Jacob M. Luber
Adversarial Hospital-Invariant Feature Learning for WSI Patch Classification
Aug 20, 2025

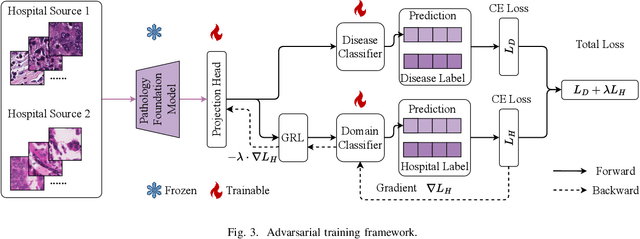

Abstract:Pathology foundation models (PFMs) have demonstrated remarkable potential in whole-slide image (WSI) diagnosis. However, pathology images from different hospitals often vary due to differences in scanning hardware and preprocessing styles, which may lead PFMs to inadvertently learn hospital-specific features, posing risks for clinical deployment. In this work, we present the first systematic study of domain bias in PFMs arising from hospital source characteristics. Specifically, we (1) construct a pipeline for quantifying domain bias in PFMs, (2) evaluate and compare the performance of multiple models, and (3) propose a lightweight adversarial framework that removes latent hospital-specific features from frozen representations without modifying the encoder itself. By introducing a trainable adapter and a domain classifier connected through a gradient reversal layer (GRL), our method learns task-discriminative yet domain-invariant representations. Experiments on multi-center histopathology datasets demonstrate that our approach substantially reduces domain predictability while maintaining or even improving disease classification performance, particularly in out-of-domain (unseen hospital) scenarios. Further analyses, including hospital detection and feature space visualization, confirm the effectiveness of our method in mitigating hospital bias. We will provide our code based on acceptance.
Beyond the Monitor: Mixed Reality Visualization and AI for Enhanced Digital Pathology Workflow
May 05, 2025Abstract:Pathologists rely on gigapixel whole-slide images (WSIs) to diagnose diseases like cancer, yet current digital pathology tools hinder diagnosis. The immense scale of WSIs, often exceeding 100,000 X 100,000 pixels, clashes with the limited views traditional monitors offer. This mismatch forces constant panning and zooming, increasing pathologist cognitive load, causing diagnostic fatigue, and slowing pathologists' adoption of digital methods. PathVis, our mixed-reality visualization platform for Apple Vision Pro, addresses these challenges. It transforms the pathologist's interaction with data, replacing cumbersome mouse-and-monitor navigation with intuitive exploration using natural hand gestures, eye gaze, and voice commands in an immersive workspace. PathVis integrates AI to enhance diagnosis. An AI-driven search function instantly retrieves and displays the top five similar patient cases side-by-side, improving diagnostic precision and efficiency through rapid comparison. Additionally, a multimodal conversational AI assistant offers real-time image interpretation support and aids collaboration among pathologists across multiple Apple devices. By merging the directness of traditional pathology with advanced mixed-reality visualization and AI, PathVis improves diagnostic workflows, reduces cognitive strain, and makes pathology practice more effective and engaging. The PathVis source code and a demo video are publicly available at: https://github.com/jaiprakash1824/Path_Vis
Dual Codebook VQ: Enhanced Image Reconstruction with Reduced Codebook Size
Mar 13, 2025Abstract:Vector Quantization (VQ) techniques face significant challenges in codebook utilization, limiting reconstruction fidelity in image modeling. We introduce a Dual Codebook mechanism that effectively addresses this limitation by partitioning the representation into complementary global and local components. The global codebook employs a lightweight transformer for concurrent updates of all code vectors, while the local codebook maintains precise feature representation through deterministic selection. This complementary approach is trained from scratch without requiring pre-trained knowledge. Experimental evaluation across multiple standard benchmark datasets demonstrates state-of-the-art reconstruction quality while using a compact codebook of size 512 - half the size of previous methods that require pre-training. Our approach achieves significant FID improvements across diverse image domains, particularly excelling in scene and face reconstruction tasks. These results establish Dual Codebook VQ as an efficient paradigm for high-fidelity image reconstruction with significantly reduced computational requirements.
Peptide Sequencing Via Protein Language Models
Aug 01, 2024



Abstract:We introduce a protein language model for determining the complete sequence of a peptide based on measurement of a limited set of amino acids. To date, protein sequencing relies on mass spectrometry, with some novel edman degregation based platforms able to sequence non-native peptides. Current protein sequencing techniques face limitations in accurately identifying all amino acids, hindering comprehensive proteome analysis. Our method simulates partial sequencing data by selectively masking amino acids that are experimentally difficult to identify in protein sequences from the UniRef database. This targeted masking mimics real-world sequencing limitations. We then modify and finetune a ProtBert derived transformer-based model, for a new downstream task predicting these masked residues, providing an approximation of the complete sequence. Evaluating on three bacterial Escherichia species, we achieve per-amino-acid accuracy up to 90.5% when only four amino acids ([KCYM]) are known. Structural assessment using AlphaFold and TM-score validates the biological relevance of our predictions. The model also demonstrates potential for evolutionary analysis through cross-species performance. This integration of simulated experimental constraints with computational predictions offers a promising avenue for enhancing protein sequence analysis, potentially accelerating advancements in proteomics and structural biology by providing a probabilistic reconstruction of the complete protein sequence from limited experimental data.
Vulnerabilities Unveiled: Adversarially Attacking a Multimodal Vision Language Model for Pathology Imaging
Jan 08, 2024



Abstract:In the dynamic landscape of medical artificial intelligence, this study explores the vulnerabilities of the Pathology Language-Image Pretraining (PLIP) model, a Vision Language Foundation model, under targeted adversarial conditions. Leveraging the Kather Colon dataset with 7,180 H&E images across nine tissue types, our investigation employs Projected Gradient Descent (PGD) adversarial attacks to intentionally induce misclassifications. The outcomes reveal a 100% success rate in manipulating PLIP's predictions, underscoring its susceptibility to adversarial perturbations. The qualitative analysis of adversarial examples delves into the interpretability challenges, shedding light on nuanced changes in predictions induced by adversarial manipulations. These findings contribute crucial insights into the interpretability, domain adaptation, and trustworthiness of Vision Language Models in medical imaging. The study emphasizes the pressing need for robust defenses to ensure the reliability of AI models.
Predicting Future States with Spatial Point Processes in Single Molecule Resolution Spatial Transcriptomics
Jan 04, 2024



Abstract:In this paper, we introduce a pipeline based on Random Forest Regression to predict the future distribution of cells that are expressed by the Sog-D gene (active cells) in both the Anterior to posterior (AP) and the Dorsal to Ventral (DV) axis of the Drosophila in embryogenesis process. This method provides insights about how cells and living organisms control gene expression in super resolution whole embryo spatial transcriptomics imaging at sub cellular, single molecule resolution. A Random Forest Regression model was used to predict the next stage active distribution based on the previous one. To achieve this goal, we leveraged temporally resolved, spatial point processes by including Ripley's K-function in conjunction with the cell's state in each stage of embryogenesis, and found average predictive accuracy of active cell distribution. This tool is analogous to RNA Velocity for spatially resolved developmental biology, from one data point we can predict future spatially resolved gene expression using features from the spatial point processes.
The State of Applying Artificial Intelligence to Tissue Imaging for Cancer Research and Early Detection
Jun 29, 2023



Abstract:Artificial intelligence represents a new frontier in human medicine that could save more lives and reduce the costs, thereby increasing accessibility. As a consequence, the rate of advancement of AI in cancer medical imaging and more particularly tissue pathology has exploded, opening it to ethical and technical questions that could impede its adoption into existing systems. In order to chart the path of AI in its application to cancer tissue imaging, we review current work and identify how it can improve cancer pathology diagnostics and research. In this review, we identify 5 core tasks that models are developed for, including regression, classification, segmentation, generation, and compression tasks. We address the benefits and challenges that such methods face, and how they can be adapted for use in cancer prevention and treatment. The studies looked at in this paper represent the beginning of this field and future experiments will build on the foundations that we highlight.
Histopathology Slide Indexing and Search: Are We There Yet?
Jun 29, 2023


Abstract:The search and retrieval of digital histopathology slides is an important task that has yet to be solved. In this case study, we investigate the clinical readiness of three state-of-the-art histopathology slide search engines, Yottixel, SISH, and RetCCL, on three patients with solid tumors. We provide a qualitative assessment of each model's performance in providing retrieval results that are reliable and useful to pathologists. We found that all three image search engines fail to produce consistently reliable results and have difficulties in capturing granular and subtle features of malignancy, limiting their diagnostic accuracy. Based on our findings, we also propose a minimal set of requirements to further advance the development of accurate and reliable histopathology image search engines for successful clinical adoption.
Clinically Relevant Latent Space Embedding of Cancer Histopathology Slides through Variational Autoencoder Based Image Compression
Mar 23, 2023Abstract:In this paper, we introduce a Variational Autoencoder (VAE) based training approach that can compress and decompress cancer pathology slides at a compression ratio of 1:512, which is better than the previously reported state of the art (SOTA) in the literature, while still maintaining accuracy in clinical validation tasks. The compression approach was tested on more common computer vision datasets such as CIFAR10, and we explore which image characteristics enable this compression ratio on cancer imaging data but not generic images. We generate and visualize embeddings from the compressed latent space and demonstrate how they are useful for clinical interpretation of data, and how in the future such latent embeddings can be used to accelerate search of clinical imaging data.
A SSIM Guided cGAN Architecture For Clinically Driven Generative Image Synthesis of Multiplexed Spatial Proteomics Channels
May 20, 2022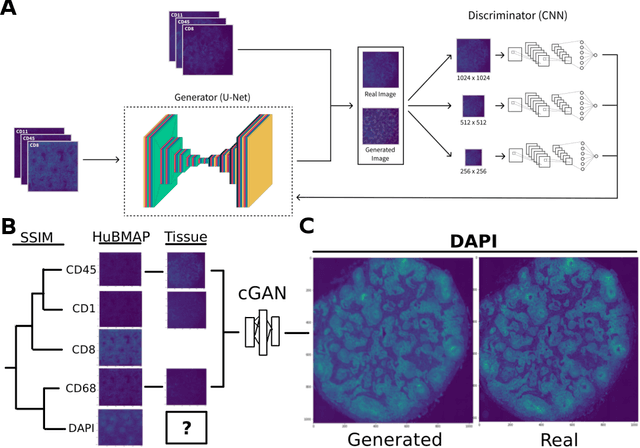
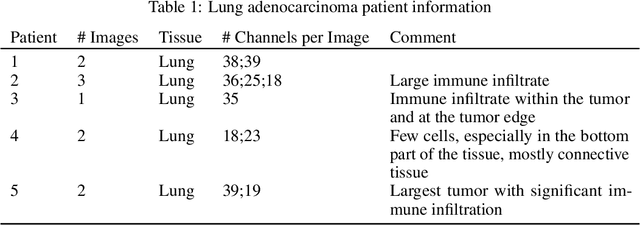


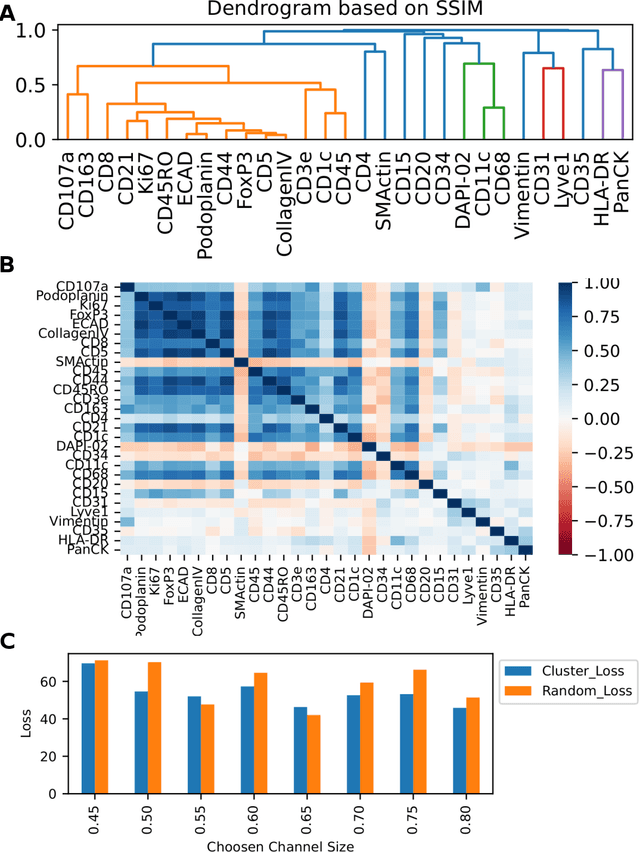
Abstract:Here we present a structural similarity index measure (SSIM) guided conditional Generative Adversarial Network (cGAN) that generatively performs image-to-image (i2i) synthesis to generate photo-accurate protein channels in multiplexed spatial proteomics images. This approach can be utilized to accurately generate missing spatial proteomics channels that were not included during experimental data collection either at the bench or the clinic. Experimental spatial proteomic data from the Human BioMolecular Atlas Program (HuBMAP) was used to generate spatial representations of missing proteins through a U-Net based image synthesis pipeline. HuBMAP channels were hierarchically clustered by the (SSIM) as a heuristic to obtain the minimal set needed to recapitulate the underlying biology represented by the spatial landscape of proteins. We subsequently prove that our SSIM based architecture allows for scaling of generative image synthesis to slides with up to 100 channels, which is better than current state of the art algorithms which are limited to data with 11 channels. We validate these claims by generating a new experimental spatial proteomics data set from human lung adenocarcinoma tissue sections and show that a model trained on HuBMAP can accurately synthesize channels from our new data set. The ability to recapitulate experimental data from sparsely stained multiplexed histological slides containing spatial proteomic will have tremendous impact on medical diagnostics and drug development, and also raises important questions on the medical ethics of utilizing data produced by generative image synthesis in the clinical setting. The algorithm that we present in this paper will allow researchers and clinicians to save time and costs in proteomics based histological staining while also increasing the amount of data that they can generate through their experiments.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge