic properties. We provide a systematic assessment on the ability of LLMs to encode and express these relations, verifying for fluency, prompt-alignment, semantic coherence, factual knowledge and specificity of generated responses. The systematic analysis is applied to nine state-of-the-art models (including ChatGPT and GPT-4) in two prompting-based tasks: chemical compound definition generation and chemical compound-fungus relation determination. Results show that while recent models have improved in fluency, factual accuracy is still low and models are biased towards over-represented entities. The ability of LLMs to serve as biomedical knowledge bases is questioned, and the need for additional systematic evaluation frameworks is highlighted. The best performing GPT-4 produced a factual definition for 70% of chemical compounds and 43.6% factual relations to fungi, whereas the best open source model BioGPT-large 30% of the compounds and 30% of the relations for the best-performing prompt. The results show that while LLMs are currently not fit for purpose to be used as biomedical factual knowledge bases, there is a promising emerging property in the direction of factuality as the models become domain specialised, scale-up in size and level of human feedback.
On the Visualisation of Argumentation Graphs to Support Text Interpretation
Mar 06, 2023Abstract:The recent evolution in Natural Language Processing (NLP) methods, in particular in the field of argumentation mining, has the potential to transform the way we interact with text, supporting the interpretation and analysis of complex discourse and debates. Can a graphic visualisation of complex argumentation enable a more critical interpretation of the arguments? This study focuses on analysing the impact of argumentation graphs (AGs) compared with regular texts for supporting argument interpretation. We found that AGs outperformed the extrinsic metrics throughout most UEQ scales as well as the NASA-TLX workload in all the terms but not in temporal or physical demand. The AG model was liked by a more significant number of participants, despite the fact that both the text-based and AG models yielded comparable outcomes in the critical interpretation in terms of working memory and altering participants decisions. The interpretation process involves reference to argumentation schemes (linked to critical questions (CQs)) in AGs. Interestingly, we found that the participants chose more CQs (using argument schemes in AGs) when they were less familiar with the argument topics, making AG schemes on some scales (relatively) supportive of the interpretation process. Therefore, AGs were considered to deliver a more critical approach to argument interpretation, especially with unfamiliar topics. Based on the 25 participants conducted in this study, it appears that AG has demonstrated an overall positive effect on the argument interpretation process.
Biologically-informed deep learning models for cancer: fundamental trends for encoding and interpreting oncology data
Jul 02, 2022



Abstract:In this paper we provide a structured literature analysis focused on Deep Learning (DL) models used to support inference in cancer biology with a particular emphasis on multi-omics analysis. The work focuses on how existing models address the need for better dialogue with prior knowledge, biological plausibility and interpretability, fundamental properties in the biomedical domain. We discuss the recent evolutionary arch of DL models in the direction of integrating prior biological relational and network knowledge to support better generalisation (e.g. pathways or Protein-Protein-Interaction networks) and interpretability. This represents a fundamental functional shift towards models which can integrate mechanistic and statistical inference aspects. We discuss representational methodologies for the integration of domain prior knowledge in such models. The paper also provides a critical outlook into contemporary methods for explainability and interpretabiltiy. This analysis points in the direction of a convergence between encoding prior knowledge and improved interpretability.
Metareview-informed Explainable Cytokine Storm Detection during CAR-T cell Therapy
Jun 20, 2022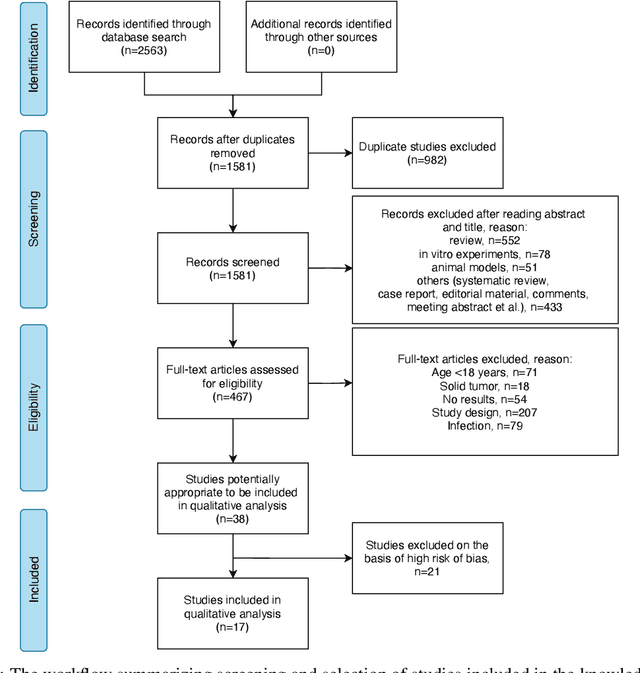
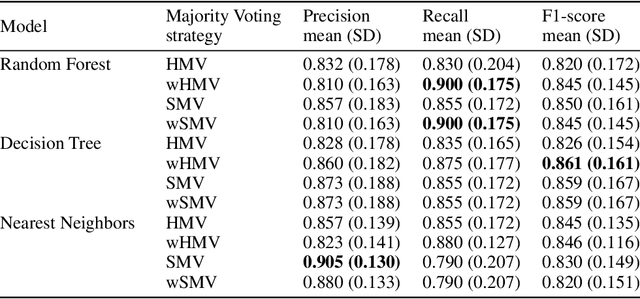
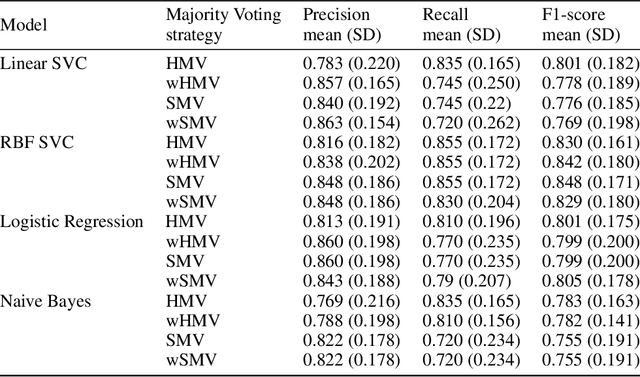
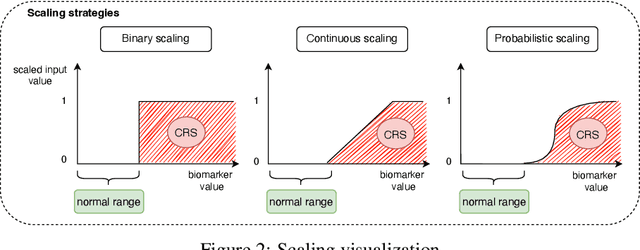
Abstract:Cytokine release syndrome (CRS), also known as cytokine storm, is one of the most consequential adverse effects of chimeric antigen receptor therapies that have shown promising results in cancer treatment. When emerging, CRS could be identified by the analysis of specific cytokine and chemokine profiles that tend to exhibit similarities across patients. In this paper, we exploit these similarities using machine learning algorithms and set out to pioneer a meta--review informed method for the identification of CRS based on specific cytokine peak concentrations and evidence from previous clinical studies. We argue that such methods could support clinicians in analyzing suspect cytokine profiles by matching them against CRS knowledge from past clinical studies, with the ultimate aim of swift CRS diagnosis. During evaluation with real--world CRS clinical data, we emphasize the potential of our proposed method of producing interpretable results, in addition to being effective in identifying the onset of cytokine storm.
Assessing the communication gap between AI models and healthcare professionals: explainability, utility and trust in AI-driven clinical decision-making
Apr 13, 2022



Abstract:This paper contributes with a pragmatic evaluation framework for explainable Machine Learning (ML) models for clinical decision support. The study revealed a more nuanced role for ML explanation models, when these are pragmatically embedded in the clinical context. Despite the general positive attitude of healthcare professionals (HCPs) towards explanations as a safety and trust mechanism, for a significant set of participants there were negative effects associated with confirmation bias, accentuating model over-reliance and increased effort to interact with the model. Also, contradicting one of its main intended functions, standard explanatory models showed limited ability to support a critical understanding of the limitations of the model. However, we found new significant positive effects which repositions the role of explanations within a clinical context: these include reduction of automation bias, addressing ambiguous clinical cases (cases where HCPs were not certain about their decision) and support of less experienced HCPs in the acquisition of new domain knowledge.
Transformers and the representation of biomedical background knowledge
Feb 04, 2022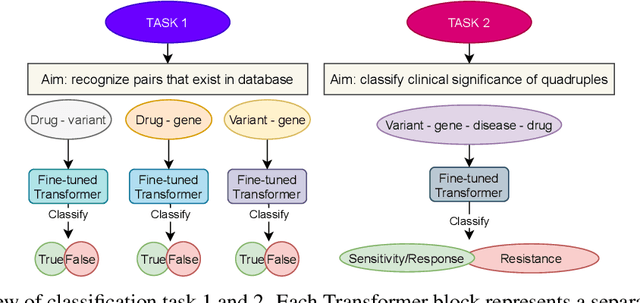

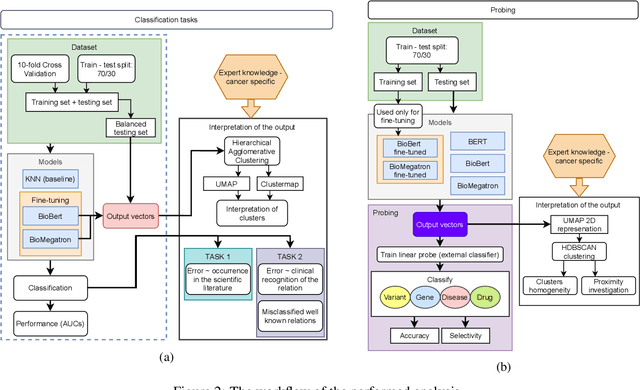
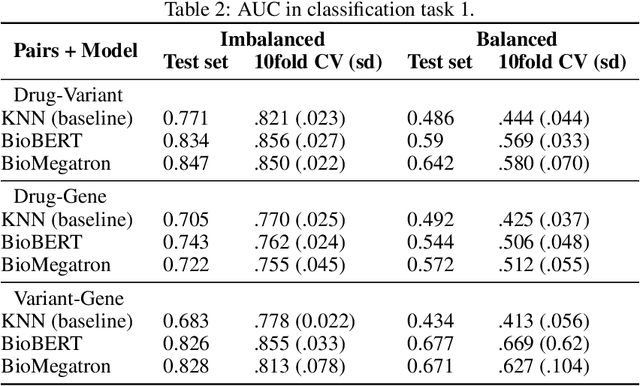
Abstract:BioBERT and BioMegatron are Transformers models adapted for the biomedical domain based on publicly available biomedical corpora. As such, they have the potential to encode large-scale biological knowledge. We investigate the encoding and representation of biological knowledge in these models, and its potential utility to support inference in cancer precision medicine - namely, the interpretation of the clinical significance of genomic alterations. We compare the performance of different transformer baselines; we use probing to determine the consistency of encodings for distinct entities; and we use clustering methods to compare and contrast the internal properties of the embeddings for genes, variants, drugs and diseases. We show that these models do indeed encode biological knowledge, although some of this is lost in fine-tuning for specific tasks. Finally, we analyse how the models behave with regard to biases and imbalances in the dataset.
Architectures of Meaning, A Systematic Corpus Analysis of NLP Systems
Jul 16, 2021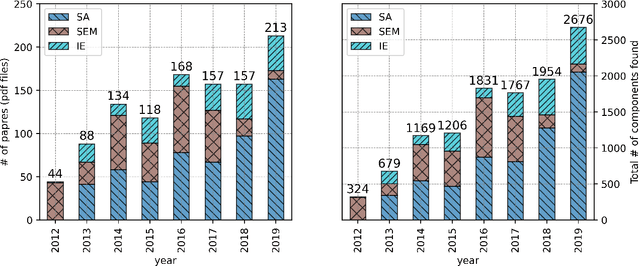
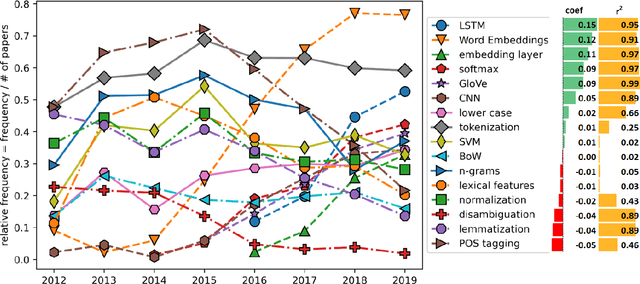
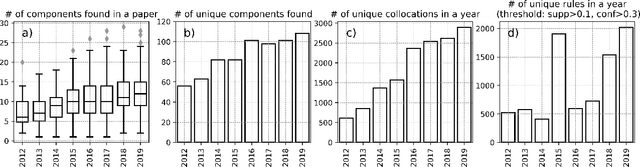
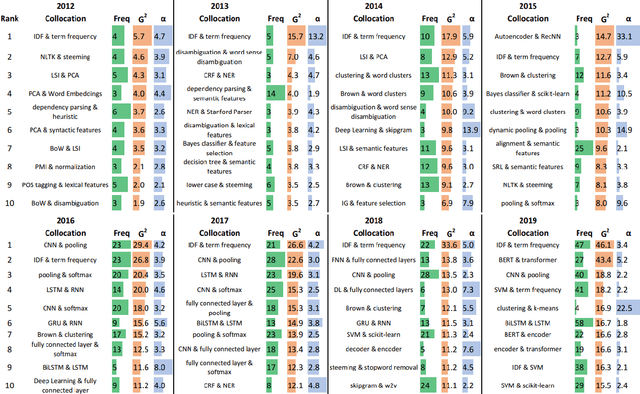
Abstract:This paper proposes a novel statistical corpus analysis framework targeted towards the interpretation of Natural Language Processing (NLP) architectural patterns at scale. The proposed approach combines saturation-based lexicon construction, statistical corpus analysis methods and graph collocations to induce a synthesis representation of NLP architectural patterns from corpora. The framework is validated in the full corpus of Semeval tasks and demonstrated coherent architectural patterns which can be used to answer architectural questions on a data-driven fashion, providing a systematic mechanism to interpret a largely dynamic and exponentially growing field.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge