Nicholas Tustison
QU-BraTS: MICCAI BraTS 2020 Challenge on Quantifying Uncertainty in Brain Tumor Segmentation -- Analysis of Ranking Metrics and Benchmarking Results
Dec 19, 2021
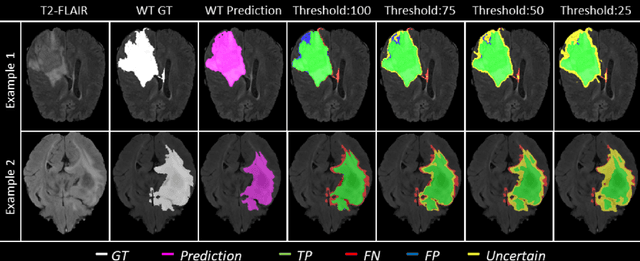
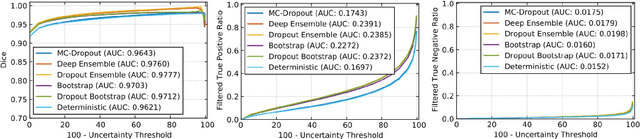
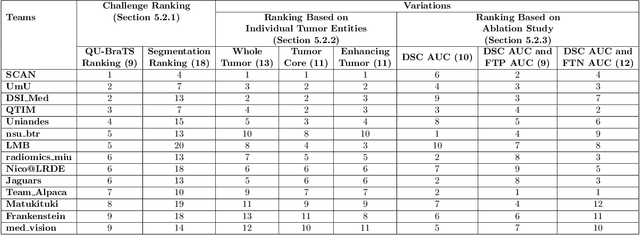
Abstract:Deep learning (DL) models have provided the state-of-the-art performance in a wide variety of medical imaging benchmarking challenges, including the Brain Tumor Segmentation (BraTS) challenges. However, the task of focal pathology multi-compartment segmentation (e.g., tumor and lesion sub-regions) is particularly challenging, and potential errors hinder the translation of DL models into clinical workflows. Quantifying the reliability of DL model predictions in the form of uncertainties, could enable clinical review of the most uncertain regions, thereby building trust and paving the way towards clinical translation. Recently, a number of uncertainty estimation methods have been introduced for DL medical image segmentation tasks. Developing metrics to evaluate and compare the performance of uncertainty measures will assist the end-user in making more informed decisions. In this study, we explore and evaluate a metric developed during the BraTS 2019-2020 task on uncertainty quantification (QU-BraTS), and designed to assess and rank uncertainty estimates for brain tumor multi-compartment segmentation. This metric (1) rewards uncertainty estimates that produce high confidence in correct assertions, and those that assign low confidence levels at incorrect assertions, and (2) penalizes uncertainty measures that lead to a higher percentages of under-confident correct assertions. We further benchmark the segmentation uncertainties generated by 14 independent participating teams of QU-BraTS 2020, all of which also participated in the main BraTS segmentation task. Overall, our findings confirm the importance and complementary value that uncertainty estimates provide to segmentation algorithms, and hence highlight the need for uncertainty quantification in medical image analyses. Our evaluation code is made publicly available at https://github.com/RagMeh11/QU-BraTS.
Brain Tumor Segmentation using an Ensemble of 3D U-Nets and Overall Survival Prediction using Radiomic Features
Dec 03, 2018
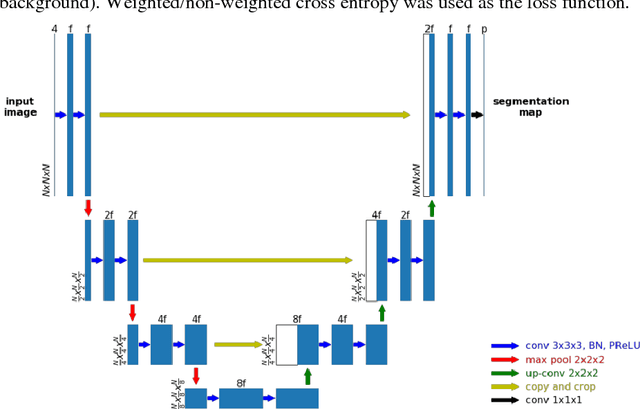
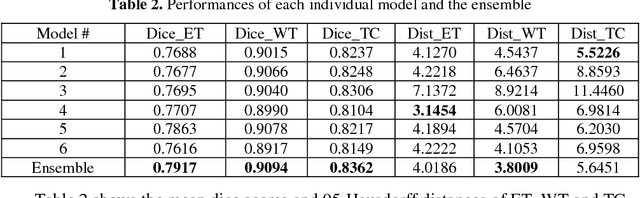
Abstract:Accurate segmentation of different sub-regions of gliomas including peritumoral edema, necrotic core, enhancing and non-enhancing tumor core from multimodal MRI scans has important clinical relevance in diagnosis, prognosis and treatment of brain tumors. However, due to the highly heterogeneous appearance and shape, segmentation of the sub-regions is very challenging. Recent development using deep learning models has proved its effectiveness in the past several brain segmentation challenges as well as other semantic and medical image segmentation problems. Most models in brain tumor segmentation use a 2D/3D patch to predict the class label for the center voxel and variant patch sizes and scales are used to improve the model performance. However, it has low computation efficiency and also has limited receptive field. U-Net is a widely used network structure for end-to-end segmentation and can be used on the entire image or extracted patches to provide classification labels over the entire input voxels so that it is more efficient and expect to yield better performance with larger input size. Furthermore, instead of picking the best network structure, an ensemble of multiple models, trained on different dataset or different hyper-parameters, can generally improve the segmentation performance. In this study we propose to use an ensemble of 3D U-Nets with different hyper-parameters for brain tumor segmentation. Preliminary results showed effectiveness of this model. In addition, we developed a linear model for survival prediction using extracted imaging and non-imaging features, which, despite the simplicity, can effectively reduce overfitting and regression errors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge