Jiahua Shi
FedDifRC: Unlocking the Potential of Text-to-Image Diffusion Models in Heterogeneous Federated Learning
Jul 09, 2025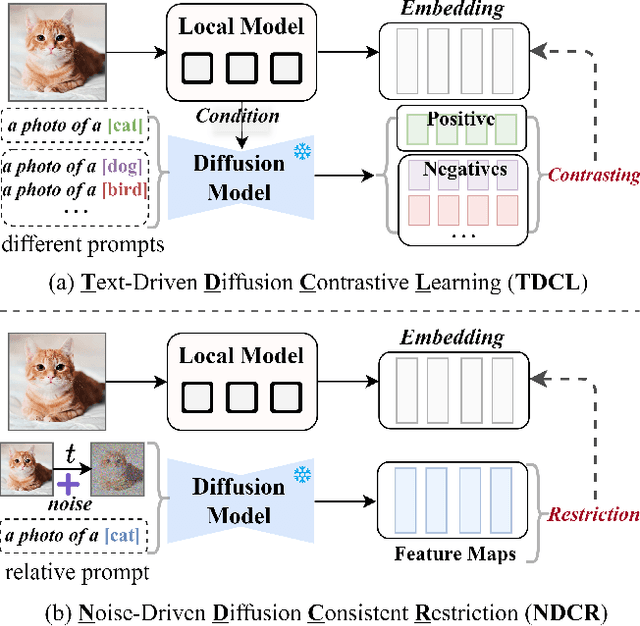



Abstract:Federated learning aims at training models collaboratively across participants while protecting privacy. However, one major challenge for this paradigm is the data heterogeneity issue, where biased data preferences across multiple clients, harming the model's convergence and performance. In this paper, we first introduce powerful diffusion models into the federated learning paradigm and show that diffusion representations are effective steers during federated training. To explore the possibility of using diffusion representations in handling data heterogeneity, we propose a novel diffusion-inspired Federated paradigm with Diffusion Representation Collaboration, termed FedDifRC, leveraging meaningful guidance of diffusion models to mitigate data heterogeneity. The key idea is to construct text-driven diffusion contrasting and noise-driven diffusion regularization, aiming to provide abundant class-related semantic information and consistent convergence signals. On the one hand, we exploit the conditional feedback from the diffusion model for different text prompts to build a text-driven contrastive learning strategy. On the other hand, we introduce a noise-driven consistency regularization to align local instances with diffusion denoising representations, constraining the optimization region in the feature space. In addition, FedDifRC can be extended to a self-supervised scheme without relying on any labeled data. We also provide a theoretical analysis for FedDifRC to ensure convergence under non-convex objectives. The experiments on different scenarios validate the effectiveness of FedDifRC and the efficiency of crucial components.
FedSC: Federated Learning with Semantic-Aware Collaboration
Jun 26, 2025



Abstract:Federated learning (FL) aims to train models collaboratively across clients without sharing data for privacy-preserving. However, one major challenge is the data heterogeneity issue, which refers to the biased labeling preferences at multiple clients. A number of existing FL methods attempt to tackle data heterogeneity locally (e.g., regularizing local models) or globally (e.g., fine-tuning global model), often neglecting inherent semantic information contained in each client. To explore the possibility of using intra-client semantically meaningful knowledge in handling data heterogeneity, in this paper, we propose Federated Learning with Semantic-Aware Collaboration (FedSC) to capture client-specific and class-relevant knowledge across heterogeneous clients. The core idea of FedSC is to construct relational prototypes and consistent prototypes at semantic-level, aiming to provide fruitful class underlying knowledge and stable convergence signals in a prototype-wise collaborative way. On the one hand, FedSC introduces an inter-contrastive learning strategy to bring instance-level embeddings closer to relational prototypes with the same semantics and away from distinct classes. On the other hand, FedSC devises consistent prototypes via a discrepancy aggregation manner, as a regularization penalty to constrain the optimization region of the local model. Moreover, a theoretical analysis for FedSC is provided to ensure a convergence guarantee. Experimental results on various challenging scenarios demonstrate the effectiveness of FedSC and the efficiency of crucial components.
FedSKC: Federated Learning with Non-IID Data via Structural Knowledge Collaboration
May 25, 2025Abstract:With the advancement of edge computing, federated learning (FL) displays a bright promise as a privacy-preserving collaborative learning paradigm. However, one major challenge for FL is the data heterogeneity issue, which refers to the biased labeling preferences among multiple clients, negatively impacting convergence and model performance. Most previous FL methods attempt to tackle the data heterogeneity issue locally or globally, neglecting underlying class-wise structure information contained in each client. In this paper, we first study how data heterogeneity affects the divergence of the model and decompose it into local, global, and sampling drift sub-problems. To explore the potential of using intra-client class-wise structural knowledge in handling these drifts, we thus propose Federated Learning with Structural Knowledge Collaboration (FedSKC). The key idea of FedSKC is to extract and transfer domain preferences from inter-client data distributions, offering diverse class-relevant knowledge and a fair convergent signal. FedSKC comprises three components: i) local contrastive learning, to prevent weight divergence resulting from local training; ii) global discrepancy aggregation, which addresses the parameter deviation between the server and clients; iii) global period review, correcting for the sampling drift introduced by the server randomly selecting devices. We have theoretically analyzed FedSKC under non-convex objectives and empirically validated its superiority through extensive experimental results.
Vox-UDA: Voxel-wise Unsupervised Domain Adaptation for Cryo-Electron Subtomogram Segmentation with Denoised Pseudo Labeling
Jun 25, 2024Abstract:Cryo-Electron Tomography (cryo-ET) is a 3D imaging technology facilitating the study of macromolecular structures at near-atomic resolution. Recent volumetric segmentation approaches on cryo-ET images have drawn widespread interest in biological sector. However, existing methods heavily rely on manually labeled data, which requires highly professional skills, thereby hindering the adoption of fully-supervised approaches for cryo-ET images. Some unsupervised domain adaptation (UDA) approaches have been designed to enhance the segmentation network performance using unlabeled data. However, applying these methods directly to cryo-ET images segmentation tasks remains challenging due to two main issues: 1) the source data, usually obtained through simulation, contain a certain level of noise, while the target data, directly collected from raw-data from real-world scenario, have unpredictable noise levels. 2) the source data used for training typically consists of known macromoleculars, while the target domain data are often unknown, causing the model's segmenter to be biased towards these known macromolecules, leading to a domain shift problem. To address these challenges, in this work, we introduce the first voxel-wise unsupervised domain adaptation approach, termed Vox-UDA, specifically for cryo-ET subtomogram segmentation. Vox-UDA incorporates a noise generation module to simulate target-like noises in the source dataset for cross-noise level adaptation. Additionally, we propose a denoised pseudo-labeling strategy based on improved Bilateral Filter to alleviate the domain shift problem. Experimental results on both simulated and real cryo-ET subtomogram datasets demonstrate the superiority of our proposed approach compared to state-of-the-art UDA methods.
FDNet: Frequency Domain Denoising Network For Cell Segmentation in Astrocytes Derived From Induced Pluripotent Stem Cells
Feb 05, 2024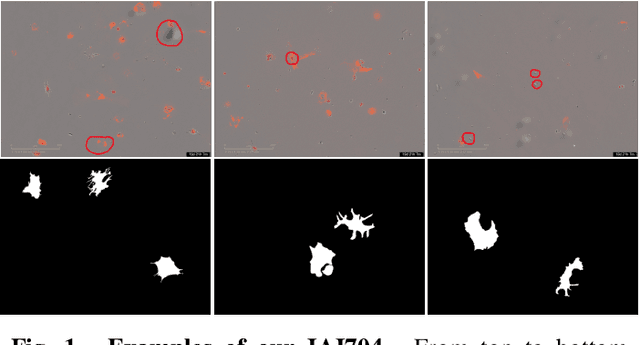
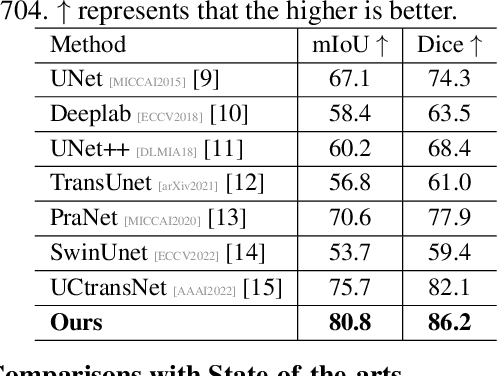
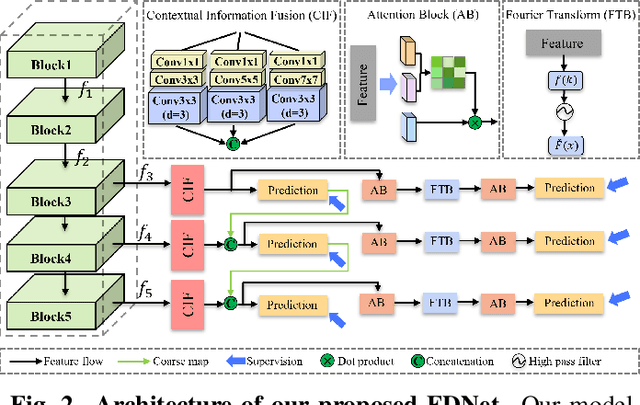
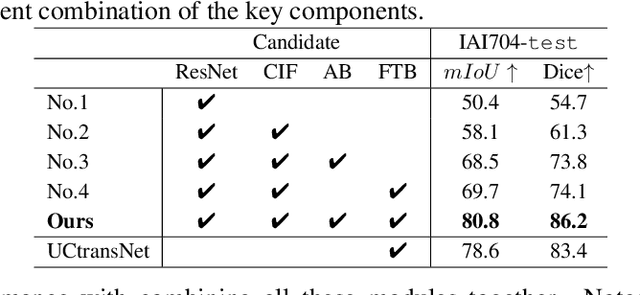
Abstract:Artificially generated induced pluripotent stem cells (iPSCs) from somatic cells play an important role for disease modeling and drug screening of neurodegenerative diseases. Astrocytes differentiated from iPSCs are important targets to investigate neuronal metabolism. The astrocyte differentiation progress can be monitored through the variations of morphology observed from microscopy images at different differentiation stages, then determined by molecular biology techniques upon maturation. However, the astrocytes usually ``perfectly'' blend into the background and some of them are covered by interference information (i.e., dead cells, media sediments, and cell debris), which makes astrocytes difficult to observe. Due to the lack of annotated datasets, the existing state-of-the-art deep learning approaches cannot be used to address this issue. In this paper, we introduce a new task named astrocyte segmentation with a novel dataset, called IAI704, which contains 704 images and their corresponding pixel-level annotation masks. Moreover, a novel frequency domain denoising network, named FDNet, is proposed for astrocyte segmentation. In detail, our FDNet consists of a contextual information fusion module (CIF), an attention block (AB), and a Fourier transform block (FTB). CIF and AB fuse multi-scale feature embeddings to localize the astrocytes. FTB transforms feature embeddings into the frequency domain and conducts a high-pass filter to eliminate interference information. Experimental results demonstrate the superiority of our proposed FDNet over the state-of-the-art substitutes in astrocyte segmentation, shedding insights for iPSC differentiation progress prediction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge