Hua Xie
Pre-Training Graph Contrastive Masked Autoencoders are Strong Distillers for EEG
Nov 28, 2024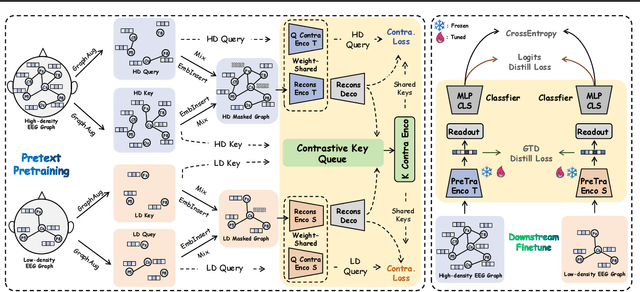
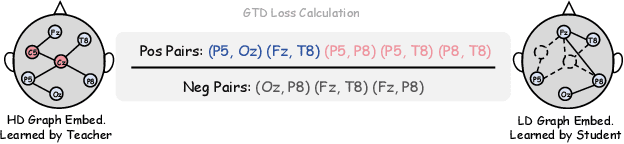
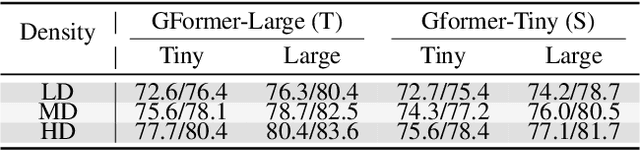
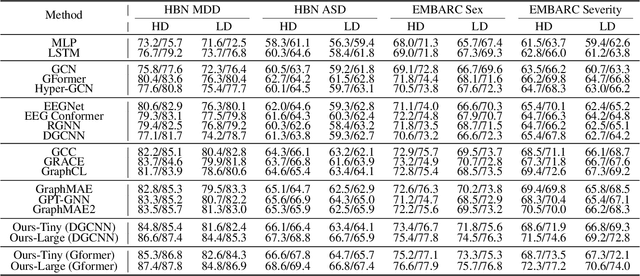


Abstract:Effectively utilizing extensive unlabeled high-density EEG data to improve performance in scenarios with limited labeled low-density EEG data presents a significant challenge. In this paper, we address this by framing it as a graph transfer learning and knowledge distillation problem. We propose a Unified Pre-trained Graph Contrastive Masked Autoencoder Distiller, named EEG-DisGCMAE, to bridge the gap between unlabeled/labeled and high/low-density EEG data. To fully leverage the abundant unlabeled EEG data, we introduce a novel unified graph self-supervised pre-training paradigm, which seamlessly integrates Graph Contrastive Pre-training and Graph Masked Autoencoder Pre-training. This approach synergistically combines contrastive and generative pre-training techniques by reconstructing contrastive samples and contrasting the reconstructions. For knowledge distillation from high-density to low-density EEG data, we propose a Graph Topology Distillation loss function, allowing a lightweight student model trained on low-density data to learn from a teacher model trained on high-density data, effectively handling missing electrodes through contrastive distillation. To integrate transfer learning and distillation, we jointly pre-train the teacher and student models by contrasting their queries and keys during pre-training, enabling robust distillers for downstream tasks. We demonstrate the effectiveness of our method on four classification tasks across two clinical EEG datasets with abundant unlabeled data and limited labeled data. The experimental results show that our approach significantly outperforms contemporary methods in both efficiency and accuracy.
Multi-modal Cross-domain Self-supervised Pre-training for fMRI and EEG Fusion
Sep 27, 2024Abstract:Neuroimaging techniques including functional magnetic resonance imaging (fMRI) and electroencephalogram (EEG) have shown promise in detecting functional abnormalities in various brain disorders. However, existing studies often focus on a single domain or modality, neglecting the valuable complementary information offered by multiple domains from both fMRI and EEG, which is crucial for a comprehensive representation of disorder pathology. This limitation poses a challenge in effectively leveraging the synergistic information derived from these modalities. To address this, we propose a Multi-modal Cross-domain Self-supervised Pre-training Model (MCSP), a novel approach that leverages self-supervised learning to synergize multi-modal information across spatial, temporal, and spectral domains. Our model employs cross-domain self-supervised loss that bridges domain differences by implementing domain-specific data augmentation and contrastive loss, enhancing feature discrimination. Furthermore, MCSP introduces cross-modal self-supervised loss to capitalize on the complementary information of fMRI and EEG, facilitating knowledge distillation within domains and maximizing cross-modal feature convergence. We constructed a large-scale pre-training dataset and pretrained MCSP model by leveraging proposed self-supervised paradigms to fully harness multimodal neuroimaging data. Through comprehensive experiments, we have demonstrated the superior performance and generalizability of our model on multiple classification tasks. Our study contributes a significant advancement in the fusion of fMRI and EEG, marking a novel integration of cross-domain features, which enriches the existing landscape of neuroimaging research, particularly within the context of mental disorder studies.
Improving EEG Decoding via Clustering-based Multi-task Feature Learning
Dec 12, 2020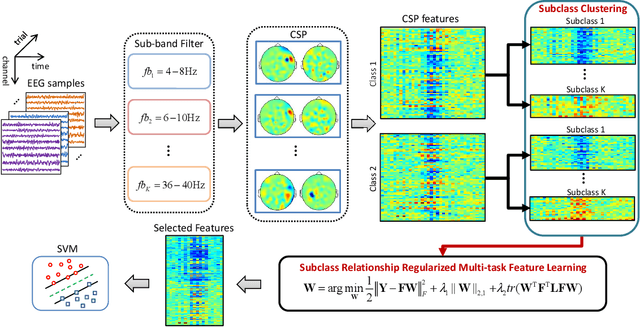
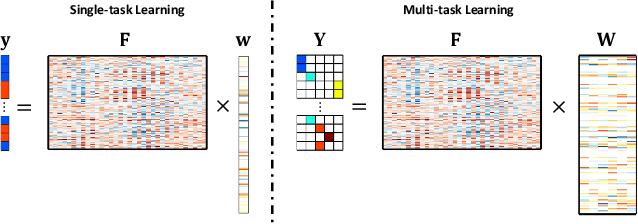
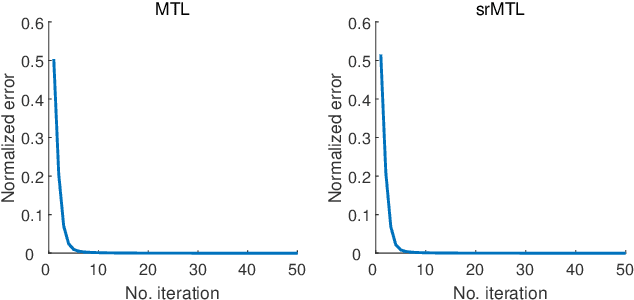
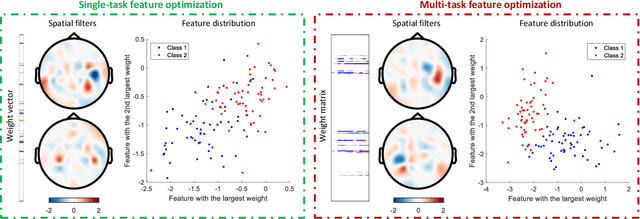
Abstract:Accurate electroencephalogram (EEG) pattern decoding for specific mental tasks is one of the key steps for the development of brain-computer interface (BCI), which is quite challenging due to the considerably low signal-to-noise ratio of EEG collected at the brain scalp. Machine learning provides a promising technique to optimize EEG patterns toward better decoding accuracy. However, existing algorithms do not effectively explore the underlying data structure capturing the true EEG sample distribution, and hence can only yield a suboptimal decoding accuracy. To uncover the intrinsic distribution structure of EEG data, we propose a clustering-based multi-task feature learning algorithm for improved EEG pattern decoding. Specifically, we perform affinity propagation-based clustering to explore the subclasses (i.e., clusters) in each of the original classes, and then assign each subclass a unique label based on a one-versus-all encoding strategy. With the encoded label matrix, we devise a novel multi-task learning algorithm by exploiting the subclass relationship to jointly optimize the EEG pattern features from the uncovered subclasses. We then train a linear support vector machine with the optimized features for EEG pattern decoding. Extensive experimental studies are conducted on three EEG datasets to validate the effectiveness of our algorithm in comparison with other state-of-the-art approaches. The improved experimental results demonstrate the outstanding superiority of our algorithm, suggesting its prominent performance for EEG pattern decoding in BCI applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge