Beatrice S. Knudsen
Building Trust in Virtual Immunohistochemistry: Automated Assessment of Image Quality
Nov 06, 2025
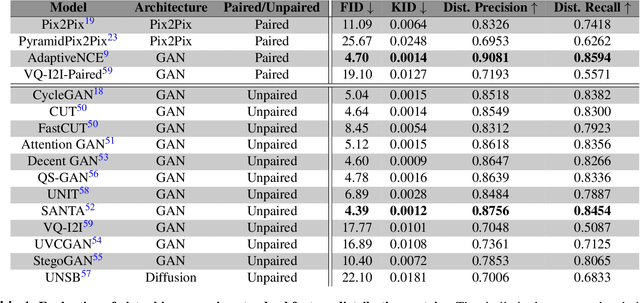
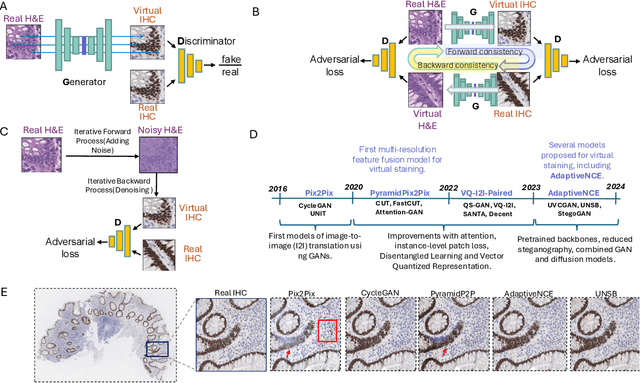

Abstract:Deep learning models can generate virtual immunohistochemistry (IHC) stains from hematoxylin and eosin (H&E) images, offering a scalable and low-cost alternative to laboratory IHC. However, reliable evaluation of image quality remains a challenge as current texture- and distribution-based metrics quantify image fidelity rather than the accuracy of IHC staining. Here, we introduce an automated and accuracy grounded framework to determine image quality across sixteen paired or unpaired image translation models. Using color deconvolution, we generate masks of pixels stained brown (i.e., IHC-positive) as predicted by each virtual IHC model. We use the segmented masks of real and virtual IHC to compute stain accuracy metrics (Dice, IoU, Hausdorff distance) that directly quantify correct pixel - level labeling without needing expert manual annotations. Our results demonstrate that conventional image fidelity metrics, including Frechet Inception Distance (FID), peak signal-to-noise ratio (PSNR), and structural similarity (SSIM), correlate poorly with stain accuracy and pathologist assessment. Paired models such as PyramidPix2Pix and AdaptiveNCE achieve the highest stain accuracy, whereas unpaired diffusion- and GAN-based models are less reliable in providing accurate IHC positive pixel labels. Moreover, whole-slide images (WSI) reveal performance declines that are invisible in patch-based evaluations, emphasizing the need for WSI-level benchmarks. Together, this framework defines a reproducible approach for assessing the quality of virtual IHC models, a critical step to accelerate translation towards routine use by pathologists.
DuoFormer: Leveraging Hierarchical Representations by Local and Global Attention Vision Transformer
Jun 15, 2025Abstract:Despite the widespread adoption of transformers in medical applications, the exploration of multi-scale learning through transformers remains limited, while hierarchical representations are considered advantageous for computer-aided medical diagnosis. We propose a novel hierarchical transformer model that adeptly integrates the feature extraction capabilities of Convolutional Neural Networks (CNNs) with the advanced representational potential of Vision Transformers (ViTs). Addressing the lack of inductive biases and dependence on extensive training datasets in ViTs, our model employs a CNN backbone to generate hierarchical visual representations. These representations are adapted for transformer input through an innovative patch tokenization process, preserving the inherited multi-scale inductive biases. We also introduce a scale-wise attention mechanism that directly captures intra-scale and inter-scale associations. This mechanism complements patch-wise attention by enhancing spatial understanding and preserving global perception, which we refer to as local and global attention, respectively. Our model significantly outperforms baseline models in terms of classification accuracy, demonstrating its efficiency in bridging the gap between Convolutional Neural Networks (CNNs) and Vision Transformers (ViTs). The components are designed as plug-and-play for different CNN architectures and can be adapted for multiple applications. The code is available at https://github.com/xiaoyatang/DuoFormer.git.
WeakSupCon: Weakly Supervised Contrastive Learning for Encoder Pre-training
Mar 06, 2025


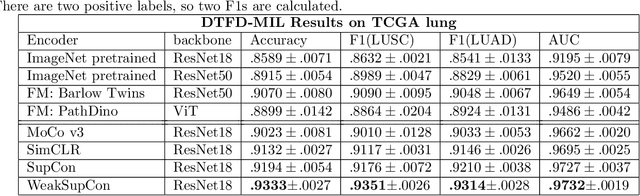
Abstract:Weakly supervised multiple instance learning (MIL) is a challenging task given that only bag-level labels are provided, while each bag typically contains multiple instances. This topic has been extensively studied in histopathological image analysis, where labels are usually available only at the whole slide image (WSI) level, while each whole slide image can be divided into thousands of small image patches for training. The dominant MIL approaches take fixed patch features as inputs to address computational constraints and ensure model stability. These features are commonly generated by encoders pre-trained on ImageNet, foundation encoders pre-trained on large datasets, or through self-supervised learning on local datasets. While the self-supervised encoder pre-training on the same dataset as downstream MIL tasks helps mitigate domain shift and generate better features, the bag-level labels are not utilized during the process, and the features of patches from different categories may cluster together, reducing classification performance on MIL tasks. Recently, pre-training with supervised contrastive learning (SupCon) has demonstrated superior performance compared to self-supervised contrastive learning and even end-to-end training on traditional image classification tasks. In this paper, we propose a novel encoder pre-training method for downstream MIL tasks called Weakly Supervised Contrastive Learning (WeakSupCon) that utilizes bag-level labels. In our method, we employ multi-task learning and define distinct contrastive learning losses for samples with different bag labels. Our experiments demonstrate that the features generated using WeakSupCon significantly enhance MIL classification performance compared to self-supervised approaches across three datasets.
SRA: A Novel Method to Improve Feature Embedding in Self-supervised Learning for Histopathological Images
Oct 31, 2024



Abstract:Self-supervised learning has become a cornerstone in various areas, particularly histopathological image analysis. Image augmentation plays a crucial role in self-supervised learning, as it generates variations in image samples. However, traditional image augmentation techniques often overlook the unique characteristics of histopathological images. In this paper, we propose a new histopathology-specific image augmentation method called stain reconstruction augmentation (SRA). We integrate our SRA with MoCo v3, a leading model in self-supervised contrastive learning, along with our additional contrastive loss terms, and call the new model SRA-MoCo v3. We demonstrate that our SRA-MoCo v3 always outperforms the standard MoCo v3 across various downstream tasks and achieves comparable or superior performance to other foundation models pre-trained on significantly larger histopathology datasets.
PathMoCo: A Novel Framework to Improve Feature Embedding in Self-supervised Contrastive Learning for Histopathological Images
Oct 23, 2024



Abstract:Self-supervised contrastive learning has become a cornerstone in various areas, particularly histopathological image analysis. Image augmentation plays a crucial role in self-supervised contrastive learning, as it generates variations in image samples. However, traditional image augmentation techniques often overlook the unique characteristics of histopathological images. In this paper, we propose a new histopathology-specific image augmentation method called stain reconstruction augmentation (SRA). We integrate our SRA with MoCo v3, a leading model in self-supervised contrastive learning, along with our additional contrastive loss terms, and call the new model PathMoCo. We demonstrate that our PathMoCo always outperforms the standard MoCo v3 across various downstream tasks and achieves comparable or superior performance to other foundation models pre-trained on significantly larger histopathology datasets.
DuoFormer: Leveraging Hierarchical Visual Representations by Local and Global Attention
Jul 18, 2024
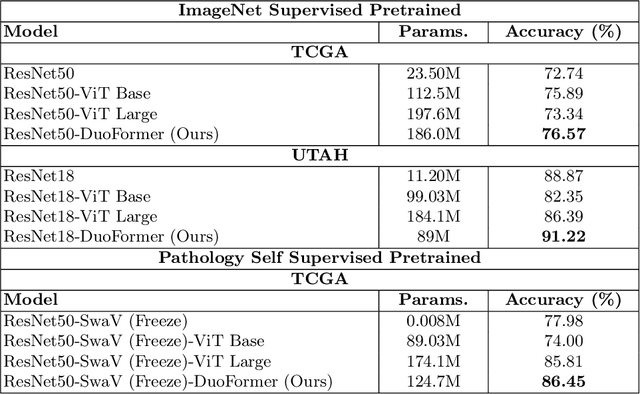

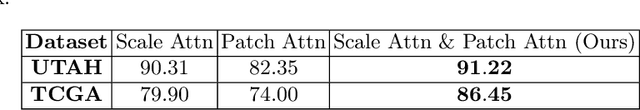
Abstract:We here propose a novel hierarchical transformer model that adeptly integrates the feature extraction capabilities of Convolutional Neural Networks (CNNs) with the advanced representational potential of Vision Transformers (ViTs). Addressing the lack of inductive biases and dependence on extensive training datasets in ViTs, our model employs a CNN backbone to generate hierarchical visual representations. These representations are then adapted for transformer input through an innovative patch tokenization. We also introduce a 'scale attention' mechanism that captures cross-scale dependencies, complementing patch attention to enhance spatial understanding and preserve global perception. Our approach significantly outperforms baseline models on small and medium-sized medical datasets, demonstrating its efficiency and generalizability. The components are designed as plug-and-play for different CNN architectures and can be adapted for multiple applications. The code is available at https://github.com/xiaoyatang/DuoFormer.git.
CLASS-M: Adaptive stain separation-based contrastive learning with pseudo-labeling for histopathological image classification
Jan 04, 2024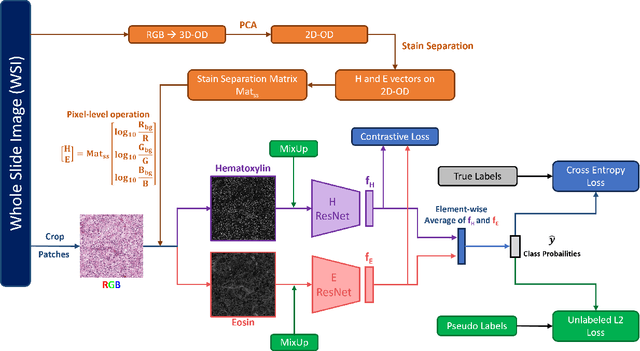
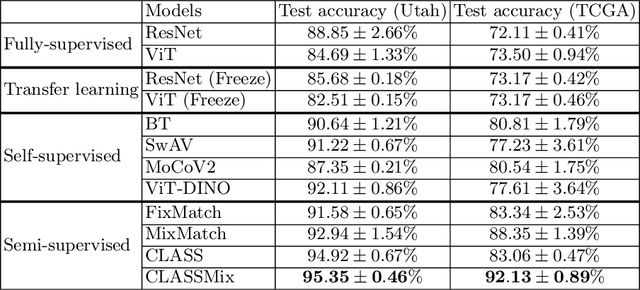
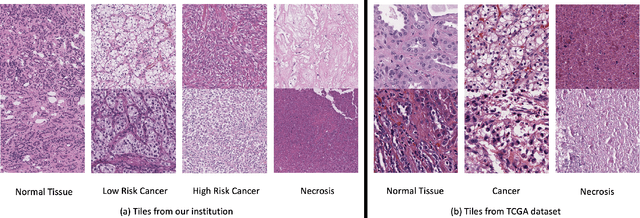
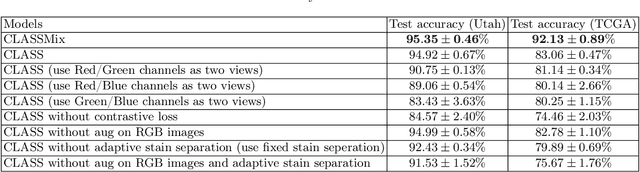
Abstract:Histopathological image classification is an important task in medical image analysis. Recent approaches generally rely on weakly supervised learning due to the ease of acquiring case-level labels from pathology reports. However, patch-level classification is preferable in applications where only a limited number of cases are available or when local prediction accuracy is critical. On the other hand, acquiring extensive datasets with localized labels for training is not feasible. In this paper, we propose a semi-supervised patch-level histopathological image classification model, named CLASS-M, that does not require extensively labeled datasets. CLASS-M is formed by two main parts: a contrastive learning module that uses separated Hematoxylin and Eosin images generated through an adaptive stain separation process, and a module with pseudo-labels using MixUp. We compare our model with other state-of-the-art models on two clear cell renal cell carcinoma datasets. We demonstrate that our CLASS-M model has the best performance on both datasets. Our code is available at github.com/BzhangURU/Paper_CLASS-M/tree/main
An attention-based multi-resolution model for prostate whole slide imageclassification and localization
May 30, 2019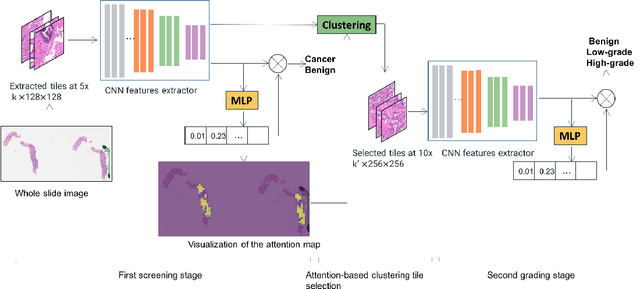

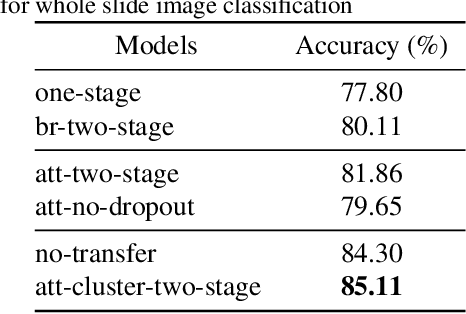
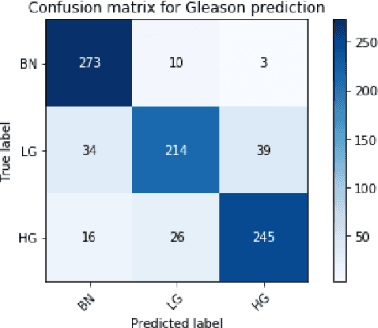
Abstract:Histology review is often used as the `gold standard' for disease diagnosis. Computer aided diagnosis tools can potentially help improve current pathology workflows by reducing examination time and interobserver variability. Previous work in cancer grading has focused mainly on classifying pre-defined regions of interest (ROIs), or relied on large amounts of fine-grained labels. In this paper, we propose a two-stage attention-based multiple instance learning model for slide-level cancer grading and weakly-supervised ROI detection and demonstrate its use in prostate cancer. Compared with existing Gleason classification models, our model goes a step further by utilizing visualized saliency maps to select informative tiles for fine-grained grade classification. The model was primarily developed on a large-scale whole slide dataset consisting of 3,521 prostate biopsy slides with only slide-level labels from 718 patients. The model achieved state-of-the-art performance for prostate cancer grading with an accuracy of 85.11\% for classifying benign, low-grade (Gleason grade 3+3 or 3+4), and high-grade (Gleason grade 4+3 or higher) slides on an independent test set.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge