Michelle Williams
MorphiNet: A Graph Subdivision Network for Adaptive Bi-ventricle Surface Reconstruction
Dec 14, 2024



Abstract:Cardiac Magnetic Resonance (CMR) imaging is widely used for heart modelling and digital twin computational analysis due to its ability to visualize soft tissues and capture dynamic functions. However, the anisotropic nature of CMR images, characterized by large inter-slice distances and misalignments from cardiac motion, poses significant challenges to accurate model reconstruction. These limitations result in data loss and measurement inaccuracies, hindering the capture of detailed anatomical structures. This study introduces MorphiNet, a novel network that enhances heart model reconstruction by leveraging high-resolution Computer Tomography (CT) images, unpaired with CMR images, to learn heart anatomy. MorphiNet encodes anatomical structures as gradient fields, transforming template meshes into patient-specific geometries. A multi-layer graph subdivision network refines these geometries while maintaining dense point correspondence. The proposed method achieves high anatomy fidelity, demonstrating approximately 40% higher Dice scores, half the Hausdorff distance, and around 3 mm average surface error compared to state-of-the-art methods. MorphiNet delivers superior results with greater inference efficiency. This approach represents a significant advancement in addressing the challenges of CMR-based heart model reconstruction, potentially improving digital twin computational analyses of cardiac structure and functions.
Factorised Representation Learning in Cardiac Image Analysis
Mar 22, 2019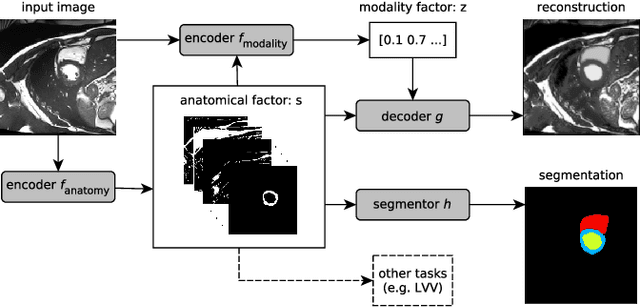
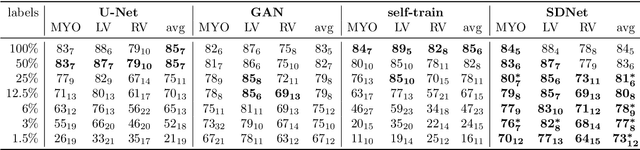
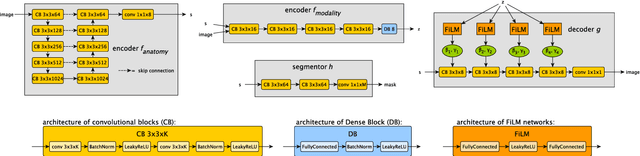
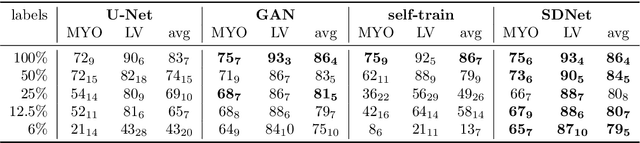
Abstract:Typically, a medical image offers spatial information on the anatomy (and pathology) modulated by imaging specific characteristics. Many imaging modalities including Magnetic Resonance Imaging (MRI) and Computed Tomography (CT) can be interpreted in this way. We can venture further and consider that a medical image naturally factors into some spatial factors depicting anatomy and factors that denote the imaging characteristics. Here, we explicitly learn this decomposed (factorised) representation of imaging data, focusing in particular on cardiac images. We propose Spatial Decomposition Network (SDNet), which factorises 2D medical images into spatial anatomical factors and non-spatial imaging factors. We demonstrate that this high-level representation is ideally suited for several medical image analysis tasks, such as semi-supervised segmentation, multi-task segmentation and regression, and image-to-image synthesis. Specifically, we show that our model can match the performance of fully supervised segmentation models, using only a fraction of the labelled images. Critically, we show that our factorised representation also benefits from supervision obtained either when we use auxiliary tasks to train the model in a multi-task setting (e.g. regressing to known cardiac indices), or when aggregating multimodal data from different sources (e.g. pooling together MRI and CT data). To explore the properties of the learned factorisation, we perform latent-space arithmetic and show that we can synthesise CT from MR and vice versa, by swapping the modality factors. We also demonstrate that the factor holding image specific information can be used to predict the input modality with high accuracy.
Factorised spatial representation learning: application in semi-supervised myocardial segmentation
Nov 02, 2018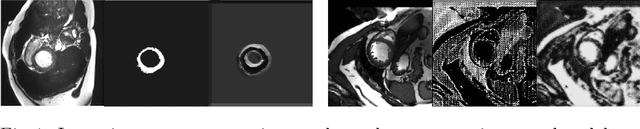
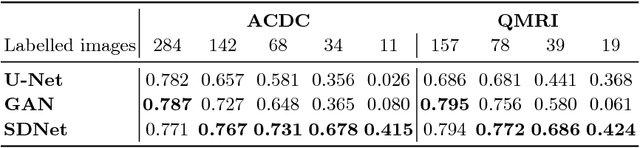
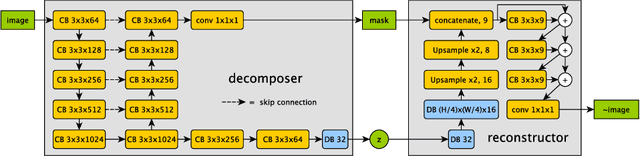
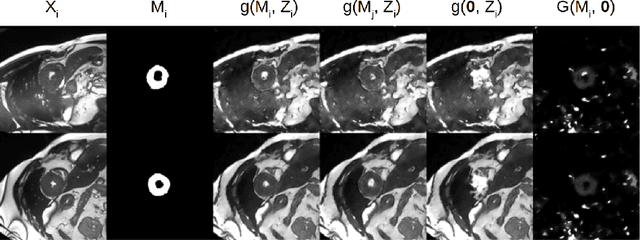
Abstract:The success and generalisation of deep learning algorithms heavily depend on learning good feature representations. In medical imaging this entails representing anatomical information, as well as properties related to the specific imaging setting. Anatomical information is required to perform further analysis, whereas imaging information is key to disentangle scanner variability and potential artefacts. The ability to factorise these would allow for training algorithms only on the relevant information according to the task. To date, such factorisation has not been attempted. In this paper, we propose a methodology of latent space factorisation relying on the cycle-consistency principle. As an example application, we consider cardiac MR segmentation, where we separate information related to the myocardium from other features related to imaging and surrounding substructures. We demonstrate the proposed method's utility in a semi-supervised setting: we use very few labelled images together with many unlabelled images to train a myocardium segmentation neural network. Specifically, we achieve comparable performance to fully supervised networks using a fraction of labelled images in experiments on ACDC and a dataset from Edinburgh Imaging Facility QMRI. Code will be made available at https://github.com/agis85/spatial_factorisation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge