Corwyn Rowsell
HistoKT: Cross Knowledge Transfer in Computational Pathology
Jan 27, 2022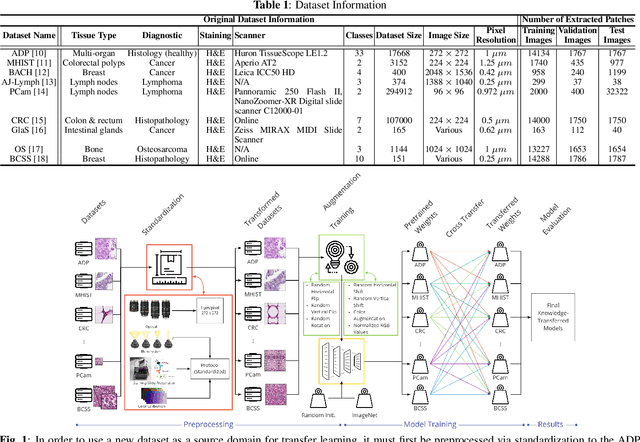



Abstract:The lack of well-annotated datasets in computational pathology (CPath) obstructs the application of deep learning techniques for classifying medical images. %Since pathologist time is expensive, dataset curation is intrinsically difficult. Many CPath workflows involve transferring learned knowledge between various image domains through transfer learning. Currently, most transfer learning research follows a model-centric approach, tuning network parameters to improve transfer results over few datasets. In this paper, we take a data-centric approach to the transfer learning problem and examine the existence of generalizable knowledge between histopathological datasets. First, we create a standardization workflow for aggregating existing histopathological data. We then measure inter-domain knowledge by training ResNet18 models across multiple histopathological datasets, and cross-transferring between them to determine the quantity and quality of innate shared knowledge. Additionally, we use weight distillation to share knowledge between models without additional training. We find that hard to learn, multi-class datasets benefit most from pretraining, and a two stage learning framework incorporating a large source domain such as ImageNet allows for better utilization of smaller datasets. Furthermore, we find that weight distillation enables models trained on purely histopathological features to outperform models using external natural image data.
Probeable DARTS with Application to Computational Pathology
Aug 16, 2021



Abstract:AI technology has made remarkable achievements in computational pathology (CPath), especially with the help of deep neural networks. However, the network performance is highly related to architecture design, which commonly requires human experts with domain knowledge. In this paper, we combat this challenge with the recent advance in neural architecture search (NAS) to find an optimal network for CPath applications. In particular, we use differentiable architecture search (DARTS) for its efficiency. We first adopt a probing metric to show that the original DARTS lacks proper hyperparameter tuning on the CIFAR dataset, and how the generalization issue can be addressed using an adaptive optimization strategy. We then apply our searching framework on CPath applications by searching for the optimum network architecture on a histological tissue type dataset (ADP). Results show that the searched network outperforms state-of-the-art networks in terms of prediction accuracy and computation complexity. We further conduct extensive experiments to demonstrate the transferability of the searched network to new CPath applications, the robustness against downscaled inputs, as well as the reliability of predictions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge