Lena Simine
Diversifying Design of Nucleic Acid Aptamers Using Unsupervised Machine Learning
Aug 10, 2022
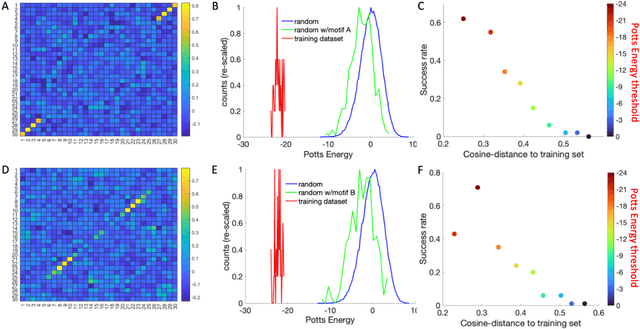
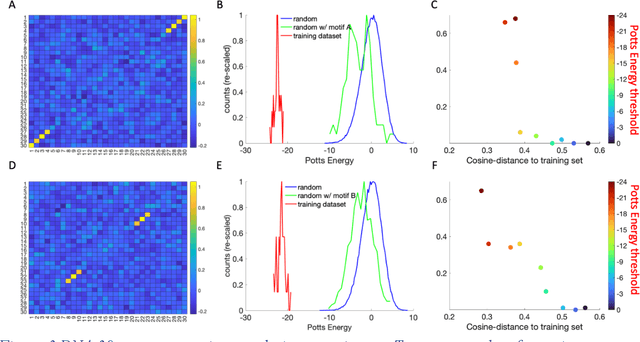
Abstract:Inverse design of short single-stranded RNA and DNA sequences (aptamers) is the task of finding sequences that satisfy a set of desired criteria. Relevant criteria may be, for example, the presence of specific folding motifs, binding to molecular ligands, sensing properties, etc. Most practical approaches to aptamer design identify a small set of promising candidate sequences using high-throughput experiments (e.g. SELEX), and then optimize performance by introducing only minor modifications to the empirically found candidates. Sequences that possess the desired properties but differ drastically in chemical composition will add diversity to the search space and facilitate the discovery of useful nucleic acid aptamers. Systematic diversification protocols are needed. Here we propose to use an unsupervised machine learning model known as the Potts model to discover new, useful sequences with controllable sequence diversity. We start by training a Potts model using the maximum entropy principle on a small set of empirically identified sequences unified by a common feature. To generate new candidate sequences with a controllable degree of diversity, we take advantage of the model's spectral feature: an energy bandgap separating sequences that are similar to the training set from those that are distinct. By controlling the Potts energy range that is sampled, we generate sequences that are distinct from the training set yet still likely to have the encoded features. To demonstrate performance, we apply our approach to design diverse pools of sequences with specified secondary structure motifs in 30-mer RNA and DNA aptamers.
Biological Sequence Design with GFlowNets
Mar 02, 2022


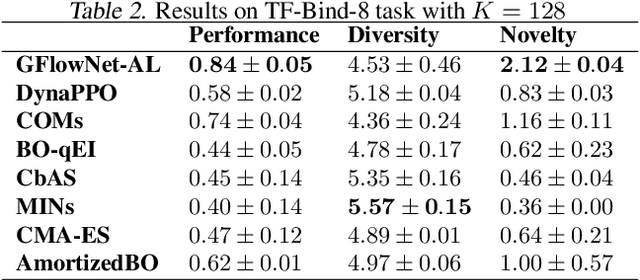
Abstract:Design of de novo biological sequences with desired properties, like protein and DNA sequences, often involves an active loop with several rounds of molecule ideation and expensive wet-lab evaluations. These experiments can consist of multiple stages, with increasing levels of precision and cost of evaluation, where candidates are filtered. This makes the diversity of proposed candidates a key consideration in the ideation phase. In this work, we propose an active learning algorithm leveraging epistemic uncertainty estimation and the recently proposed GFlowNets as a generator of diverse candidate solutions, with the objective to obtain a diverse batch of useful (as defined by some utility function, for example, the predicted anti-microbial activity of a peptide) and informative candidates after each round. We also propose a scheme to incorporate existing labeled datasets of candidates, in addition to a reward function, to speed up learning in GFlowNets. We present empirical results on several biological sequence design tasks, and we find that our method generates more diverse and novel batches with high scoring candidates compared to existing approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge