Mohammad Bakhtiari
Privacy-Preserving Multi-Center Differential Protein Abundance Analysis with FedProt
Jul 21, 2024



Abstract:Quantitative mass spectrometry has revolutionized proteomics by enabling simultaneous quantification of thousands of proteins. Pooling patient-derived data from multiple institutions enhances statistical power but raises significant privacy concerns. Here we introduce FedProt, the first privacy-preserving tool for collaborative differential protein abundance analysis of distributed data, which utilizes federated learning and additive secret sharing. In the absence of a multicenter patient-derived dataset for evaluation, we created two, one at five centers from LFQ E.coli experiments and one at three centers from TMT human serum. Evaluations using these datasets confirm that FedProt achieves accuracy equivalent to DEqMS applied to pooled data, with completely negligible absolute differences no greater than $\text{$4 \times 10^{-12}$}$. In contrast, -log10(p-values) computed by the most accurate meta-analysis methods diverged from the centralized analysis results by up to 25-27. FedProt is available as a web tool with detailed documentation as a FeatureCloud App.
The FeatureCloud AI Store for Federated Learning in Biomedicine and Beyond
May 12, 2021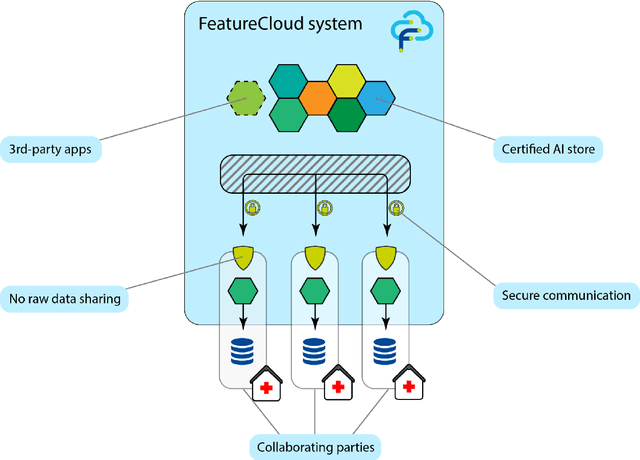
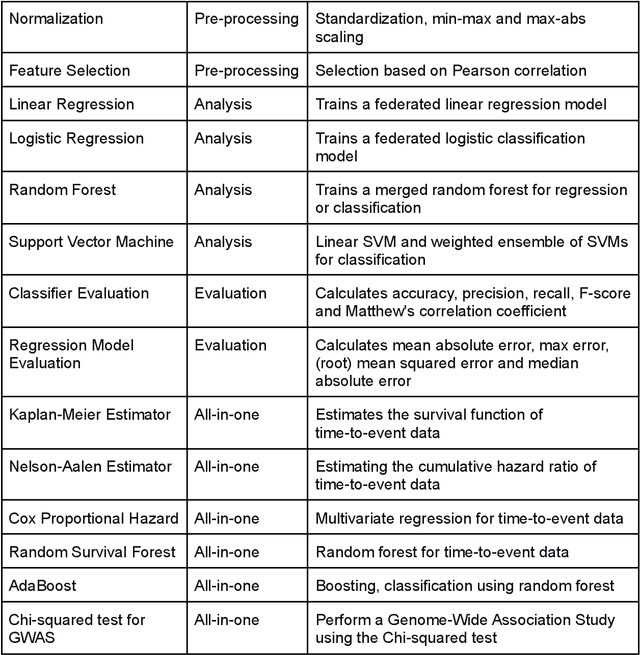
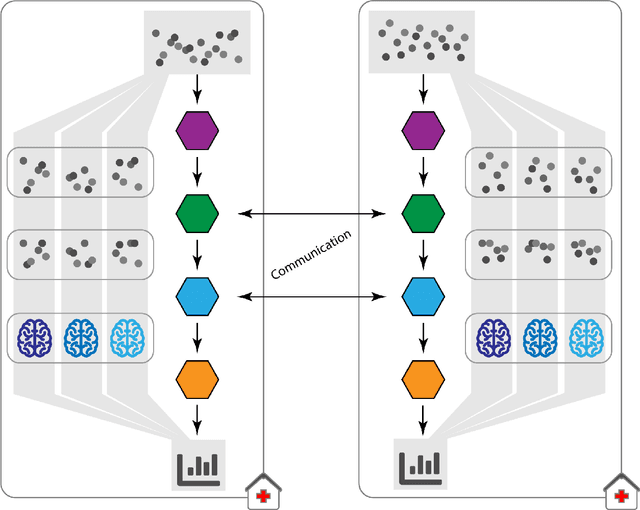
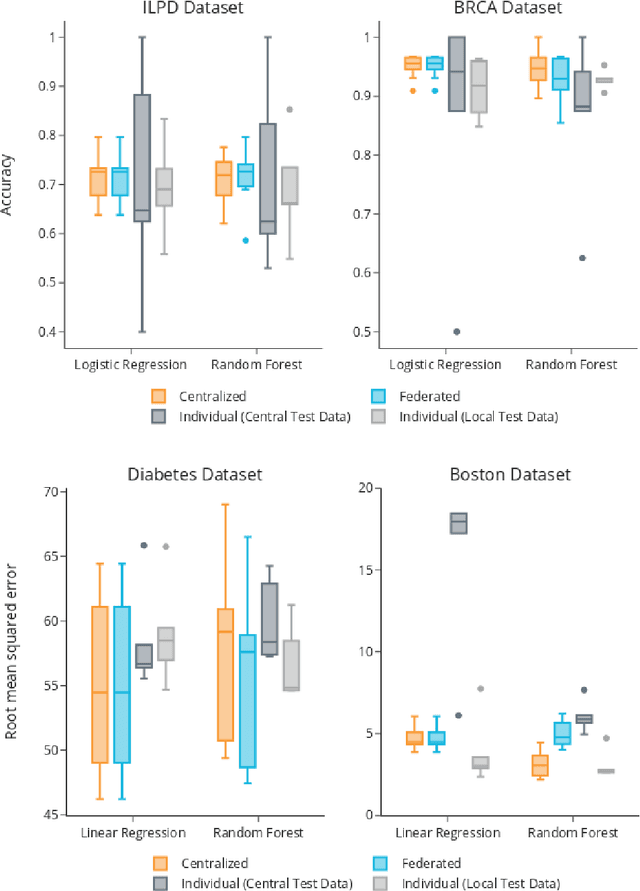
Abstract:Machine Learning (ML) and Artificial Intelligence (AI) have shown promising results in many areas and are driven by the increasing amount of available data. However, this data is often distributed across different institutions and cannot be shared due to privacy concerns. Privacy-preserving methods, such as Federated Learning (FL), allow for training ML models without sharing sensitive data, but their implementation is time-consuming and requires advanced programming skills. Here, we present the FeatureCloud AI Store for FL as an all-in-one platform for biomedical research and other applications. It removes large parts of this complexity for developers and end-users by providing an extensible AI Store with a collection of ready-to-use apps. We show that the federated apps produce similar results to centralized ML, scale well for a typical number of collaborators and can be combined with Secure Multiparty Computation (SMPC), thereby making FL algorithms safely and easily applicable in biomedical and clinical environments.
Federated Multi-Mini-Batch: An Efficient Training Approach to Federated Learning in Non-IID Environments
Nov 13, 2020
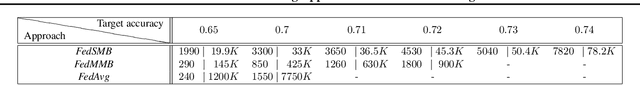
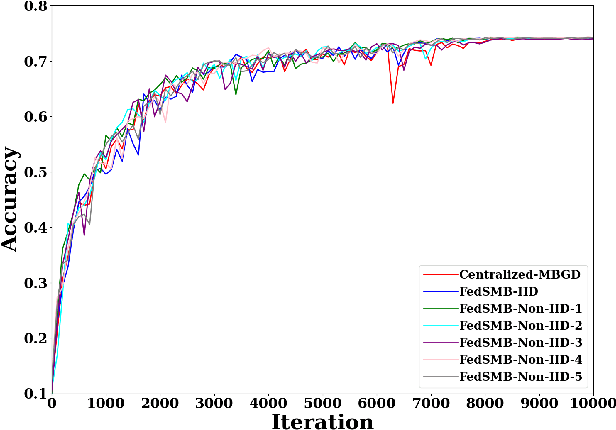
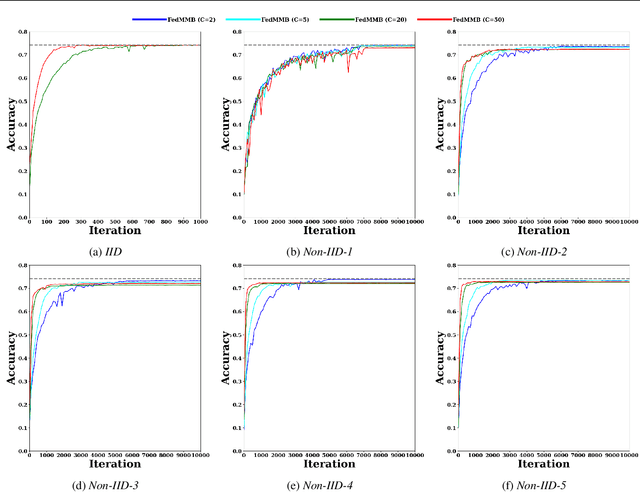
Abstract:Federated learning is a well-established approach to privacy-preserving training of a joint model on heavily distributed data. Federated averaging (FedAvg) is a well-known communication-efficient algorithm for federated learning, which performs well if the data distribution across the clients is independently and identically distributed (IID). However, FedAvg provides a lower accuracy and still requires a large number of communication rounds to achieve a target accuracy when it comes to Non-IID environments. To address the former limitation, we present federated single mini-batch (FedSMB), where the clients train the model on a single mini-batch from their dataset in each iteration. We show that FedSMB achieves the accuracy of the centralized training in Non-IID configurations, but in a considerable number of iterations. To address the latter limitation, we introduce federated multi-mini-batch (FedMMB) as a generalization of FedSMB, where the clients train the model on multiple mini-batches (specified by the batch count) in each communication round. FedMMB decouples the batch size from the batch count and provides a trade-off between the accuracy and communication efficiency in Non-IID settings. This is not possible with FedAvg, in which a single parameter determines both the batch size and batch count. The simulation results illustrate that FedMMB outperforms FedAvg in terms of the accuracy, communication efficiency, as well as computational efficiency and is an efficient training approach to federated learning in Non-IID environments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge