Yusong Lin
CLI-Gym: Scalable CLI Task Generation via Agentic Environment Inversion
Feb 11, 2026Abstract:Agentic coding requires agents to effectively interact with runtime environments, e.g., command line interfaces (CLI), so as to complete tasks like resolving dependency issues, fixing system problems, etc. But it remains underexplored how such environment-intensive tasks can be obtained at scale to enhance agents' capabilities. To address this, based on an analogy between the Dockerfile and the agentic task, we propose to employ agents to simulate and explore environment histories, guided by execution feedback. By tracing histories of a healthy environment, its state can be inverted to an earlier one with runtime failures, from which a task can be derived by packing the buggy state and the corresponding error messages. With our method, named CLI-Gym, a total of 1,655 environment-intensive tasks are derived, being the largest collection of its kind. Moreover, with curated successful trajectories, our fine-tuned model, named LiberCoder, achieves substantial absolute improvements of +21.1% (to 46.1%) on Terminal-Bench, outperforming various strong baselines. To our knowledge, this is the first public pipeline for scalable derivation of environment-intensive tasks.
Adaptive PromptNet For Auxiliary Glioma Diagnosis without Contrast-Enhanced MRI
Nov 15, 2022Abstract:Multi-contrast magnetic resonance imaging (MRI)-based automatic auxiliary glioma diagnosis plays an important role in the clinic. Contrast-enhanced MRI sequences (e.g., contrast-enhanced T1-weighted imaging) were utilized in most of the existing relevant studies, in which remarkable diagnosis results have been reported. Nevertheless, acquiring contrast-enhanced MRI data is sometimes not feasible due to the patients physiological limitations. Furthermore, it is more time-consuming and costly to collect contrast-enhanced MRI data in the clinic. In this paper, we propose an adaptive PromptNet to address these issues. Specifically, a PromptNet for glioma grading utilizing only non-enhanced MRI data has been constructed. PromptNet receives constraints from features of contrast-enhanced MR data during training through a designed prompt loss. To further boost the performance, an adaptive strategy is designed to dynamically weight the prompt loss in a sample-based manner. As a result, PromptNet is capable of dealing with more difficult samples. The effectiveness of our method is evaluated on a widely-used BraTS2020 dataset, and competitive glioma grading performance on NE-MRI data is achieved.
Expert Knowledge-guided Geometric Representation Learning for Magnetic Resonance Imaging-based Glioma Grading
Jan 08, 2022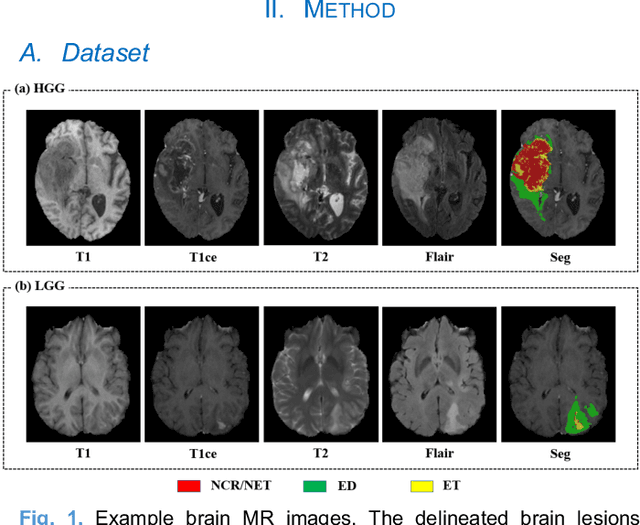
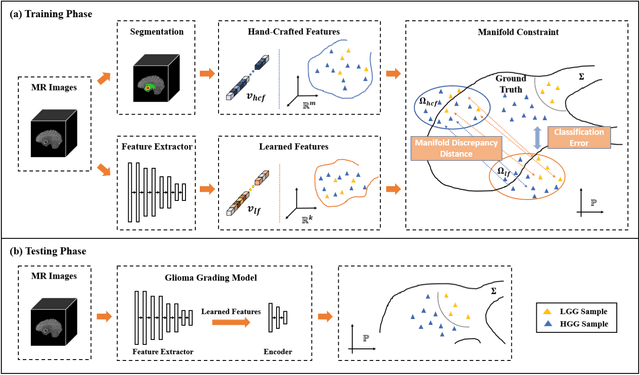
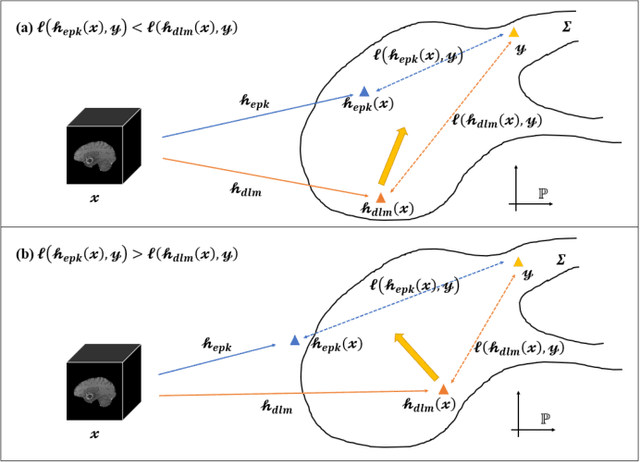
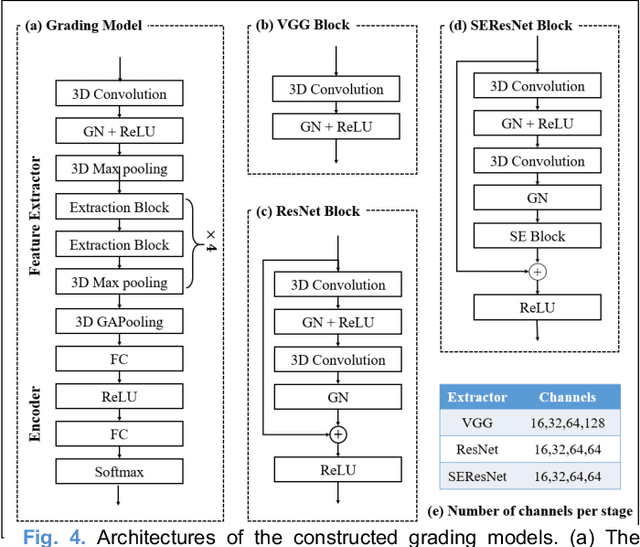
Abstract:Radiomics and deep learning have shown high popularity in automatic glioma grading. Radiomics can extract hand-crafted features that quantitatively describe the expert knowledge of glioma grades, and deep learning is powerful in extracting a large number of high-throughput features that facilitate the final classification. However, the performance of existing methods can still be improved as their complementary strengths have not been sufficiently investigated and integrated. Furthermore, lesion maps are usually needed for the final prediction at the testing phase, which is very troublesome. In this paper, we propose an expert knowledge-guided geometric representation learning (ENROL) framework . Geometric manifolds of hand-crafted features and learned features are constructed to mine the implicit relationship between deep learning and radiomics, and therefore to dig mutual consent and essential representation for the glioma grades. With a specially designed manifold discrepancy measurement, the grading model can exploit the input image data and expert knowledge more effectively in the training phase and get rid of the requirement of lesion segmentation maps at the testing phase. The proposed framework is flexible regarding deep learning architectures to be utilized. Three different architectures have been evaluated and five models have been compared, which show that our framework can always generate promising results.
Radiomic biomarker extracted from PI-RADS 3 patients support more eìcient and robust prostate cancer diagnosis: a multi-center study
Dec 23, 2021Abstract:Prostate Imaging Reporting and Data System (PI-RADS) based on multi-parametric MRI classi\^ees patients into 5 categories (PI-RADS 1-5) for routine clinical diagnosis guidance. However, there is no consensus on whether PI-RADS 3 patients should go through biopsies. Mining features from these hard samples (HS) is meaningful for physicians to achieve accurate diagnoses. Currently, the mining of HS biomarkers is insu\`icient, and the e\'eectiveness and robustness of HS biomarkers for prostate cancer diagnosis have not been explored. In this study, biomarkers from di\'eerent data distributions are constructed. Results show that HS biomarkers can achieve better performances in di\'eerent data distributions.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge