Anita Burgun
HeKA, UPCité, CRC
Spectral structure learning for clinical time series
Feb 17, 2025
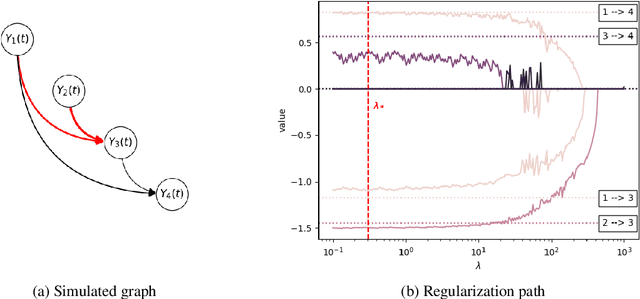

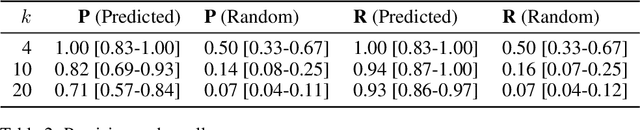
Abstract:We develop and evaluate a structure learning algorithm for clinical time series. Clinical time series are multivariate time series observed in multiple patients and irregularly sampled, challenging existing structure learning algorithms. We assume that our times series are realizations of StructGP, a k-dimensional multi-output or multi-task stationary Gaussian process (GP), with independent patients sharing the same covariance function. StructGP encodes ordered conditional relations between time series, represented in a directed acyclic graph. We implement an adapted NOTEARS algorithm, which based on a differentiable definition of acyclicity, recovers the graph by solving a series of continuous optimization problems. Simulation results show that up to mean degree 3 and 20 tasks, we reach a median recall of 0.93% [IQR, 0.86, 0.97] while keeping a median precision of 0.71% [0.57-0.84], for recovering directed edges. We further show that the regularization path is key to identifying the graph. With StructGP, we proposed a model of time series dependencies, that flexibly adapt to different time series regularity, while enabling us to learn these dependencies from observations.
The Smart Data Extractor, a Clinician Friendly Solution to Accelerate and Improve the Data Collection During Clinical Trials
Aug 31, 2023Abstract:In medical research, the traditional way to collect data, i.e. browsing patient files, has been proven to induce bias, errors, human labor and costs. We propose a semi-automated system able to extract every type of data, including notes. The Smart Data Extractor pre-populates clinic research forms by following rules. We performed a cross-testing experiment to compare semi-automated to manual data collection. 20 target items had to be collected for 79 patients. The average time to complete one form was 6'81'' for manual data collection and 3'22'' with the Smart Data Extractor. There were also more mistakes during manual data collection (163 for the whole cohort) than with the Smart Data Extractor (46 for the whole cohort). We present an easy to use, understandable and agile solution to fill out clinical research forms. It reduces human effort and provides higher quality data, avoiding data re-entry and fatigue induced errors.
Learning the grammar of prescription: recurrent neural network grammars for medication information extraction in clinical texts
Apr 24, 2020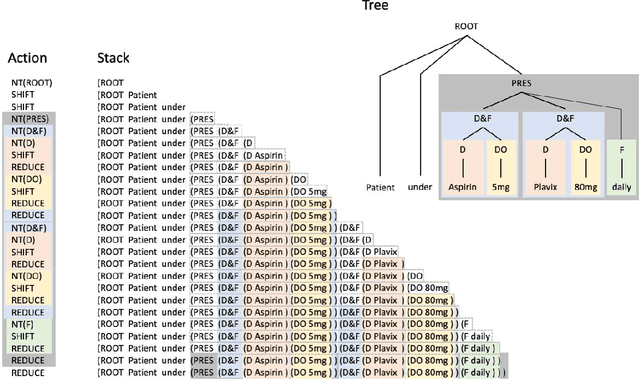
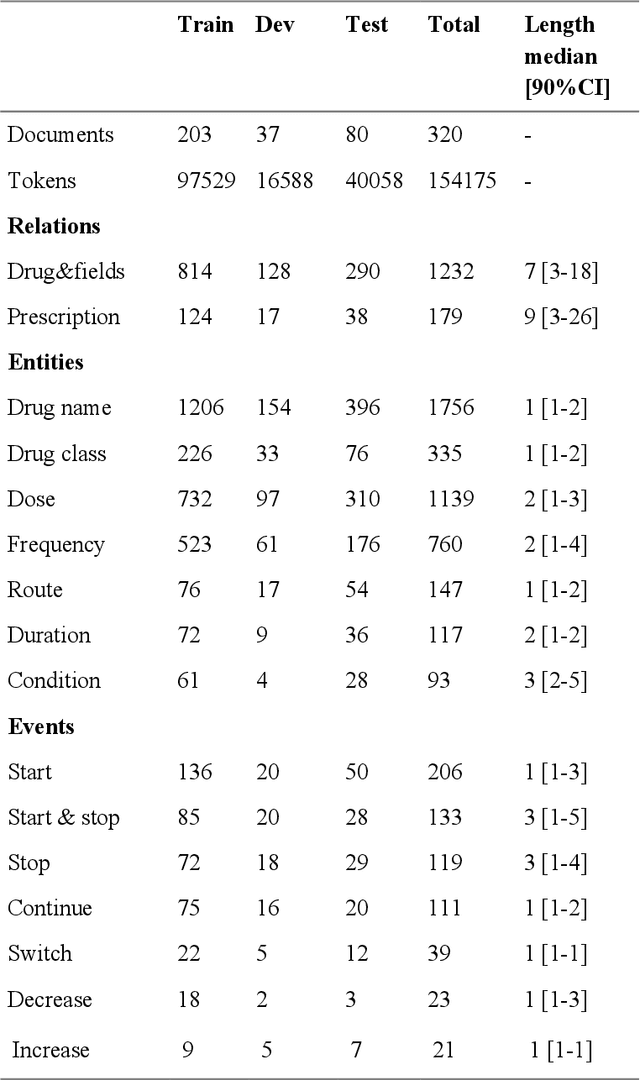
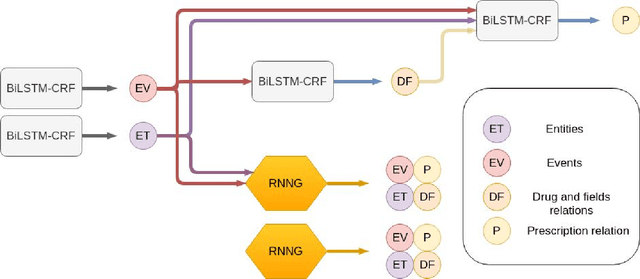
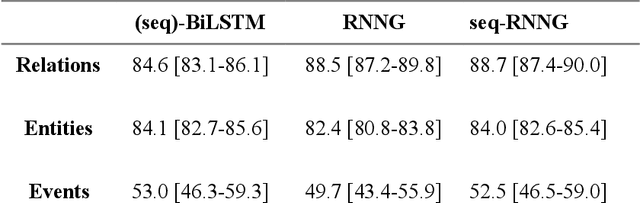
Abstract:In this study, we evaluated the RNNG, a neural top-down transition based parser, for medication information extraction in clinical texts. We evaluated this model on a French clinical corpus. The task was to extract the name of a drug (or class of drug), as well as fields informing its administration: frequency, dosage, duration, condition and route of administration. We compared the RNNG model that jointly identify entities and their relations with separate BiLSTMs models for entities and relations as baselines. We call seq-BiLSTMs the baseline models for relations extraction that takes as extra-input the output of the BiLSTMs for entities. RNNG outperforms seq-BiLSTM for identifying relations, with on average 88.5% [87.2-89.8] versus 84.6 [83.1-86.1] F-measure. However, RNNG is weaker than the baseline BiLSTM on detecting entities, with on average 82.4 [80.8-83.8] versus 84.1 [82.7-85.6] % F- measure. RNNG trained only for detecting relations is weaker than RNNG with the joint modelling objective, 87.4 [85.8-88.8] versus 88.5% [87.2-89.8]. The performance of RNNG on relations can be explained both by the model architecture, which provides shortcut between distant parts of the sentence, and the joint modelling objective which allow the RNNG to learn richer representations. RNNG is efficient for modeling relations between entities in medical texts and its performances are close to those of a BiLSTM for entity detection.
Natural language understanding for task oriented dialog in the biomedical domain in a low resources context
Nov 29, 2018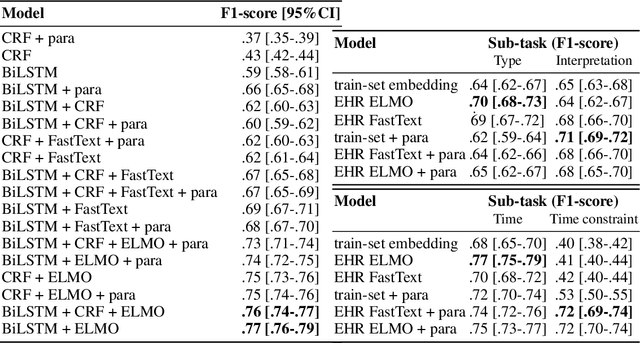
Abstract:In the biomedical domain, the lack of sharable datasets often limit the possibility of developing natural language processing systems, especially dialogue applications and natural language understanding models. To overcome this issue, we explore data generation using templates and terminologies and data augmentation approaches. Namely, we report our experiments using paraphrasing and word representations learned on a large EHR corpus with Fasttext and ELMo, to learn a NLU model without any available dataset. We evaluate on a NLU task of natural language queries in EHRs divided in slot-filling and intent classification sub-tasks. On the slot-filling task, we obtain a F-score of 0.76 with the ELMo representation; and on the classification task, a mean F-score of 0.71. Our results show that this method could be used to develop a baseline system.
Comparison of methods for early-readmission prediction in a high-dimensional heterogeneous covariates and time-to-event outcome framework
Jul 25, 2018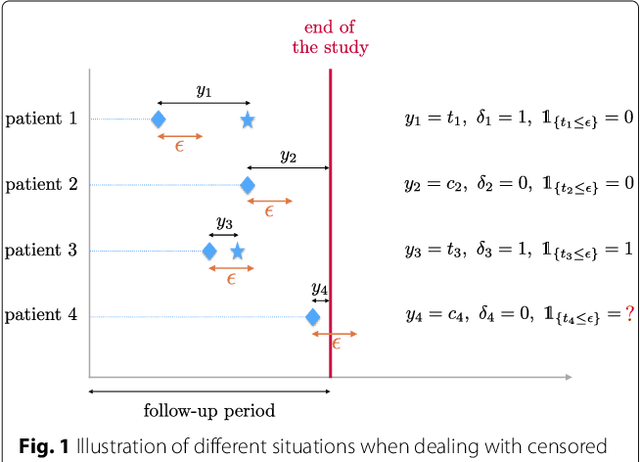
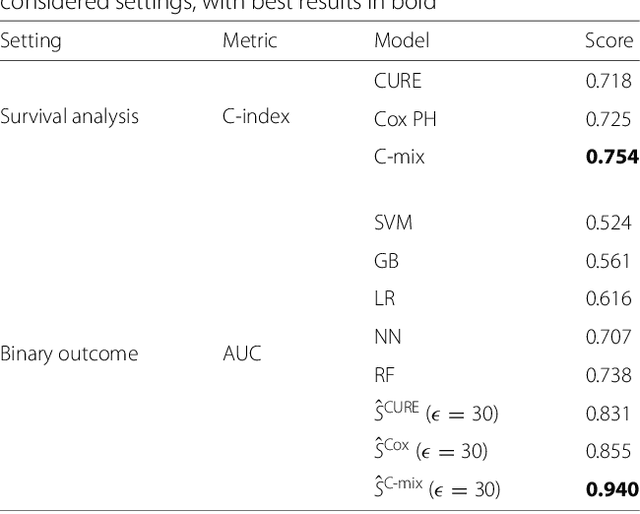
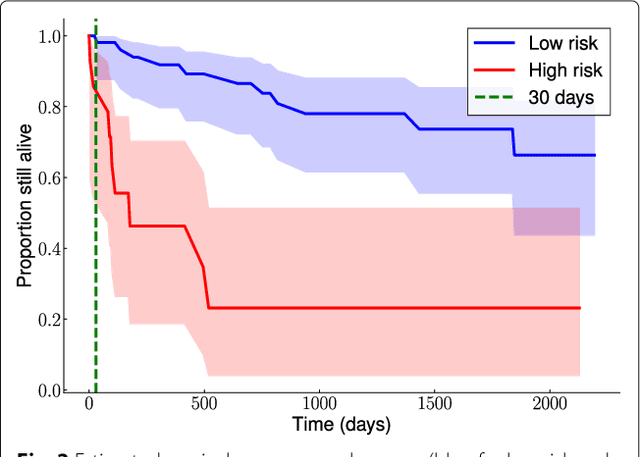
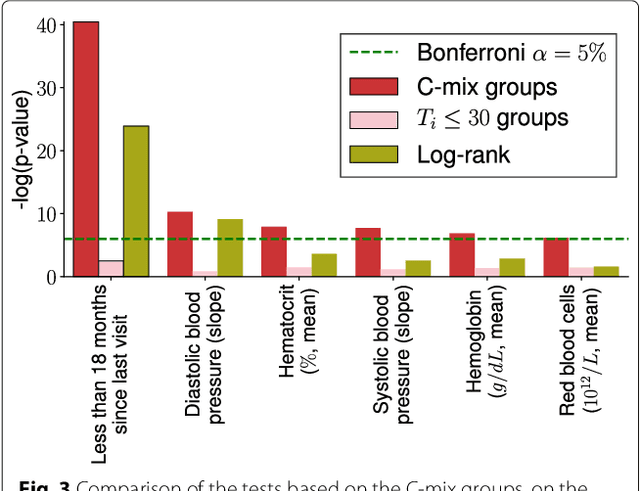
Abstract:Background: Choosing the most performing method in terms of outcome prediction or variables selection is a recurring problem in prognosis studies, leading to many publications on methods comparison. But some aspects have received little attention. First, most comparison studies treat prediction performance and variable selection aspects separately. Second, methods are either compared within a binary outcome setting (based on an arbitrarily chosen delay) or within a survival setting, but not both. In this paper, we propose a comparison methodology to weight up those different settings both in terms of prediction and variables selection, while incorporating advanced machine learning strategies. Methods: Using a high-dimensional case study on a sickle-cell disease (SCD) cohort, we compare 8 statistical methods. In the binary outcome setting, we consider logistic regression (LR), support vector machine (SVM), random forest (RF), gradient boosting (GB) and neural network (NN); while on the survival analysis setting, we consider the Cox Proportional Hazards (PH), the CURE and the C-mix models. We then compare performances of all methods both in terms of risk prediction and variable selection, with a focus on the use of Elastic-Net regularization technique. Results: Among all assessed statistical methods assessed, the C-mix model yields the better performances in both the two considered settings, as well as interesting interpretation aspects. There is some consistency in selected covariates across methods within a setting, but not much across the two settings. Conclusions: It appears that learning withing the survival setting first, and then going back to a binary prediction using the survival estimates significantly enhance binary predictions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge