Alexander Baras
AMIGO: Sparse Multi-Modal Graph Transformer with Shared-Context Processing for Representation Learning of Giga-pixel Images
Mar 01, 2023



Abstract:Processing giga-pixel whole slide histopathology images (WSI) is a computationally expensive task. Multiple instance learning (MIL) has become the conventional approach to process WSIs, in which these images are split into smaller patches for further processing. However, MIL-based techniques ignore explicit information about the individual cells within a patch. In this paper, by defining the novel concept of shared-context processing, we designed a multi-modal Graph Transformer (AMIGO) that uses the celluar graph within the tissue to provide a single representation for a patient while taking advantage of the hierarchical structure of the tissue, enabling a dynamic focus between cell-level and tissue-level information. We benchmarked the performance of our model against multiple state-of-the-art methods in survival prediction and showed that ours can significantly outperform all of them including hierarchical Vision Transformer (ViT). More importantly, we show that our model is strongly robust to missing information to an extent that it can achieve the same performance with as low as 20% of the data. Finally, in two different cancer datasets, we demonstrated that our model was able to stratify the patients into low-risk and high-risk groups while other state-of-the-art methods failed to achieve this goal. We also publish a large dataset of immunohistochemistry images (InUIT) containing 1,600 tissue microarray (TMA) cores from 188 patients along with their survival information, making it one of the largest publicly available datasets in this context.
Weakly Supervised Prostate TMA Classification via Graph Convolutional Networks
Nov 06, 2019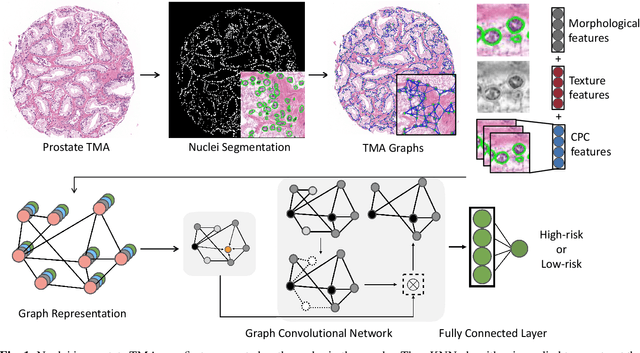

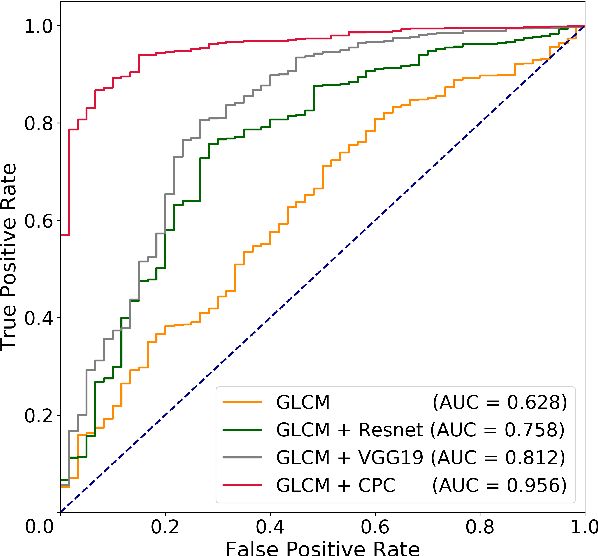
Abstract:Histology-based grade classification is clinically important for many cancer types in stratifying patients distinct treatment groups. In prostate cancer, the Gleason score is a grading system used to measure the aggressiveness of prostate cancer from the spatial organization of cells and the distribution of glands. However, the subjective interpretation of Gleason score often suffers from large interobserver and intraobserver variability. Previous work in deep learning-based objective Gleason grading requires manual pixel-level annotation. In this work, we propose a weakly-supervised approach for grade classification in tissue micro-arrays (TMA) using graph convolutional networks (GCNs), in which we model the spatial organization of cells as a graph to better capture the proliferation and community structure of tumor cells. As node-level features in our graph representation, we learn the morphometry of each cell using a contrastive predictive coding (CPC)-based self-supervised approach. We demonstrate that on a five-fold cross validation our method can achieve $0.9659\pm0.0096$ AUC using only TMA-level labels. Our method demonstrates a 39.80\% improvement over standard GCNs with texture features and a 29.27% improvement over GCNs with VGG19 features. Our proposed pipeline can be used to objectively stratify low and high risk cases, reducing inter- and intra-observer variability and pathologist workload.
Deep Adversarial Training for Multi-Organ Nuclei Segmentation in Histopathology Images
Oct 19, 2018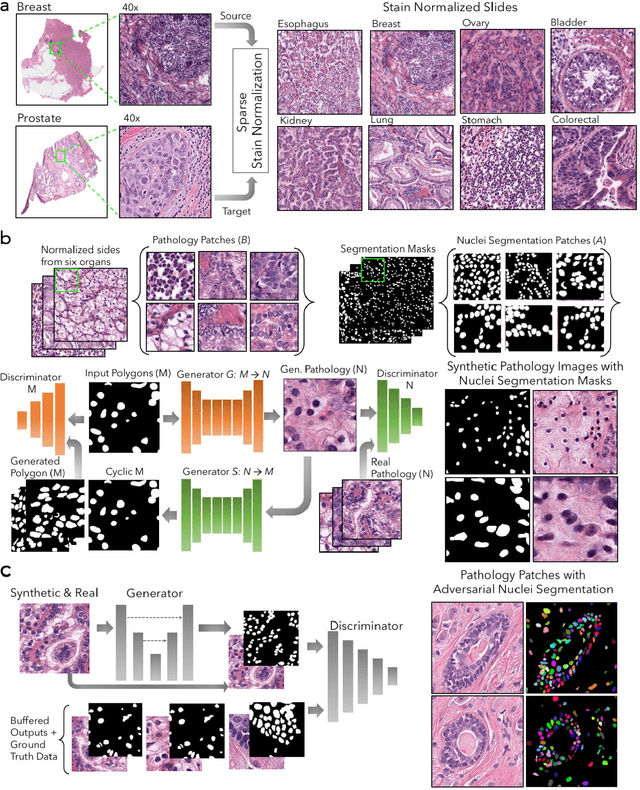
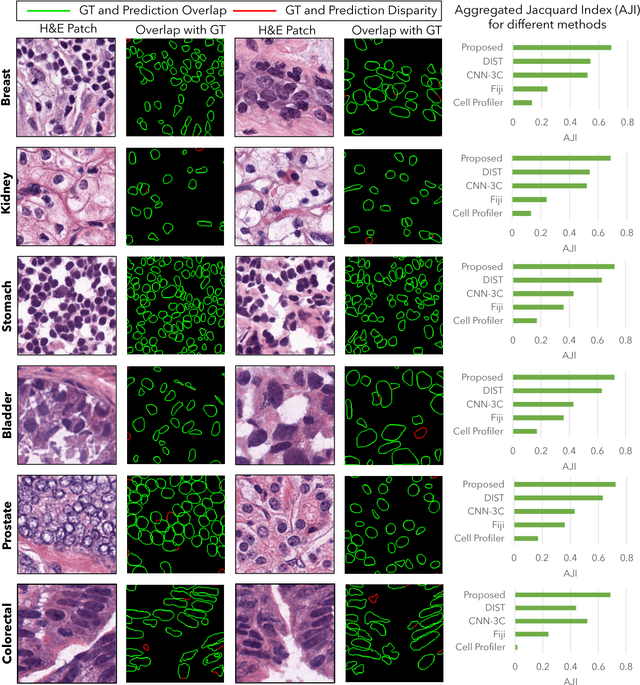
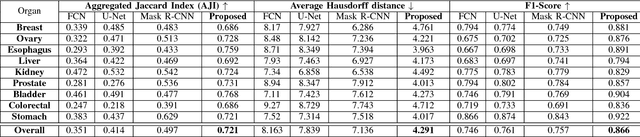
Abstract:Nuclei segmentation is a fundamental task that is critical for various computational pathology applications including nuclei morphology analysis, cell type classification, and cancer grading. Conventional vision-based methods for nuclei segmentation struggle in challenging cases and deep learning approaches have proven to be more robust and generalizable. However, CNNs require large amounts of labeled histopathology data. Moreover, conventional CNN-based approaches lack structured prediction capabilities which are required to distinguish overlapping and clumped nuclei. Here, we present an approach to nuclei segmentation that overcomes these challenges by utilizing a conditional generative adversarial network (cGAN) trained with synthetic and real data. We generate a large dataset of H&E training images with perfect nuclei segmentation labels using an unpaired GAN framework. This synthetic data along with real histopathology data from six different organs are used to train a conditional GAN with spectral normalization and gradient penalty for nuclei segmentation. This adversarial regression framework enforces higher order consistency when compared to conventional CNN models. We demonstrate that this nuclei segmentation approach generalizes across different organs, sites, patients and disease states, and outperforms conventional approaches, especially in isolating individual and overlapping nuclei.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge