Michelle Noga
Accelerated 3D-3D rigid registration of echocardiographic images obtained from apical window using particle filter
Apr 28, 2025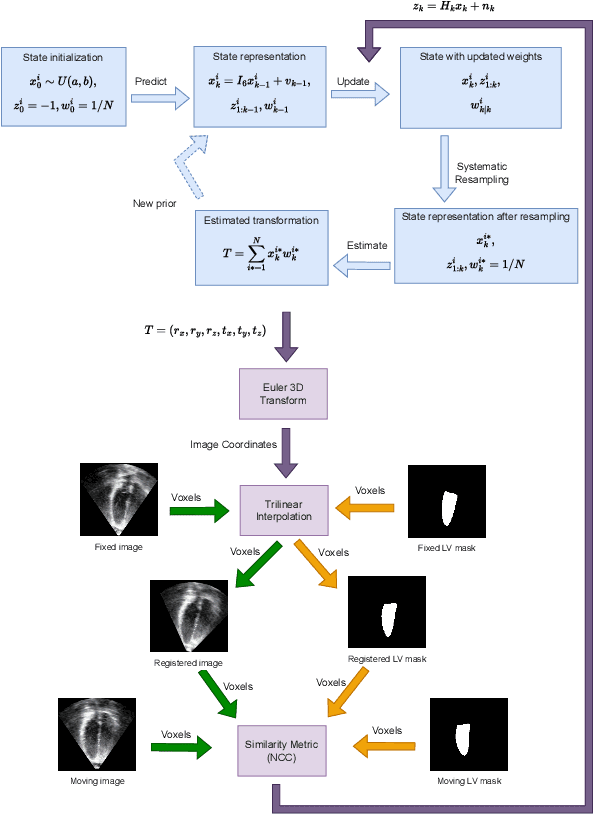



Abstract:The perfect alignment of 3D echocardiographic images captured from various angles has improved image quality and broadened the field of view. This study proposes an accelerated sequential Monte Carlo (SMC) algorithm for 3D-3D rigid registration of transthoracic echocardiographic images with significant and limited overlap taken from apical window that is robust to the noise and intensity variation in ultrasound images. The algorithm estimates the translational and rotational components of the rigid transform through an iterative process and requires an initial approximation of the rotation and translation limits. We perform registration in two ways: the image-based registration computes the transform to align the end-diastolic frame of the apical nonstandard image to the apical standard image and applies the same transform to all frames of the cardiac cycle, whereas the mask-based registration approach uses the binary masks of the left ventricle in the same way. The SMC and exhaustive search (EX) algorithms were evaluated for 4D temporal sequences recorded from 7 volunteers who participated in a study conducted at the Mazankowski Alberta Heart Institute. The evaluations demonstrate that the mask-based approach of the accelerated SMC yielded a Dice score value of 0.819 +/- 0.045 for the left ventricle and gained 16.7x speedup compared to the CPU version of the SMC algorithm.
Unsupervised diffeomorphic cardiac image registration using parameterization of the deformation field
Aug 28, 2022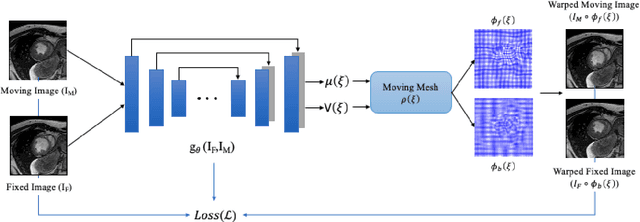

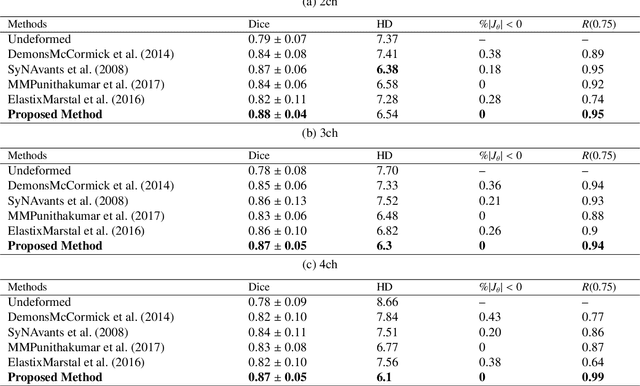
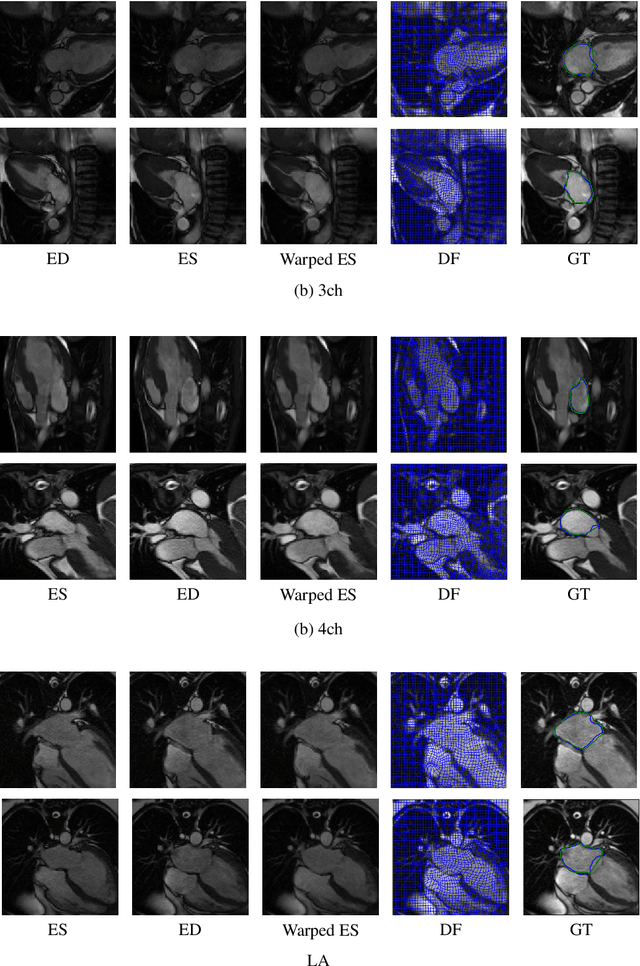
Abstract:This study proposes an end-to-end unsupervised diffeomorphic deformable registration framework based on moving mesh parameterization. Using this parameterization, a deformation field can be modeled with its transformation Jacobian determinant and curl of end velocity field. The new model of the deformation field has three important advantages; firstly, it relaxes the need for an explicit regularization term and the corresponding weight in the cost function. The smoothness is implicitly embedded in the solution which results in a physically plausible deformation field. Secondly, it guarantees diffeomorphism through explicit constraints applied to the transformation Jacobian determinant to keep it positive. Finally, it is suitable for cardiac data processing, since the nature of this parameterization is to define the deformation field in terms of the radial and rotational components. The effectiveness of the algorithm is investigated by evaluating the proposed method on three different data sets including 2D and 3D cardiac MRI scans. The results demonstrate that the proposed framework outperforms existing learning-based and non-learning-based methods while generating diffeomorphic transformations.
A training-free recursive multiresolution framework for diffeomorphic deformable image registration
Feb 01, 2022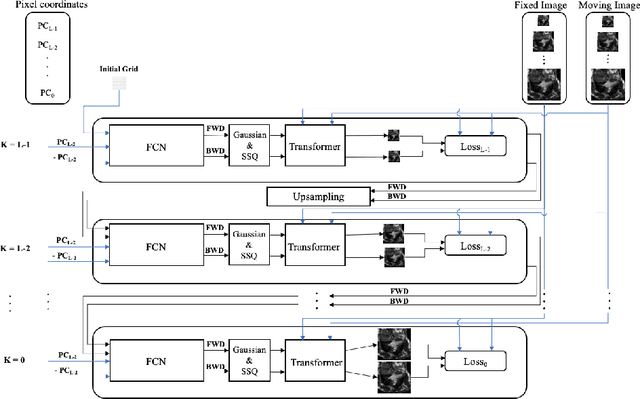
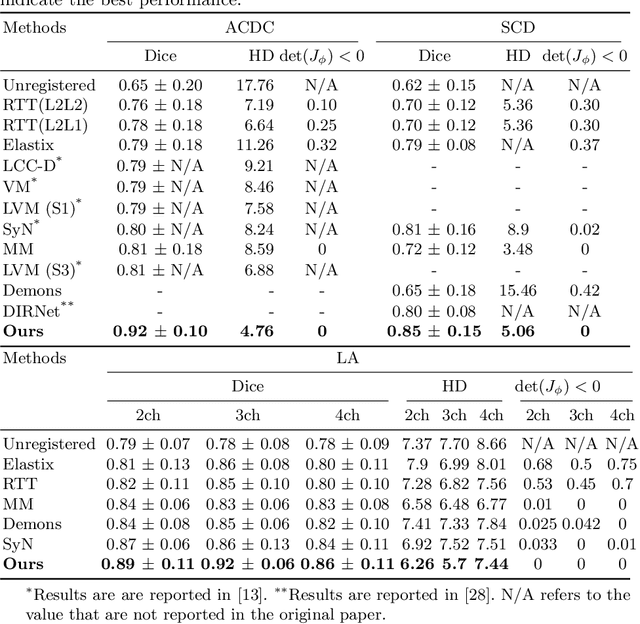
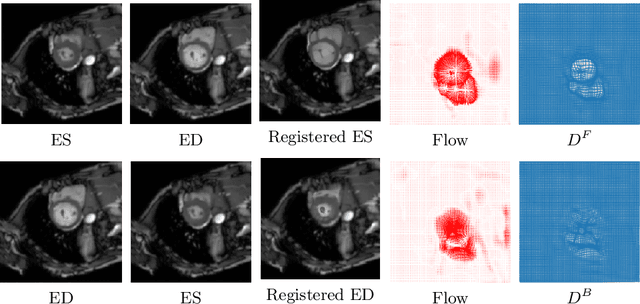

Abstract:Diffeomorphic deformable image registration is one of the crucial tasks in medical image analysis, which aims to find a unique transformation while preserving the topology and invertibility of the transformation. Deep convolutional neural networks (CNNs) have yielded well-suited approaches for image registration by learning the transformation priors from a large dataset. The improvement in the performance of these methods is related to their ability to learn information from several sample medical images that are difficult to obtain and bias the framework to the specific domain of data. In this paper, we propose a novel diffeomorphic training-free approach; this is built upon the principle of an ordinary differential equation. Our formulation yields an Euler integration type recursive scheme to estimate the changes of spatial transformations between the fixed and the moving image pyramids at different resolutions. The proposed architecture is simple in design. The moving image is warped successively at each resolution and finally aligned to the fixed image; this procedure is recursive in a way that at each resolution, a fully convolutional network (FCN) models a progressive change of deformation for the current warped image. The entire system is end-to-end and optimized for each pair of images from scratch. In comparison to learning-based methods, the proposed method neither requires a dedicated training set nor suffers from any training bias. We evaluate our method on three cardiac image datasets. The evaluation results demonstrate that the proposed method achieves state-of-the-art registration accuracy while maintaining desirable diffeomorphic properties.
A New Semi-Automated Algorithm for Volumetric Segmentation of the Left Ventricle in Temporal 3D Echocardiography Sequences
Sep 03, 2021
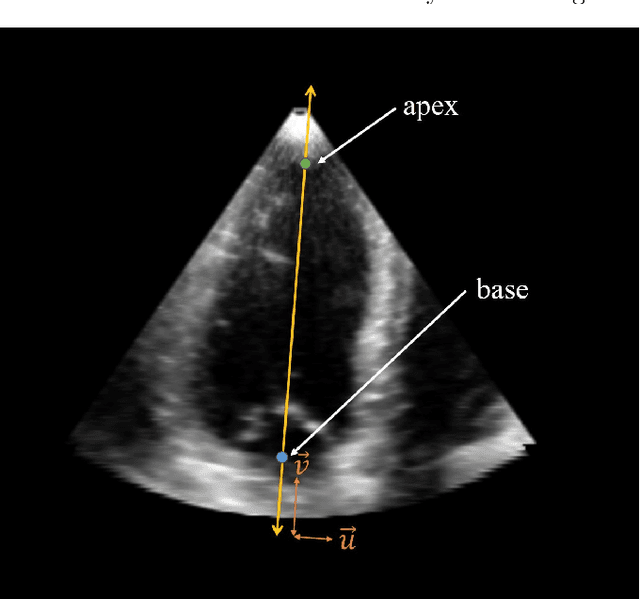
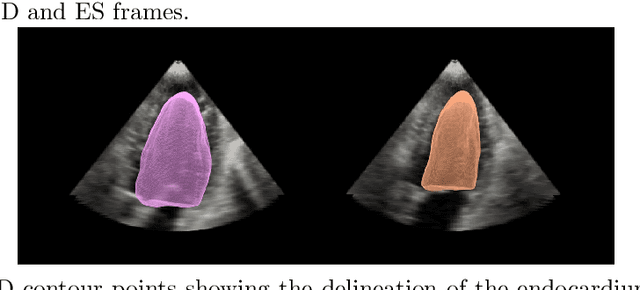
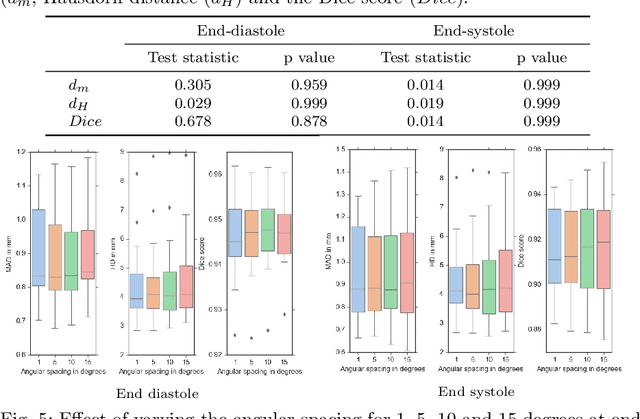
Abstract:Purpose: Echocardiography is commonly used as a non-invasive imaging tool in clinical practice for the assessment of cardiac function. However, delineation of the left ventricle is challenging due to the inherent properties of ultrasound imaging, such as the presence of speckle noise and the low signal-to-noise ratio. Methods: We propose a semi-automated segmentation algorithm for the delineation of the left ventricle in temporal 3D echocardiography sequences. The method requires minimal user interaction and relies on a diffeomorphic registration approach. Advantages of the method include no dependence on prior geometrical information, training data, or registration from an atlas. Results: The method was evaluated using three-dimensional ultrasound scan sequences from 18 patients from the Mazankowski Alberta Heart Institute, Edmonton, Canada, and compared to manual delineations provided by an expert cardiologist and four other registration algorithms. The segmentation approach yielded the following results over the cardiac cycle: a mean absolute difference of 1.01 (0.21) mm, a Hausdorff distance of 4.41 (1.43) mm, and a Dice overlap score of 0.93 (0.02). Conclusions: The method performed well compared to the four other registration algorithms.
* 22 pages, 8 figures
Fully Automated Left Atrium Segmentation from Anatomical Cine Long-axis MRI Sequences using Deep Convolutional Neural Network with Unscented Kalman Filter
Sep 28, 2020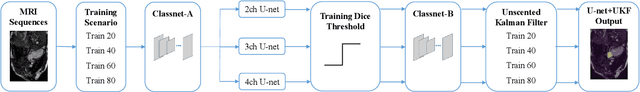
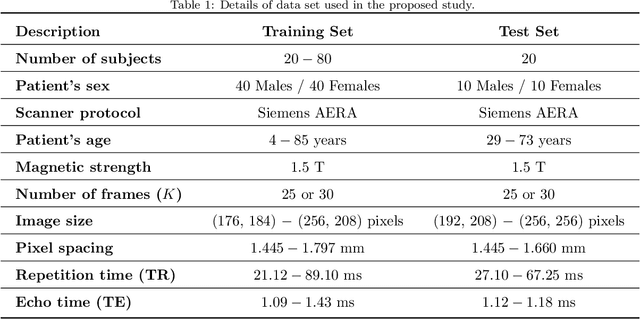
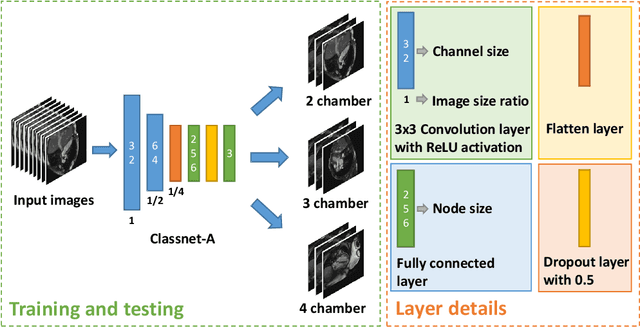
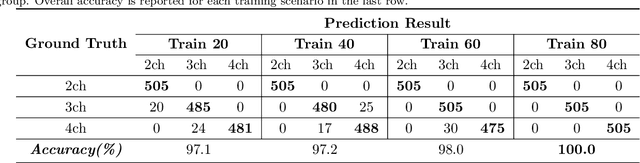
Abstract:This study proposes a fully automated approach for the left atrial segmentation from routine cine long-axis cardiac magnetic resonance image sequences using deep convolutional neural networks and Bayesian filtering. The proposed approach consists of a classification network that automatically detects the type of long-axis sequence and three different convolutional neural network models followed by unscented Kalman filtering (UKF) that delineates the left atrium. Instead of training and predicting all long-axis sequence types together, the proposed approach first identifies the image sequence type as to 2, 3 and 4 chamber views, and then performs prediction based on neural nets trained for that particular sequence type. The datasets were acquired retrospectively and ground truth manual segmentation was provided by an expert radiologist. In addition to neural net based classification and segmentation, another neural net is trained and utilized to select image sequences for further processing using UKF to impose temporal consistency over cardiac cycle. A cyclic dynamic model with time-varying angular frequency is introduced in UKF to characterize the variations in cardiac motion during image scanning. The proposed approach was trained and evaluated separately with varying amount of training data with images acquired from 20, 40, 60 and 80 patients. Evaluations over 1515 images with equal number of images from each chamber group acquired from an additional 20 patients demonstrated that the proposed model outperformed state-of-the-art and yielded a mean Dice coefficient value of 94.1%, 93.7% and 90.1% for 2, 3 and 4-chamber sequences, respectively, when trained with datasets from 80 patients.
Fully automated deep learning based segmentation of normal, infarcted and edema regions from multiple cardiac MRI sequences
Aug 18, 2020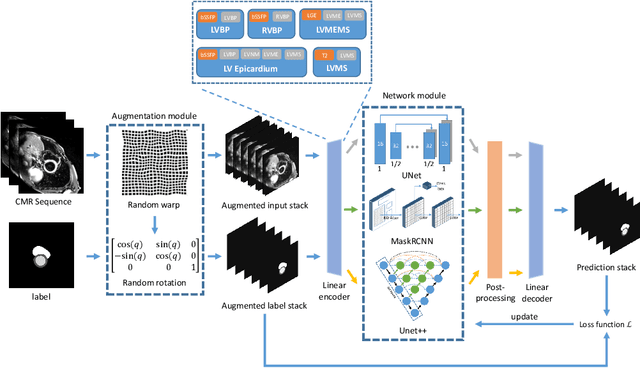
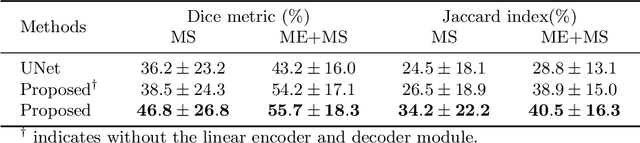

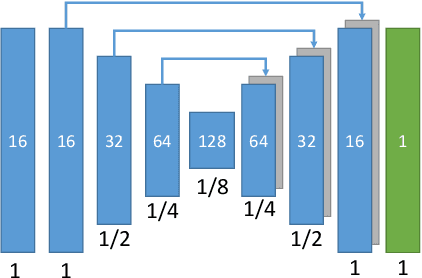
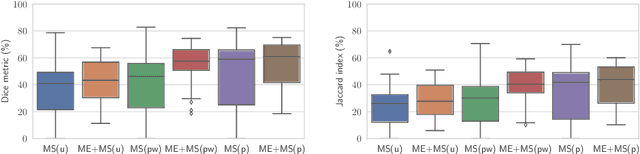
Abstract:Myocardial characterization is essential for patients with myocardial infarction and other myocardial diseases, and the assessment is often performed using cardiac magnetic resonance (CMR) sequences. In this study, we propose a fully automated approach using deep convolutional neural networks (CNN) for cardiac pathology segmentation, including left ventricular (LV) blood pool, right ventricular blood pool, LV normal myocardium, LV myocardial edema (ME) and LV myocardial scars (MS). The input to the network consists of three CMR sequences, namely, late gadolinium enhancement (LGE), T2 and balanced steady state free precession (bSSFP). The proposed approach utilized the data provided by the MyoPS challenge hosted by MICCAI 2020 in conjunction with STACOM. The training set for the CNN model consists of images acquired from 25 cases, and the gold standard labels are provided by trained raters and validated by radiologists. The proposed approach introduces a data augmentation module, linear encoder and decoder module and a network module to increase the number of training samples and improve the prediction accuracy for LV ME and MS. The proposed approach is evaluated by the challenge organizers with a test set including 20 cases and achieves a mean dice score of $46.8\%$ for LV MS and $55.7\%$ for LV ME+MS
A systematic review on the role of artificial intelligence in sonographic diagnosis of thyroid cancer: Past, present and future
Jun 10, 2020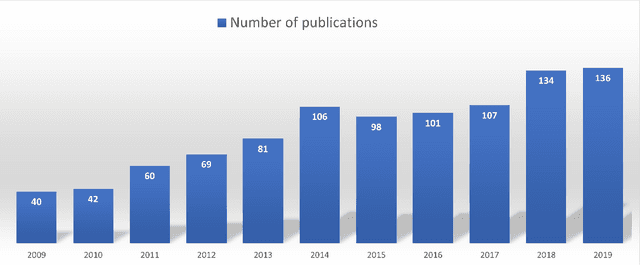
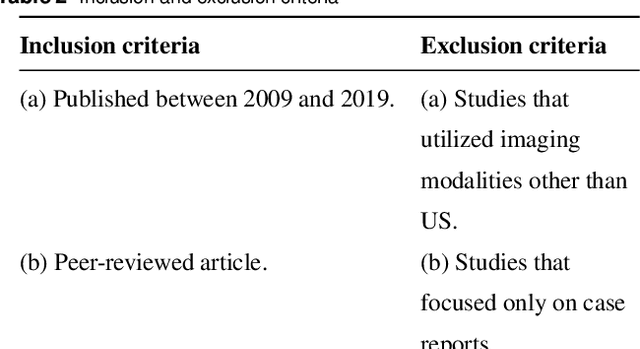


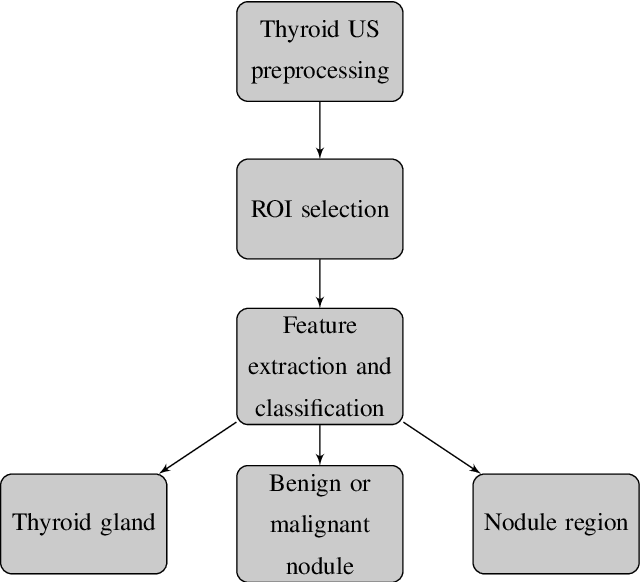
Abstract:Thyroid cancer is common worldwide, with a rapid increase in prevalence across North America in recent years. While most patients present with palpable nodules through physical examination, a large number of small and medium-sized nodules are detected by ultrasound examination. Suspicious nodules are then sent for biopsy through fine needle aspiration. Since biopsies are invasive and sometimes inconclusive, various research groups have tried to develop computer-aided diagnosis systems. Earlier approaches along these lines relied on clinically relevant features that were manually identified by radiologists. With the recent success of artificial intelligence (AI), various new methods are being developed to identify these features in thyroid ultrasound automatically. In this paper, we present a systematic review of state-of-the-art on AI application in sonographic diagnosis of thyroid cancer. This review follows a methodology-based classification of the different techniques available for thyroid cancer diagnosis. With more than 50 papers included in this review, we reflect on the trends and challenges of the field of sonographic diagnosis of thyroid malignancies and potential of computer-aided diagnosis to increase the impact of ultrasound applications on the future of thyroid cancer diagnosis. Machine learning will continue to play a fundamental role in the development of future thyroid cancer diagnosis frameworks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge