Luca Giancardo
Benchmarking the CoW with the TopCoW Challenge: Topology-Aware Anatomical Segmentation of the Circle of Willis for CTA and MRA
Dec 29, 2023



Abstract:The Circle of Willis (CoW) is an important network of arteries connecting major circulations of the brain. Its vascular architecture is believed to affect the risk, severity, and clinical outcome of serious neuro-vascular diseases. However, characterizing the highly variable CoW anatomy is still a manual and time-consuming expert task. The CoW is usually imaged by two angiographic imaging modalities, magnetic resonance angiography (MRA) and computed tomography angiography (CTA), but there exist limited public datasets with annotations on CoW anatomy, especially for CTA. Therefore we organized the TopCoW Challenge in 2023 with the release of an annotated CoW dataset and invited submissions worldwide for the CoW segmentation task, which attracted over 140 registered participants from four continents. TopCoW dataset was the first public dataset with voxel-level annotations for CoW's 13 vessel components, made possible by virtual-reality (VR) technology. It was also the first dataset with paired MRA and CTA from the same patients. TopCoW challenge aimed to tackle the CoW characterization problem as a multiclass anatomical segmentation task with an emphasis on topological metrics. The top performing teams managed to segment many CoW components to Dice scores around 90%, but with lower scores for communicating arteries and rare variants. There were also topological mistakes for predictions with high Dice scores. Additional topological analysis revealed further areas for improvement in detecting certain CoW components and matching CoW variant's topology accurately. TopCoW represented a first attempt at benchmarking the CoW anatomical segmentation task for MRA and CTA, both morphologically and topologically.
Dental CLAIRES: Contrastive LAnguage Image REtrieval Search for Dental Research
Jun 27, 2023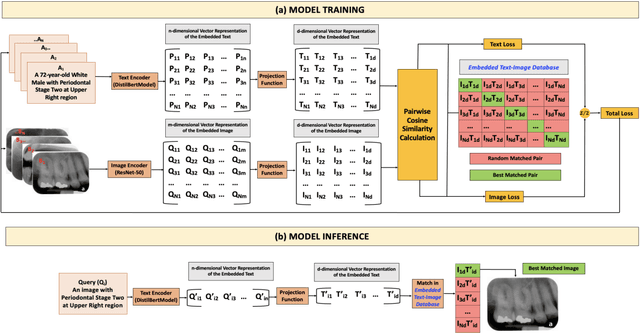
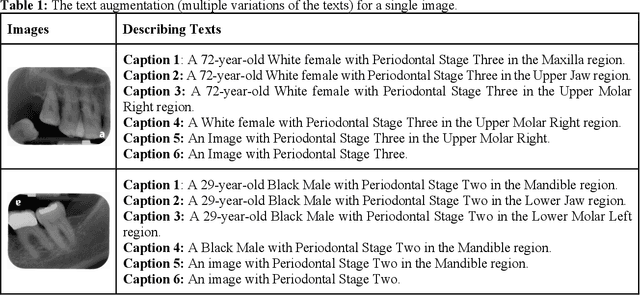

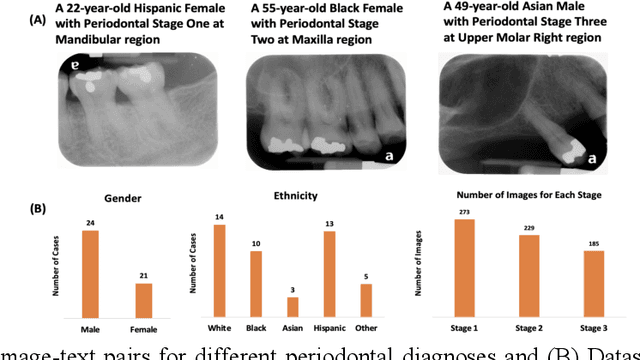
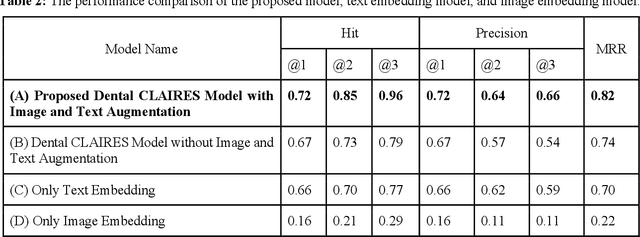
Abstract:Learning about diagnostic features and related clinical information from dental radiographs is important for dental research. However, the lack of expert-annotated data and convenient search tools poses challenges. Our primary objective is to design a search tool that uses a user's query for oral-related research. The proposed framework, Contrastive LAnguage Image REtrieval Search for dental research, Dental CLAIRES, utilizes periapical radiographs and associated clinical details such as periodontal diagnosis, demographic information to retrieve the best-matched images based on the text query. We applied a contrastive representation learning method to find images described by the user's text by maximizing the similarity score of positive pairs (true pairs) and minimizing the score of negative pairs (random pairs). Our model achieved a hit@3 ratio of 96% and a Mean Reciprocal Rank (MRR) of 0.82. We also designed a graphical user interface that allows researchers to verify the model's performance with interactions.
* 10 pages, 7 figures, 4 tables
Eye-SpatialNet: Spatial Information Extraction from Ophthalmology Notes
May 19, 2023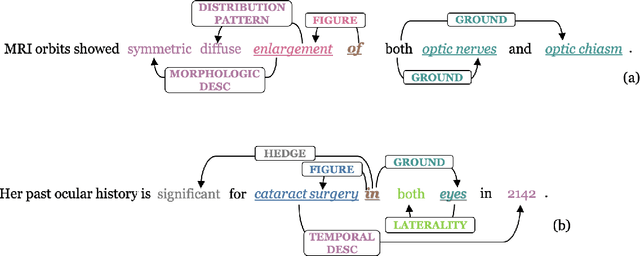



Abstract:We introduce an annotated corpus of 600 ophthalmology notes labeled with detailed spatial and contextual information of ophthalmic entities. We extend our previously proposed frame semantics-based spatial representation schema, Rad-SpatialNet, to represent spatial language in ophthalmology text, resulting in the Eye-SpatialNet schema. The spatially-grounded entities are findings, procedures, and drugs. To accurately capture all spatial details, we add some domain-specific elements in Eye-SpatialNet. The annotated corpus contains 1715 spatial triggers, 7308 findings, 2424 anatomies, and 9914 descriptors. To automatically extract the spatial information, we employ a two-turn question answering approach based on the transformer language model BERT. The results are promising, with F1 scores of 89.31, 74.86, and 88.47 for spatial triggers, Figure, and Ground frame elements, respectively. This is the first work to represent and extract a wide variety of clinical information in ophthalmology. Extracting detailed information can benefit ophthalmology applications and research targeted toward disease progression and screening.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge