Avantika Lal
Fine-Tuning Discrete Diffusion Models via Reward Optimization with Applications to DNA and Protein Design
Oct 17, 2024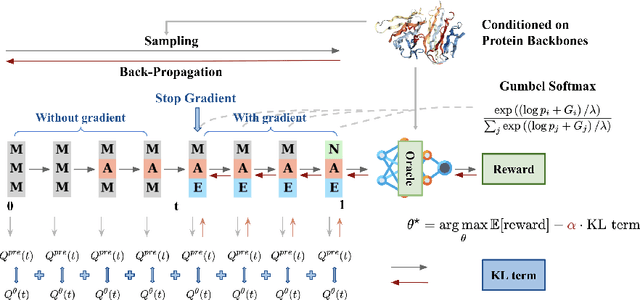
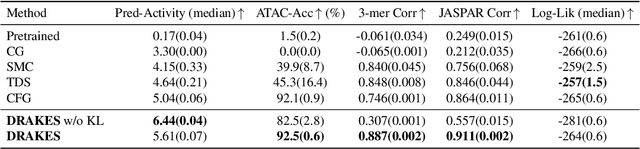
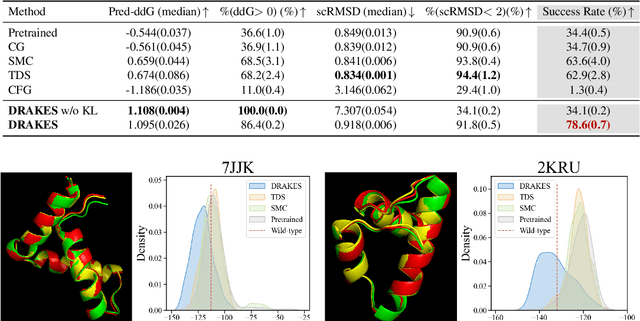

Abstract:Recent studies have demonstrated the strong empirical performance of diffusion models on discrete sequences across domains from natural language to biological sequence generation. For example, in the protein inverse folding task, conditional diffusion models have achieved impressive results in generating natural-like sequences that fold back into the original structure. However, practical design tasks often require not only modeling a conditional distribution but also optimizing specific task objectives. For instance, we may prefer protein sequences with high stability. To address this, we consider the scenario where we have pre-trained discrete diffusion models that can generate natural-like sequences, as well as reward models that map sequences to task objectives. We then formulate the reward maximization problem within discrete diffusion models, analogous to reinforcement learning (RL), while minimizing the KL divergence against pretrained diffusion models to preserve naturalness. To solve this RL problem, we propose a novel algorithm, DRAKES, that enables direct backpropagation of rewards through entire trajectories generated by diffusion models, by making the originally non-differentiable trajectories differentiable using the Gumbel-Softmax trick. Our theoretical analysis indicates that our approach can generate sequences that are both natural-like and yield high rewards. While similar tasks have been recently explored in diffusion models for continuous domains, our work addresses unique algorithmic and theoretical challenges specific to discrete diffusion models, which arise from their foundation in continuous-time Markov chains rather than Brownian motion. Finally, we demonstrate the effectiveness of DRAKES in generating DNA and protein sequences that optimize enhancer activity and protein stability, respectively, important tasks for gene therapies and protein-based therapeutics.
Bridging Model-Based Optimization and Generative Modeling via Conservative Fine-Tuning of Diffusion Models
May 31, 2024



Abstract:AI-driven design problems, such as DNA/protein sequence design, are commonly tackled from two angles: generative modeling, which efficiently captures the feasible design space (e.g., natural images or biological sequences), and model-based optimization, which utilizes reward models for extrapolation. To combine the strengths of both approaches, we adopt a hybrid method that fine-tunes cutting-edge diffusion models by optimizing reward models through RL. Although prior work has explored similar avenues, they primarily focus on scenarios where accurate reward models are accessible. In contrast, we concentrate on an offline setting where a reward model is unknown, and we must learn from static offline datasets, a common scenario in scientific domains. In offline scenarios, existing approaches tend to suffer from overoptimization, as they may be misled by the reward model in out-of-distribution regions. To address this, we introduce a conservative fine-tuning approach, BRAID, by optimizing a conservative reward model, which includes additional penalization outside of offline data distributions. Through empirical and theoretical analysis, we demonstrate the capability of our approach to outperform the best designs in offline data, leveraging the extrapolation capabilities of reward models while avoiding the generation of invalid designs through pre-trained diffusion models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge