Benedek Fabian
Pseudo-Riemannian Embedding Models for Multi-Relational Graph Representations
Dec 02, 2022
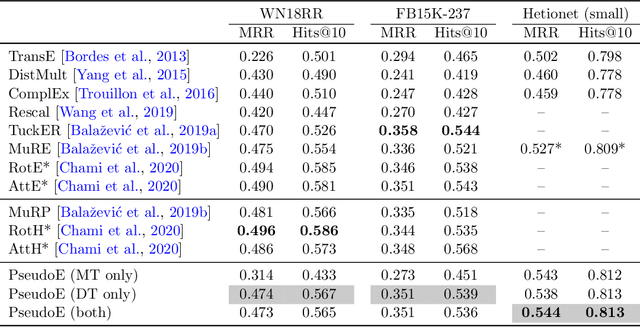
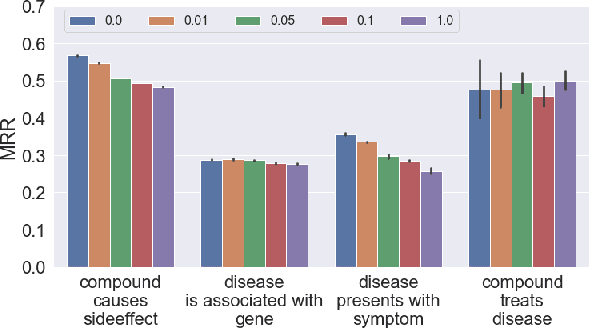
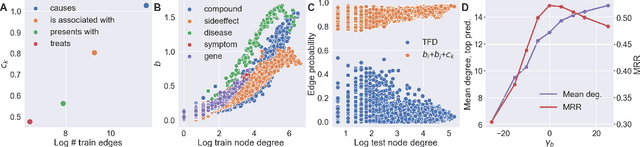
Abstract:In this paper we generalize single-relation pseudo-Riemannian graph embedding models to multi-relational networks, and show that the typical approach of encoding relations as manifold transformations translates from the Riemannian to the pseudo-Riemannian case. In addition we construct a view of relations as separate spacetime submanifolds of multi-time manifolds, and consider an interpolation between a pseudo-Riemannian embedding model and its Wick-rotated Riemannian counterpart. We validate these extensions in the task of link prediction, focusing on flat Lorentzian manifolds, and demonstrate their use in both knowledge graph completion and knowledge discovery in a biological domain.
* 11 pages, 3 figures, AKBC 2022 conference
Molecular representation learning with language models and domain-relevant auxiliary tasks
Nov 26, 2020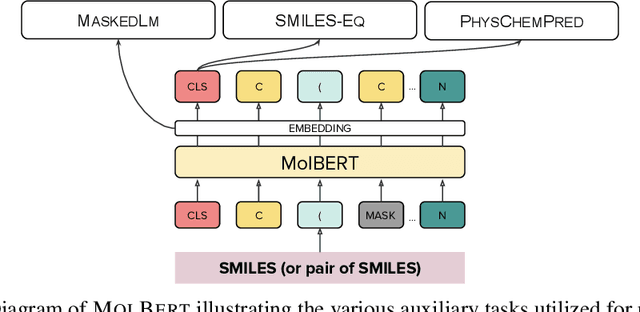
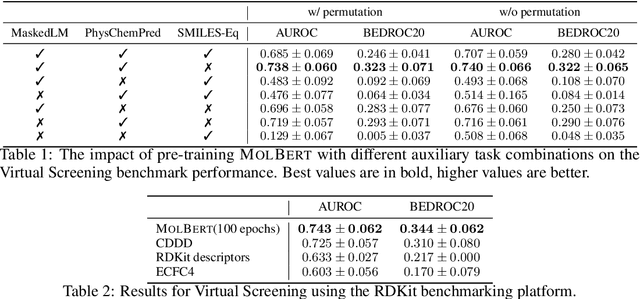

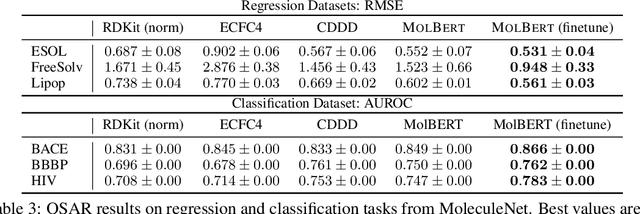
Abstract:We apply a Transformer architecture, specifically BERT, to learn flexible and high quality molecular representations for drug discovery problems. We study the impact of using different combinations of self-supervised tasks for pre-training, and present our results for the established Virtual Screening and QSAR benchmarks. We show that: i) The selection of appropriate self-supervised task(s) for pre-training has a significant impact on performance in subsequent downstream tasks such as Virtual Screening. ii) Using auxiliary tasks with more domain relevance for Chemistry, such as learning to predict calculated molecular properties, increases the fidelity of our learnt representations. iii) Finally, we show that molecular representations learnt by our model `MolBert' improve upon the current state of the art on the benchmark datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge