for the Alzheimer's Disease Neuroimaging Initiative
Reproducible evaluation of classification methods in Alzheimer's disease: framework and application to MRI and PET data
Aug 20, 2018
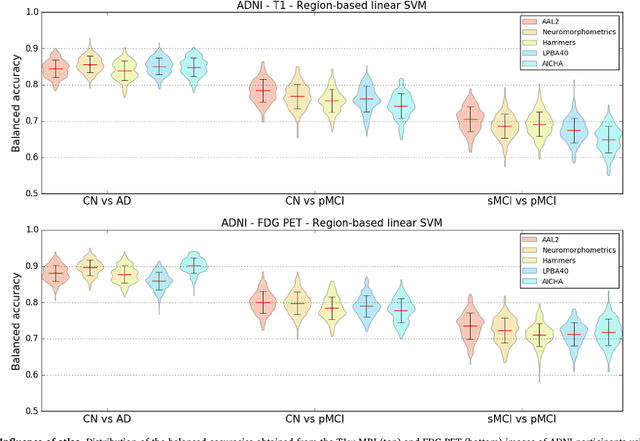
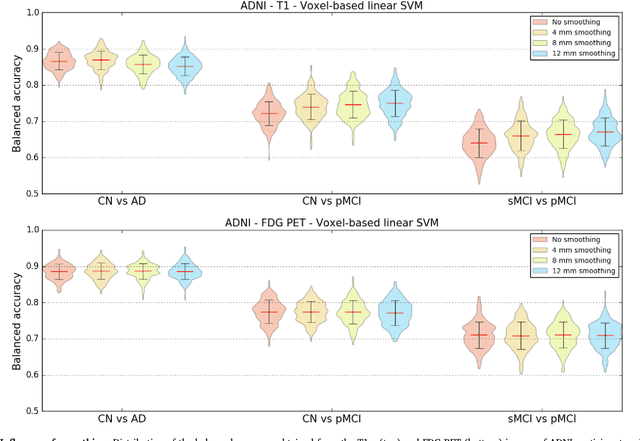
Abstract:A large number of papers have introduced novel machine learning and feature extraction methods for automatic classification of AD. However, they are difficult to reproduce because key components of the validation are often not readily available. These components include selected participants and input data, image preprocessing and cross-validation procedures. The performance of the different approaches is also difficult to compare objectively. In particular, it is often difficult to assess which part of the method provides a real improvement, if any. We propose a framework for reproducible and objective classification experiments in AD using three publicly available datasets (ADNI, AIBL and OASIS). The framework comprises: i) automatic conversion of the three datasets into BIDS format, ii) a modular set of preprocessing pipelines, feature extraction and classification methods, together with an evaluation framework, that provide a baseline for benchmarking the different components. We demonstrate the use of the framework for a large-scale evaluation on 1960 participants using T1 MRI and FDG PET data. In this evaluation, we assess the influence of different modalities, preprocessing, feature types, classifiers, training set sizes and datasets. Performances were in line with the state-of-the-art. FDG PET outperformed T1 MRI for all classification tasks. No difference in performance was found for the use of different atlases, image smoothing, partial volume correction of FDG PET images, or feature type. Linear SVM and L2-logistic regression resulted in similar performance and both outperformed random forests. The classification performance increased along with the number of subjects used for training. Classifiers trained on ADNI generalized well to AIBL and OASIS. All the code of the framework and the experiments is publicly available at: https://gitlab.icm-institute.org/aramislab/AD-ML.
Disease Progression Timeline Estimation for Alzheimer's Disease using Discriminative Event Based Modeling
Aug 10, 2018

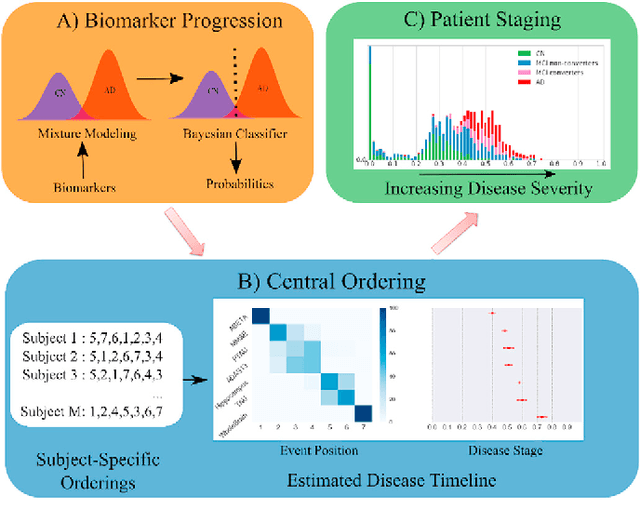
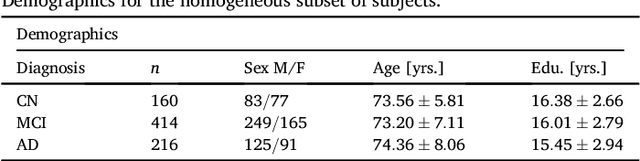
Abstract:Alzheimer's Disease (AD) is characterized by a cascade of biomarkers becoming abnormal, the pathophysiology of which is very complex and largely unknown. Event-based modeling (EBM) is a data-driven technique to estimate the sequence in which biomarkers for a disease become abnormal based on cross-sectional data. It can help in understanding the dynamics of disease progression and facilitate early diagnosis and prognosis. In this work we propose a novel discriminative approach to EBM, which is shown to be more accurate than existing state-of-the-art EBM methods. The method first estimates for each subject an approximate ordering of events. Subsequently, the central ordering over all subjects is estimated by fitting a generalized Mallows model to these approximate subject-specific orderings. We also introduce the concept of relative distance between events which helps in creating a disease progression timeline. Subsequently, we propose a method to stage subjects by placing them on the estimated disease progression timeline. We evaluated the proposed method on Alzheimer's Disease Neuroimaging Initiative (ADNI) data and compared the results with existing state-of-the-art EBM methods. We also performed extensive experiments on synthetic data simulating the progression of Alzheimer's disease. The event orderings obtained on ADNI data seem plausible and are in agreement with the current understanding of progression of AD. The proposed patient staging algorithm performed consistently better than that of state-of-the-art EBM methods. Event orderings obtained in simulation experiments were more accurate than those of other EBM methods and the estimated disease progression timeline was observed to correlate with the timeline of actual disease progression. The results of these experiments are encouraging and suggest that discriminative EBM is a promising approach to disease progression modeling.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge