Chun-Fu Yeh
AnyMAL: An Efficient and Scalable Any-Modality Augmented Language Model
Sep 27, 2023



Abstract:We present Any-Modality Augmented Language Model (AnyMAL), a unified model that reasons over diverse input modality signals (i.e. text, image, video, audio, IMU motion sensor), and generates textual responses. AnyMAL inherits the powerful text-based reasoning abilities of the state-of-the-art LLMs including LLaMA-2 (70B), and converts modality-specific signals to the joint textual space through a pre-trained aligner module. To further strengthen the multimodal LLM's capabilities, we fine-tune the model with a multimodal instruction set manually collected to cover diverse topics and tasks beyond simple QAs. We conduct comprehensive empirical analysis comprising both human and automatic evaluations, and demonstrate state-of-the-art performance on various multimodal tasks.
Self-similarity Student for Partial Label Histopathology Image Segmentation
Jul 19, 2020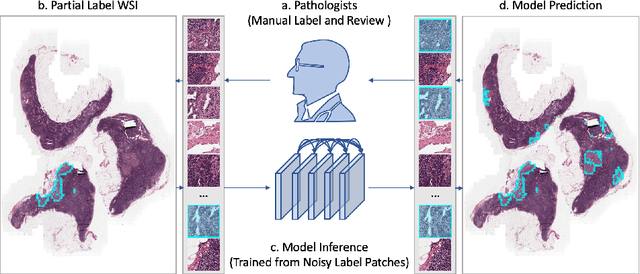
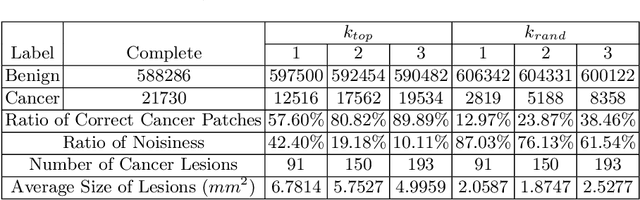
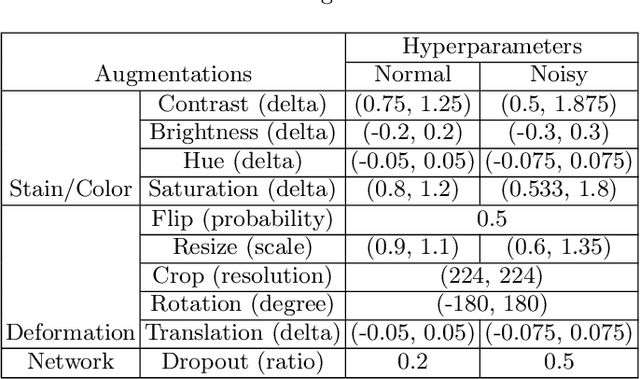
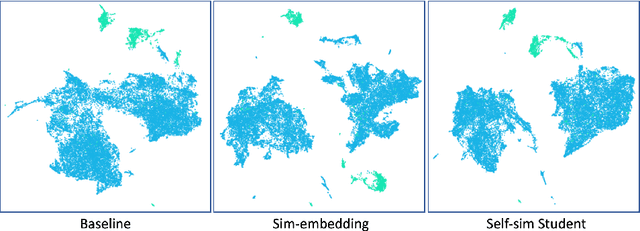
Abstract:Delineation of cancerous regions in gigapixel whole slide images (WSIs) is a crucial diagnostic procedure in digital pathology. This process is time-consuming because of the large search space in the gigapixel WSIs, causing chances of omission and misinterpretation at indistinct tumor lesions. To tackle this, the development of an automated cancerous region segmentation method is imperative. We frame this issue as a modeling problem with partial label WSIs, where some cancerous regions may be misclassified as benign and vice versa, producing patches with noisy labels. To learn from these patches, we propose Self-similarity Student, combining teacher-student model paradigm with similarity learning. Specifically, for each patch, we first sample its similar and dissimilar patches according to spatial distance. A teacher-student model is then introduced, featuring the exponential moving average on both student model weights and teacher predictions ensemble. While our student model takes patches, teacher model takes all their corresponding similar and dissimilar patches for learning robust representation against noisy label patches. Following this similarity learning, our similarity ensemble merges similar patches' ensembled predictions as the pseudo-label of a given patch to counteract its noisy label. On the CAMELYON16 dataset, our method substantially outperforms state-of-the-art noise-aware learning methods by 5$\%$ and the supervised-trained baseline by 10$\%$ in various degrees of noise. Moreover, our method is superior to the baseline on our TVGH TURP dataset with 2$\%$ improvement, demonstrating the generalizability to more clinical histopathology segmentation tasks.
A Cascaded Learning Strategy for Robust COVID-19 Pneumonia Chest X-Ray Screening
Apr 30, 2020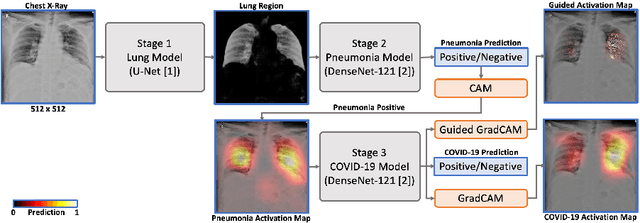
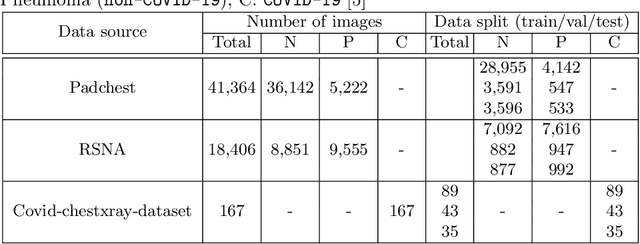
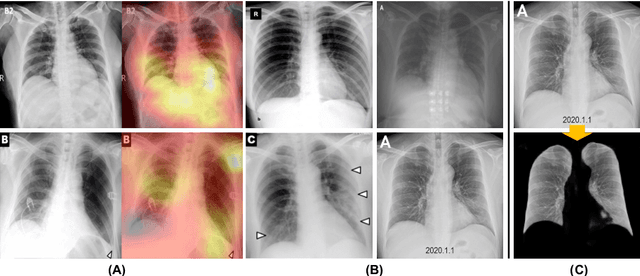
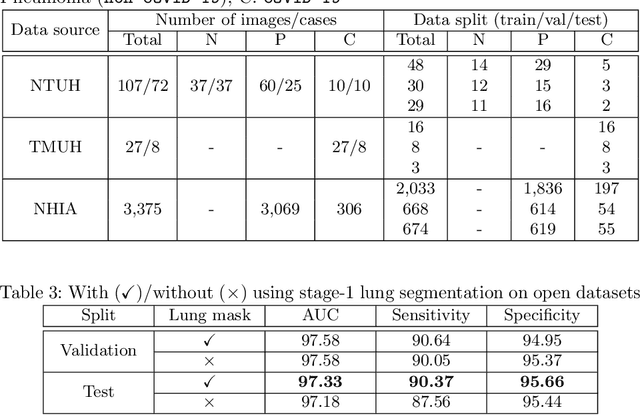
Abstract:We introduce a comprehensive screening platform for the COVID-19 (a.k.a., SARS-CoV-2) pneumonia. The proposed AI-based system works on chest x-ray (CXR) images to predict whether a patient is infected with the COVID-19 disease. Although the recent international joint effort on making the availability of all sorts of open data, the public collection of CXR images is still relatively small for reliably training a deep neural network (DNN) to carry out COVID-19 prediction. To better address such inefficiency, we design a cascaded learning strategy to improve both the sensitivity and the specificity of the resulting DNN classification model. Our approach leverages a large CXR image dataset of non-COVID-19 pneumonia to generalize the original well-trained classification model via a cascaded learning scheme. The resulting screening system is shown to achieve good classification performance on the expanded dataset, including those newly added COVID-19 CXR images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge