Jai Prakash Veerla
Beyond the Monitor: Mixed Reality Visualization and AI for Enhanced Digital Pathology Workflow
May 05, 2025Abstract:Pathologists rely on gigapixel whole-slide images (WSIs) to diagnose diseases like cancer, yet current digital pathology tools hinder diagnosis. The immense scale of WSIs, often exceeding 100,000 X 100,000 pixels, clashes with the limited views traditional monitors offer. This mismatch forces constant panning and zooming, increasing pathologist cognitive load, causing diagnostic fatigue, and slowing pathologists' adoption of digital methods. PathVis, our mixed-reality visualization platform for Apple Vision Pro, addresses these challenges. It transforms the pathologist's interaction with data, replacing cumbersome mouse-and-monitor navigation with intuitive exploration using natural hand gestures, eye gaze, and voice commands in an immersive workspace. PathVis integrates AI to enhance diagnosis. An AI-driven search function instantly retrieves and displays the top five similar patient cases side-by-side, improving diagnostic precision and efficiency through rapid comparison. Additionally, a multimodal conversational AI assistant offers real-time image interpretation support and aids collaboration among pathologists across multiple Apple devices. By merging the directness of traditional pathology with advanced mixed-reality visualization and AI, PathVis improves diagnostic workflows, reduces cognitive strain, and makes pathology practice more effective and engaging. The PathVis source code and a demo video are publicly available at: https://github.com/jaiprakash1824/Path_Vis
Vulnerabilities Unveiled: Adversarially Attacking a Multimodal Vision Language Model for Pathology Imaging
Jan 08, 2024



Abstract:In the dynamic landscape of medical artificial intelligence, this study explores the vulnerabilities of the Pathology Language-Image Pretraining (PLIP) model, a Vision Language Foundation model, under targeted adversarial conditions. Leveraging the Kather Colon dataset with 7,180 H&E images across nine tissue types, our investigation employs Projected Gradient Descent (PGD) adversarial attacks to intentionally induce misclassifications. The outcomes reveal a 100% success rate in manipulating PLIP's predictions, underscoring its susceptibility to adversarial perturbations. The qualitative analysis of adversarial examples delves into the interpretability challenges, shedding light on nuanced changes in predictions induced by adversarial manipulations. These findings contribute crucial insights into the interpretability, domain adaptation, and trustworthiness of Vision Language Models in medical imaging. The study emphasizes the pressing need for robust defenses to ensure the reliability of AI models.
Real-Time Diagnostic Integrity Meets Efficiency: A Novel Platform-Agnostic Architecture for Physiological Signal Compression
Jan 04, 2024Abstract:Head-based signals such as EEG, EMG, EOG, and ECG collected by wearable systems will play a pivotal role in clinical diagnosis, monitoring, and treatment of important brain disorder diseases. However, the real-time transmission of the significant corpus physiological signals over extended periods consumes substantial power and time, limiting the viability of battery-dependent physiological monitoring wearables. This paper presents a novel deep-learning framework employing a variational autoencoder (VAE) for physiological signal compression to reduce wearables' computational complexity and energy consumption. Our approach achieves an impressive compression ratio of 1:293 specifically for spectrogram data, surpassing state-of-the-art compression techniques such as JPEG2000, H.264, Direct Cosine Transform (DCT), and Huffman Encoding, which do not excel in handling physiological signals. We validate the efficacy of the compressed algorithms using collected physiological signals from real patients in the Hospital and deploy the solution on commonly used embedded AI chips (i.e., ARM Cortex V8 and Jetson Nano). The proposed framework achieves a 91% seizure detection accuracy using XGBoost, confirming the approach's reliability, practicality, and scalability.
Predicting Future States with Spatial Point Processes in Single Molecule Resolution Spatial Transcriptomics
Jan 04, 2024



Abstract:In this paper, we introduce a pipeline based on Random Forest Regression to predict the future distribution of cells that are expressed by the Sog-D gene (active cells) in both the Anterior to posterior (AP) and the Dorsal to Ventral (DV) axis of the Drosophila in embryogenesis process. This method provides insights about how cells and living organisms control gene expression in super resolution whole embryo spatial transcriptomics imaging at sub cellular, single molecule resolution. A Random Forest Regression model was used to predict the next stage active distribution based on the previous one. To achieve this goal, we leveraged temporally resolved, spatial point processes by including Ripley's K-function in conjunction with the cell's state in each stage of embryogenesis, and found average predictive accuracy of active cell distribution. This tool is analogous to RNA Velocity for spatially resolved developmental biology, from one data point we can predict future spatially resolved gene expression using features from the spatial point processes.
Histopathology Slide Indexing and Search: Are We There Yet?
Jun 29, 2023
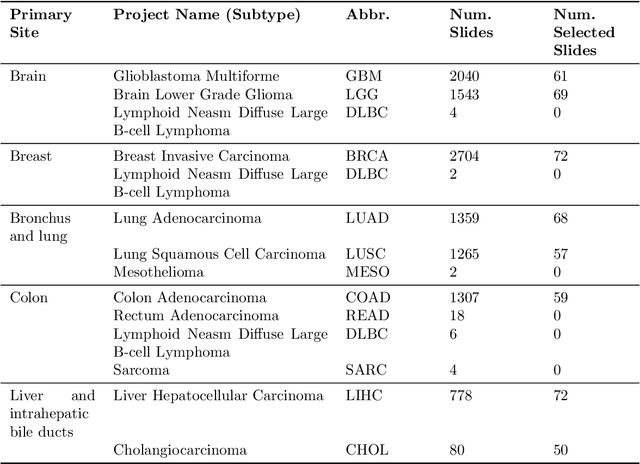


Abstract:The search and retrieval of digital histopathology slides is an important task that has yet to be solved. In this case study, we investigate the clinical readiness of three state-of-the-art histopathology slide search engines, Yottixel, SISH, and RetCCL, on three patients with solid tumors. We provide a qualitative assessment of each model's performance in providing retrieval results that are reliable and useful to pathologists. We found that all three image search engines fail to produce consistently reliable results and have difficulties in capturing granular and subtle features of malignancy, limiting their diagnostic accuracy. Based on our findings, we also propose a minimal set of requirements to further advance the development of accurate and reliable histopathology image search engines for successful clinical adoption.
Multimodal Pathology Image Search Between H&E Slides and Multiplexed Immunofluorescent Images
Jun 11, 2023



Abstract:We present an approach for multimodal pathology image search, using dynamic time warping (DTW) on Variational Autoencoder (VAE) latent space that is fed into a ranked choice voting scheme to retrieve multiplexed immunofluorescent imaging (mIF) that is most similar to a query H&E slide. Through training the VAE and applying DTW, we align and compare mIF and H&E slides. Our method improves differential diagnosis and therapeutic decisions by integrating morphological H&E data with immunophenotyping from mIF, providing clinicians a rich perspective of disease states. This facilitates an understanding of the spatial relationships in tissue samples and could revolutionize the diagnostic process, enhancing precision and enabling personalized therapy selection. Our technique demonstrates feasibility using colorectal cancer and healthy tonsil samples. An exhaustive ablation study was conducted on a search engine designed to explore the correlation between multiplexed Immunofluorescence (mIF) and Hematoxylin and Eosin (H&E) staining, in order to validate its ability to map these distinct modalities into a unified vector space. Despite extreme class imbalance, the system demonstrated robustness and utility by returning similar results across various data features, which suggests potential for future use in multimodal histopathology data analysis.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge