Zhongnan Fang
Automated Real-time Assessment of Intracranial Hemorrhage Detection AI Using an Ensembled Monitoring Model (EMM)
May 16, 2025Abstract:Artificial intelligence (AI) tools for radiology are commonly unmonitored once deployed. The lack of real-time case-by-case assessments of AI prediction confidence requires users to independently distinguish between trustworthy and unreliable AI predictions, which increases cognitive burden, reduces productivity, and potentially leads to misdiagnoses. To address these challenges, we introduce Ensembled Monitoring Model (EMM), a framework inspired by clinical consensus practices using multiple expert reviews. Designed specifically for black-box commercial AI products, EMM operates independently without requiring access to internal AI components or intermediate outputs, while still providing robust confidence measurements. Using intracranial hemorrhage detection as our test case on a large, diverse dataset of 2919 studies, we demonstrate that EMM successfully categorizes confidence in the AI-generated prediction, suggesting different actions and helping improve the overall performance of AI tools to ultimately reduce cognitive burden. Importantly, we provide key technical considerations and best practices for successfully translating EMM into clinical settings.
Merlin: A Vision Language Foundation Model for 3D Computed Tomography
Jun 10, 2024



Abstract:Over 85 million computed tomography (CT) scans are performed annually in the US, of which approximately one quarter focus on the abdomen. Given the current radiologist shortage, there is a large impetus to use artificial intelligence to alleviate the burden of interpreting these complex imaging studies. Prior state-of-the-art approaches for automated medical image interpretation leverage vision language models (VLMs). However, current medical VLMs are generally limited to 2D images and short reports, and do not leverage electronic health record (EHR) data for supervision. We introduce Merlin - a 3D VLM that we train using paired CT scans (6+ million images from 15,331 CTs), EHR diagnosis codes (1.8+ million codes), and radiology reports (6+ million tokens). We evaluate Merlin on 6 task types and 752 individual tasks. The non-adapted (off-the-shelf) tasks include zero-shot findings classification (31 findings), phenotype classification (692 phenotypes), and zero-shot cross-modal retrieval (image to findings and image to impressions), while model adapted tasks include 5-year disease prediction (6 diseases), radiology report generation, and 3D semantic segmentation (20 organs). We perform internal validation on a test set of 5,137 CTs, and external validation on 7,000 clinical CTs and on two public CT datasets (VerSe, TotalSegmentator). Beyond these clinically-relevant evaluations, we assess the efficacy of various network architectures and training strategies to depict that Merlin has favorable performance to existing task-specific baselines. We derive data scaling laws to empirically assess training data needs for requisite downstream task performance. Furthermore, unlike conventional VLMs that require hundreds of GPUs for training, we perform all training on a single GPU.
Deep Learning Super-Resolution Enables Rapid Simultaneous Morphological and Quantitative Magnetic Resonance Imaging
Aug 07, 2018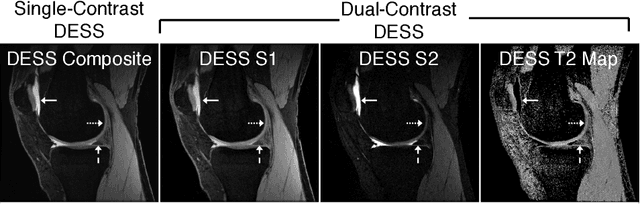


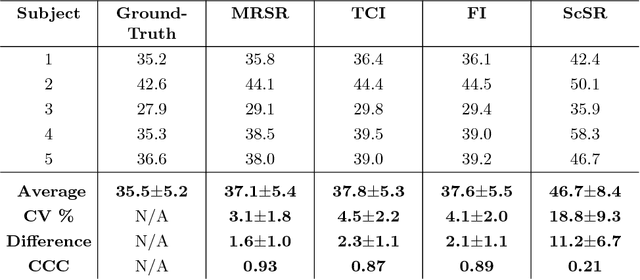
Abstract:Obtaining magnetic resonance images (MRI) with high resolution and generating quantitative image-based biomarkers for assessing tissue biochemistry is crucial in clinical and research applications. How- ever, acquiring quantitative biomarkers requires high signal-to-noise ratio (SNR), which is at odds with high-resolution in MRI, especially in a single rapid sequence. In this paper, we demonstrate how super-resolution can be utilized to maintain adequate SNR for accurate quantification of the T2 relaxation time biomarker, while simultaneously generating high- resolution images. We compare the efficacy of resolution enhancement using metrics such as peak SNR and structural similarity. We assess accuracy of cartilage T2 relaxation times by comparing against a standard reference method. Our evaluation suggests that SR can successfully maintain high-resolution and generate accurate biomarkers for accelerating MRI scans and enhancing the value of clinical and research MRI.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge